Abstract
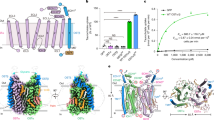
Bile acid and steroid hormone homeostasis are critical for human health, with disruptions linked to metabolic and endocrine disorders1,2. The organic solute transporter Ostα/β, essential for bile acid efflux in enterohepatic circulation3, has long defied mechanistic elucidation. Here we present cryogenic electron microscopy structures of human Ostα/β in apo and substrate-bound states at 2.6–3.1 Å resolution, revealing a distinctive membrane protein architecture that defines a new transporter class. Ostα/β forms a symmetric tetramer of heterodimers, with each Ostα subunit showing a new seven-transmembrane fold, augmented by a single transmembrane helix of Ostβ. This architecture is stabilized by extensive lipid modifications, including a palmitoylated cysteine-rich motif that forms a lateral substrate-binding groove. The structures uncover a unique transport pathway featuring two substrate-binding sites connected by an amphipathic helix-gated conduit. This design, conserved in the evolutionarily related TMEM184 family, suggests an ancient mechanism for substrate translocation. Electrophysiological studies demonstrate voltage-sensitive, bidirectional transport driven by electrochemical gradients, elucidating the efflux role of Ostα/β in vivo. Lipid interactions, notably palmitoylation-dependent trafficking, emerge as critical for stability and function. These findings clarify the molecular mechanism of Ostα/β, provide a structural basis for disease-associated mutations4,5 and establish a paradigm for lipid-modified membrane transport.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The coding sequences for human wild-type Ostα (UniProt Q86UW1) and Ostβ (UniProt Q86UW2) are available. The density maps and structure coordinates have been deposited to the Electron Microscopy Data Bank and the PDB under accession numbers EMD-64370 and 9UO2 (Ostα/β-apo complex); EMD-64364 and 9UNV (Ostα/β–TLCA complex); EMD-64369 and 9UO1 (Ostα/β–DHEAS complex). For MD simulations, the initial coordinate and simulation input files and a coordinate file of the final output are available at Zenodo (https://doi.org/10.5281/zenodo.17656875)57. Bioinformatic data and classification information were obtained from publicly available sources: TMEM184 family sequences from the National Center for Biotechnology Information, TMEM184A NP_001091089.1, TMEM184B NP_001182000.1 and TMEM184C NP_060711.2, TCDB classifications from the CDB through http://www.tcdb.org, accessed through 2.A.82.1, and domain annotations from Pfam at https://www.ebi.ac.uk/interpro/ (InterPro entry IPR005178). Foldseek is available through https://doi.org/10.1038/s41587-023-01773-0 and was used for sequence similarity searches. All other data supporting the findings of this study are available within the Article and its Supplementary Information. Any extra materials are available from the corresponding authors upon reasonable request. Source data are provided with this paper.
References
Deng, F. Y. & Bae, Y. H. Bile acid transporter-mediated oral drug delivery. J. Control. Release 327, 100–116 (2020).
Beaudoin, J. J., Brouwer, K. L. R. & Malinen, M. M. Novel insights into the organic solute transporter alpha/beta, OSTα/β: from the bench to the bedside. Pharmacol. Therapeut. https://doi.org/10.1016/j.pharmthera.2020.107542 (2020).
Ballatori, N., Christian, W. V., Wheeler, S. G. & Hammond, C. L. The heteromeric organic solute transporter, OSTα-OSTβ/SLC51: a transporter for steroid-derived molecules. Mol. Aspects Med. 34, 683–692 (2013).
Gao, E. et al. Organic solute transporter alpha deficiency: a disorder with cholestasis, liver fibrosis, and congenital diarrhea. Hepatology 71, 1879–1882 (2020).
Sultan, M. et al. Organic solute transporter-beta (SLC51B) deficiency in two brothers with congenital diarrhea and features of cholestasis. Hepatology 68, 590–598 (2018).
Collins, S. L., Stine, J. C., Bisanz, J. E., Okafor, C. D. & Patterson, A. D. Bile acids and the gut microbiota: metabolic interactions and impacts on disease. Nat. Rev. Microbiol. 21, 236–247 (2023).
Fleishman, J. S. & Kumar, S. Bile acid metabolism and signaling in health and disease: molecular mechanisms and therapeutic targets. Signal Transduct. Tar. https://doi.org/10.1038/s41392-024-01811-6 (2024).
Fang, F. et al. Neurosteroid transport by the organic solute transporter OSTα-OSTβ. J. Neurochem. 115, 220–233 (2010).
Boyer, J. L. & Soroka, C. J. Bile formation and secretion: an update. J. Hepatol. 75, 190–201 (2021).
Ferrebee, C. B. et al. Organic solute transporter α-β protects ileal enterocytes from bile acid-induced injury. Cell Mol. Gastroenter. 5, 499–522 (2018).
Goutam, K., Ielasi, F. S., Pardon, E., Steyaert, J. & Reyes, N. Structural basis of sodium-dependent bile salt uptake into the liver. Nature 606, 1015 (2022).
Liu, H. T. et al. Structure of human NTCP reveals the basis of recognition and sodium-driven transport of bile salts into the liver. Cell Res. 32, 773–776 (2022).
Shionoya, K. et al. Structural basis for hepatitis B virus restriction by a viral receptor homologue. Nat. Commun. 15, 9241 (2024).
Asami, J. et al. Structure of the bile acid transporter and HBV receptor NTCP. Nature 606, 1021–1026 (2022).
Liu, H. et al. Structure of antiviral drug bulevirtide bound to hepatitis B and D virus receptor protein NTCP. Nat. Commun. 15, 2476 (2024).
Hu, N. J., Iwata, S., Cameron, A. D. & Drew, D. Crystal structure of a bacterial homologue of the bile acid sodium symporter ASBT. Nature 478, 408 (2011).
Zhou, X. M. et al. Structural basis of the alternating-access mechanism in a bile acid transporter. Nature 505, 569–573 (2014).
Dawson, P. A. et al. The heteromeric organic solute transporter alpha-beta, Ostalpha-Ostbeta, is an ileal basolateral bile acid transporter. J. Biol. Chem. 280, 6960–6968 (2005).
Varadi, M. et al. AlphaFold Protein Structure Database: massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 50, D439–D444 (2022).
van Kempen, M. et al. Fast and accurate protein structure search with Foldseek. Nat. Biotechnol. 42, 243–246 (2024).
Gyimesi, G. & Hediger, M. A. Systematic in silico discovery of novel solute carrier-like proteins from proteomes. PLoS ONE 17, e0271062 (2022).
Pugh, R. J. et al. Transmembrane Protein 184A is a receptor required for vascular smooth muscle cell responses to heparin. J. Biol. Chem. 291, 5326–5341 (2016).
Chapman, K. A. et al. Pathogenic variants in TMEM184B cause a neurodevelopmental syndrome associated with alteration of metabolic signaling. Am. J. Hum. Genet. 112, 2381–2401 (2025).
Bhattacharya, M. R. C. et al. TMEM184b promotes axon degeneration and neuromuscular junction maintenance. J. Neurosci. 36, 4681–4689 (2016).
Suga, T., Yamaguchi, H., Ogura, J. & Mano, N. Characterization of conjugated and unconjugated bile acid transport via human organic solute transporter alpha/beta. Biochim. Biophys. Acta Biomembr. 1861, 1023–1029 (2019).
Holm, L., Laiho, A., Toronen, P. & Salgado, M. DALI shines a light on remote homologs: one hundred discoveries. Protein Sci. 32, e4519 (2023).
Kanada, S., Takeguchi, Y., Murakami, M., Ihara, K. & Kouyama, T. Crystal structures of an O-like blue form and an anion-free yellow form of pharaonis halorhodopsin. J. Mol. Biol. 413, 162–176 (2011).
Ballatori, N. Biology of a novel organic solute and steroid transporter, OSTalpha-OSTbeta. Exp. Biol. Med. 230, 689–698 (2005).
McCarthy, A. E., Yoshioka, C. & Mansoor, S. E. Full-length P2XStructures reveal how palmitoylation prevents channel desensitization. Cell 179, 659 (2019).
Drew, D., North, R. A., Nagarathinam, K. & Tanabe, M. Structures and general transport mechanisms by the Major Facilitator Superfamily (MFS). Chem. Rev. 121, 5289–5335 (2021).
Deng, D. et al. Crystal structure of the human glucose transporter GLUT1. Nature 510, 121–125 (2014).
Zeng, Y. C. et al. Structural basis of promiscuous substrate transport by Organic Cation Transporter 1. Nat. Commun. 14, 6374 (2023).
Varadi, M. et al. AlphaFold protein structure database in 2024: providing structure coverage for over 214 million protein sequences. Nucleic Acids Res. 52, D368–D375 (2024).
Padan, E. & Michel, H. NhaA: a unique structural fold of secondary active transporters. Isr. J. Chem. 55, 1233–1239 (2015).
Sakuragi, T. & Nagata, S. Regulation of phospholipid distribution in the lipid bilayer by flippases and scramblases (Apr 2023, 10.1038/s41580-02300604-z). Nat. Rev. Mol. Cell Bio. 24, 597–597 (2023).
Bushell, K. S. R. et al. The structural basis of lipid scrambling and inactivation in the endoplasmic reticulum scramblase TMEM16K. Nat. Commun. 10, 3956 (2019).
Sakuragi, T. et al. The tertiary structure of the human Xkr8-Basigin complex that scrambles phospholipids at plasma membranes. Nat. Struct. Mol. Biol. 28, 825 (2021).
Kasimova, M. A., Lindahl, E. & Delemotte, L. Determining the molecular basis of voltage sensitivity in membrane proteins. J. Gen. Physiol. 150, 1444–1458 (2018).
English, N. J. & Waldron, C. J. Perspectives on external electric fields in molecular simulation: progress, prospects and challenges. Phys. Chem. Chem. Phys. 17, 12407–12440 (2015).
Wang, W., Seward, D. J., Li, L., Boyer, J. L. & Ballatori, N. Expression cloning of two genes that together mediate organic solute and steroid transport in the liver of a marine vertebrate. Proc. Natl Acad. Sci. USA 98, 9431–9436 (2001).
Ballatori, N. et al. OSTalpha-OSTbeta: a major basolateral bile acid and steroid transporter in human intestinal, renal, and biliary epithelia. Hepatology 42, 1270–1279 (2005).
Yan, R., Zhao, X., Lei, J. & Zhou, Q. Structure of the human LAT1-4F2hc heteromeric amino acid transporter complex. Nature 568, 127–130 (2019).
Maxfield, F. R. & Wustner, D. Intracellular cholesterol transport. J. Clin. Invest. 110, 891–898 (2002).
Lu, Y. et al. Characterization of a novel organic solute transporter homologue from Clonorchis sinensis. PLoS Negl. Trop. Dis. 12, e0006459 (2018).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Malinen, M. M., Ali, I., Bezencon, J., Beaudoin, J. J. & Brouwer, K. L. R. Organic solute transporter OSTalpha/beta is overexpressed in nonalcoholic steatohepatitis and modulated by drugs associated with liver injury. Am. J. Physiol. Gastrointest. Liver Physiol. 314, G597–G609 (2018).
Schott-Verdugo, S. & Gohlke, H. PACKMOL-Memgen: a simple-to-use, generalized workflow for membrane-protein-lipid-bilayer system building. J. Chem. Inf. Model. 59, 2522–2528 (2019).
Tian, C. et al. ff19SB: amino-acid-specific protein backbone parameters trained against quantum mechanics energy surfaces in solution. J. Chem. Theory Comput. 16, 528–552 (2020).
Dickson, C. J., Walker, R. C. & Gould, I. R. Lipid21: complex lipid membrane simulations with AMBER. J. Chem. Theory Comput. 18, 1726–1736 (2022).
He, X., Man, V. H., Yang, W., Lee, T. S. & Wang, J. A fast and high-quality charge model for the next generation general AMBER force field. J. Chem. Phys. 153, 114502 (2020).
Alenaizan, A., Burns, L. A. & Sherrill, C. D. Python implementation of the restrained electrostatic potential charge model. Int. J. Quantum Chem. 120, e26035 (2020).
Olsson, M. H. M., Søndergaard, C. R., Rostkowski, M. & Jensen, J. H. PROPKA3: consistent treatment of internal and surface residues in empirical pKa predictions. J. Chem. Theory Comput. 7, 525–537 (2011).
Lu, S. et al. Activation pathway of a G protein-coupled receptor uncovers conformational intermediates as targets for allosteric drug design. Nat. Commun. 12, 4721 (2021).
He, X. et al. Conformational selection mechanism provides structural insights into the optimization of APC-Asef inhibitors. Molecules https://doi.org/10.3390/molecules26040962 (2021).
Salomon-Ferrer, R., Götz, A. W., Poole, D., Le Grand, S. & Walker, R. C. Routine microsecond molecular dynamics simulations with AMBER on GPUs. 2. explicit solvent particle mesh Ewald. J. Chem. Theory Comput. 9, 3878–3888 (2013).
Roe, D. R. & Cheatham, T. E. III. PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
He, X. Simulation files for Ostα/β. Zenodo https://doi.org/10.5281/zenodo.17656875 (2025).
Acknowledgements
The cryo-EM data were collected at the Advanced Center for Electron Microscopy, Shanghai Institute of Materia Medica (SIMM). We thank all staff, especially W. Hu, K. Wu and S. Li at the institution, for their assistance in cryo-EM data collection. Thank you to J. Li for the generous advice on data processing. We express our appreciation for the provision of experimental instruments from the Experimental Nuclear Medicine Laboratory, Core Facility of Basic Medical Sciences, Shanghai Jiao Tong University School of Medicine. We express our appreciation to H. Song for providing experimental instruments. This work was supported by the National Natural Science Foundation of China (82495184, 32130022, 82121005 to H.E.X.; 82130018 to X.M.; 32301016 to C.W.), the National Key R&D Program of China (2022YFA1302900 to W.Y., 2022YFC2703105 to H.E.X.), National Key R&D Program “Strategic Scientific and Technological Innovation Cooperation” Key Project (2022YFE0203600) released by the Ministry of Science and Technology, CAS Strategic Priority Research Program (XDB37030103 to H.E.X.), Shanghai Municipal Science and Technology Major Project (2019SHZDZX02 to H.E.X.), Shanghai Municipal Science and Technology Major Project (H.E.X.) and the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB0830000 to H.E.X.).
Author information
Authors and Affiliations
Contributions
X.Y. designed the expression constructs, purified the Ostα/β protein samples, prepared cryo-EM grids, calculated cryo-EM data, built and refined structural models, and prepared figures and wrote the initial article draft. N.C. conducted the [3H]TCA transporter uptake assay, flow cytometry and immunofluorescence analysis, co-immunoprecipitation and prepared corresponding figures and methods supervised by X.M. T.L. conducted the electrophysiological recordings and prepared corresponding methods and figures, supervised by Y.L. X.H. performed the molecular dynamics simulations and prepared corresponding methods, figures and videos. H.Z. participated in the discussion on experiment design and structural analysis. C.W. participated in protein purification and sample preparation. H.E.X., in collaboration with X.M., supervised the project. H.E.X. modified the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Palmitoylation influences Ostα subcellular localization.
a Subcellular localization of Ostα (green), Ostβ-mCherry (red), plasma membrane (orange) and nuclei (blue). Scale bar = 10 μm. Results are representative of three independent experiments. b Plasma membrane Ostα expression in WT and mutants quantified by flow cytometry (FACS). Data are mean ± s.e.m. (n = 4 biological replicates per group, ****P < 0.0001). c Co-immunoprecipitation (IP) analysis of overexpressed Ostβ and Ostα mutants in HEK293T cells. Results are representative of two independent experiments. d Quantification of total Ostα expression levels assessed by FACS in WT and mutants. Data are mean ± s.e.m. (n = 3 biological replicates per group). The P values for comparisons between Ostα/β and the respective conditions were 0.9936, 0.4073, 0.3491, and 0.9703. e Inhibition of Ostα/β transport activity by 2-BP. [³H]-TCA uptake measured after 2-BP treatment. Data are mean ± s.e.m. (n = 6 biological replicates per group, **P = 0.0015; ****P < 0.0001). f Plasma membrane Ostα expression levels assessed by FACS in WT and 2-BP treatment, normalized to untreated WT Ostα/β. Data are mean ± s.e.m. (n = 3 biological replicates per group, ***P = 0.0009; ****P < 0.0001). g Dose-dependent degradation of Ostα in response to 2-BP treatment. β-actin and Na+/K+ ATPase served as controls. h Total Ostα expression levels assessed by FACS in WT and 2-BP treatment, normalized to untreated WT Ostα/β. Data are mean ± s.e.m. (n = 3 biological replicates per group, ****P < 0.0001). b, d-f, h Results are representative of three independent experiments. Data analyzed by one-way ANOVA with Fisher’s LSD test.
Extended Data Fig. 2 Cholic acids binding site in different transporters.
a-b Surface representation of Ostα/βTLCA structure. The TLCA in magenta are shown in ball-stick model. c Hydrophilic surface of Ostα/βTLCA, same view as (b). d Inward conformation comparison between NTCP (PDB: 7ZYI) and ASBTnm (PDB: 3ZUX). e Surface representation of NTCP. Two glyco-chenodeoxycholic acid (GCDC) are coloured in yellow in ball-stick model. Cholesterol is coloured in light gray. f Hydrophilic surface of NTCP. g Surface representation of ASBTnm. Taurocholate is coloured in yellow in ball-stick model. Lipids are coloured in light gray in stick model. h Hydrophilic surface of ASBTnm.
Extended Data Fig. 3 Comparison of 3D maps and models refined with C1 and C2 symmetry.
Density maps and their corresponding atomic models were independently refined under C1 and C2 symmetry. The C2 maps are shown in gray with 50% transparency. C1 maps were generated de novo from the same dataset. All maps are visualized at a contour level of 0.35 with dust removal threshold set to 5. Subtle local differences between the two reconstructions are indicated by red arrows.
Extended Data Fig. 4 Structure alignment and ICH domain interaction with surrounding environment.
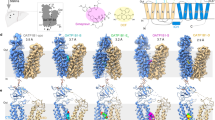
a Overall structure alignment among Ostα/βapo (blue), Ostα/βTLCA (violet) and Ostα/βDHEAS (green). The RMSD of Ostα/βTLCA and Ostα/βDHEAS compared with Ostα/βapo is 0.264 Å and 0.204 Å, correspondingly. b Lateral view from the extracellular side of substrate binding site. Surface is coloured based on Ostα/βapo structure hydrophobic properties. c The amphiphilic helix ICH is located between TM4 and TM5.
Extended Data Fig. 5 Sequence alignment of Ostα/β from different species.
Secondary structure assignments are based on the resolved Ostα (a) and Ostβ (b). Hom.s, Homo sapiens, Mus.m, Mus musculus, Mon.m, Monodon monoceros, Rou.a, Rousettus aegyptiacus, Dan.r, Danio rerio, Aha.p, Ahaetulla prasine, Gal.g, Gallus gallus, Dan.r, Danio rerio, Dro.m, Drosophila melanogaster, Cae.e, Caenorhabditis elegans.
Extended Data Fig. 6 Inner face key residues at the potential extracellular binding site.
a Hydrophilic surface of inner lateral lobe. b Hydrophilic surface of inner main lobe. Effects of mutants at the inner cavity on Ostα/β transport activity. c, d Ligands interaction with Ostα/β near the extracellular side. LigPlot+ analysis for cholesterol (blue, c) and DHEAS (green, d). Hydrogen bonds were defined using a donor–acceptor distance cutoff of 3.9 Å, and hydrophobic (non-bonded) contacts were identified within a 2.2–3.9 Å range. e MD simulation distance traces from Ostα/βDHEAS tetramer structure: Lig–G279 (DHEAS sulfur–G279 Cα), Lig–K191 (DHEAS sulfur–K191 side chain nitrogen), P209–E116 (minimum distance), and Lig–palm (minimum distance DHEAS sulfur-7 palmitoylated cysteines).
Extended Data Fig. 7 Lipids binding at the Ostα-Ostα’ interface.
a Head group of PE -like lipids located at the interface with positive potential. The surface electrostatic potentials were calculated by APBS. b Local densities of PE-like lipids in C2 Ostα/βapo map (contour level 0.2).
Extended Data Fig. 8 Cryo-EM structure of Ostα/βapo purified in the absence of CHS.
a, Cryo-EM map of Ostα/βapo purified without CHS, coloured by local resolution. The previously determined Ostα/βapo model (protein shown in green, four cholesterol molecules in yellow) was rigid-body fitted into the map for comparison (contour level 0.28). b, Fourier shell correlation (FSC) curve of the final reconstruction. c, Enlarged views of sterol-like densities observed at three sites, consistent with cholesterol binding. d, Angular distribution of particle orientations contributing to the reconstruction.
Extended Data Fig. 9 Representative lipid-like densities in the Ostα/βapo structure.
a Representative non-protein densities surrounding the Ostα/βapo structure (contour = 0.21). Putative cholesterols (excluding those at the substrate binding sites) shown in transparent gray, phospholipid-like densities in transparent purple. b Enlarged view of density resembling lysophosphatidylglycerol (LPG). c Enlarged view of density resembling PE. d Lipid-Ostα/β interaction interface 1 (IF1) with surrounding residues with extended hydrophobic side chains. e-g, Cholesterol-Ostα/β interaction interface 2-5 (IF2-5), with surrounding residues with extended hydrophobic side chains. h, [³H]-TCA uptake activity of IF mutants compared to WT Ostα/β. Data are mean ± s.e.m. (n = 6 biological replicates per group, ***P = 0.0001; ****P < 0.0001. i, FASC analysis of IFs mutants showing differential effects on Ostα cell surface expression. Data are mean ± s.e.m. (n = 3 biological replicates per group, ****P < 0.0001, ns = not significant P > 0.05, **P < 0.01. The exact P-values for the respective IF-mutants 1-5: 0.3014, 0.1436, 0.0067, 0.6246, >0.999). j, Quantification of plasma membrane (green) and total (gray) Ostα expression levels in WT and IF-mutants backgrounds, normalized to WT Ostα/β membrane expression. Data are mean ± s.e.m. (n = 3 biological replicates per group). Unpaired t-tests were used for statistical comparisons; “nd” indicates no significant difference. The exact P values were: >0.9999, 0.0194, 0.7840, 0.6885, 0.0233, and 0.4872. h, i, Data analyzed by one-way ANOVA with Fisher’s LSD test. h-j, Results are representative of three independent experiments.
Supplementary information
Supplementary Information (download PDF )
This file contains: Supplementary Information 1 (Figs 1–11 and Tables 1 and 2); Supplementary Information 2 (raw SDS–PAGE and western blot source data for Extended Data Fig. 1c) and Supplementary Information 3 (chemical structures of ligands in the figures).
Supplementary Video 1 (download MP4 )
DHEAS upward movement in the tetrameric Ostα/β complex, trajectory 2. Time-evolution video from MD simulations of the tetrameric Ostα/β complex (trajectory 2), illustrating the upward movement of DHEAS during the production phase.
Supplementary Video 2 (download MP4 )
DHEAS upward movement in the tetrameric Ostα/β complex, trajectory 5. Time-evolution video from MD simulations of the tetrameric Ostα/β complex (trajectory 5), showing the upward movement of DHEAS during the production phase.
Supplementary Video 3 (download MP4 )
DHEAS upward movement in the heterodimeric Ostα/β complex, trajectory 1. Time-evolution video from MD simulations of the heterodimeric Ostα/β complex (trajectory 1), depicting the upward movement of DHEAS during the production phase.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yang, X., Cui, N., Li, T. et al. Structures of Ostα/β reveal a unique fold and bile acid transport mechanism. Nature 651, 260–267 (2026). https://doi.org/10.1038/s41586-025-10029-7
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41586-025-10029-7