Abstract
Ancient DNA studies revealed that, in Europe from 6500 to 4000 bce, descendants of western Anatolian farmers mixed with local hunter-gatherers resulting in 70–100% ancestry turnover1, then steppe ancestry spread with the Corded Ware complex 3000–2500 bce2. Here we document an exception in the wetland, riverine and coastal areas of the Netherlands, Belgium and western Germany, using genome-wide data from 112 people 8500–1700 bce. A distinctive population with high (approximately 50%) hunter-gatherer ancestry persisted 3,000 years later than in most European regions, reflecting incorporation of female individuals of Early European Farmer ancestry into local communities. In the western Netherlands, the arrival of the Corded Ware complex was also exceptional: lowland individuals from settlements adopting Corded Ware pottery had hardly any steppe ancestry, despite a Y-chromosome characteristic of people associated with the early Corded Ware complex. These distinctive patterns may reflect the specific ecology that they inhabited, which was not amenable to full adoption of the early Neolithic type of farming introduced with Linearbandkeramik3, and resulted in distinct communities where transfer of ideas was accompanied by little gene flow. This changed with the formation of Lower Rhine–Meuse Bell Beaker users by fusion of local people (13–18%) and Corded Ware associated migrants of both sexes. Their subsequent expansion then had a disruptive impact across a much wider part of northwestern Europe, especially in Great Britain where they were the main source of a 90–100% replacement of local Neolithic ancestry.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Genotype data for individuals included in this study can be obtained from the Harvard Dataverse repository (https://reich.hms.harvard.edu/datasets). The DNA sequences reported in this paper have been deposited in the European Nucleotide Archive under accession number PRJEB105335. Other newly reported data, such as radiocarbon dates and archaeological context information, are included in the Article and its Supplementary Information.
Change history
18 February 2026
In the version of the article initially published, the text “Provinciaal Archeologisch Depot Flevoland (Tineke Roovers)” was missing from the Acknowledgements and has now been added to the HTML and PDF versions of the article.
References
Lipson, M. et al. Parallel palaeogenomic transects reveal complex genetic history of early European farmers. Nature 551, 368–372 (2017).
Allentoft, M. E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Amkreutz, L. W. S. W. Persistent traditions. A Long-Term Perspective on Communities in the Process of Neolithisation in the Lower Rhine Area (5500–2500 Cal BC) (Sidestone Press, 2013).
Lazaridis, I. et al. Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513, 409–413 (2014).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Olalde, I. et al. The Beaker phenomenon and the genomic transformation of northwest Europe. Nature 555, 190–196 (2018).
Papac, L. et al. Dynamic changes in genomic and social structures in third millennium BCE central Europe. Sci. Adv. 7, eabi6941 (2021).
Robb, J. Material culture, landscapes of action, and emergent causation: a new model for the origins of the European Neolithic. Curr. Anthropol. 54, 657–683 (2013).
Mittnik, A. et al. The genetic prehistory of the Baltic Sea region. Nat. Commun. 9, 442 (2018).
Skoglund, P. et al. Genomic diversity and admixture differs for Stone-Age Scandinavian foragers and farmers. Science 344, 747–750 (2014).
Hofmann, D. in Something Out of the Ordinary? Interpreting Diversity in the Early Neolithic Linearbandkeramik and Beyond (eds Amkreutz, L. W. S. W. et al.) 191–226 (Cambridge Scholars Press, 2016).
Kirschneck, E. The phenomena la Hoguette and Limburg—technological aspects. Open Archaeol. 7, 1295–1344 (2021).
Crombé, P. et al. New evidence on the earliest domesticated animals and possible small-scale husbandry in Atlantic NW Europe. Sci. Rep. 10, 20083 (2020).
Brusgaard, N. Ø et al. Early animal management in northern Europe: multi-proxy evidence from Swifterbant, the Netherlands. Antiquity 98, 654–671 (2024).
Kooijmans, L. P. L. & Jongste, P. F. B. A Neolithic Settlement on the Dutch North Sea Coast c. 3500 CAL BC (Analecta Praehistorica Leidensia, 2006).
Dreshaj, M., Dee, M., Brusgaard, N., Raemaekers, D. & Peeters, H. High-resolution Bayesian chronology of the earliest evidence of domesticated animals in the Dutch wetlands (Hardinxveld-Giessendam archaeological sites). PLoS ONE 18, e0280619 (2023).
Menne, J. & Brunner, M. Transition from Swifterbant to Funnelbeaker: a Bayesian chronological model. Open Archaeol. 7, 1235–1243 (2021).
Goossens, T. A. Opgraving Hellevoetsluis-Ossenhoek. Een Nederzetting van de Vlaardingen-Groep op een Kwelderrug in de Gemeente Hellevoetsluis (ARCHOL, 2009).
Bourgeois, Q. Monuments on the Horizon: The Formation of the Barrow Landscape throughout the 3rd and 2nd Millennium BC (Sidestone Press, 2013).
Beckerman, S. M. Corded Ware Coastal Communities: Using Ceramic Analysis to Reconstruct Third Millennium BC Societies in the Netherlands (Sidestone Press, 2015).
Fokkens, H., Steffens, B. J. W. & van As, S. F. M. Farmers, fishers, fowlers, hunters: knowledge generated by development-led archaeology about the Late Neolithic, the Early Bronze Age and the start of the Middle Bronze Age (2850–1500 cal BC) in the Netherlands. Ned. Archeol. Rapp. 53, 978–990 (2016).
Furholt, M. Re-integrating archaeology: a contribution to aDNA studies and the migration discourse on the 3rd millennium BC in Europe. Proc. Prehist. Soc. 85, 115–129 (2019).
Kroon, E. J. Serial Learners. Interactions between Funnel Beaker West and Corded Ware Communities in the Netherlands during the Third Millennium BCE from the Perspective of Ceramic Technology (Sidestone Press, 2024).
Wentink, K. Stereotype. The Role of Grave Sets in Corded Ware and Bell Beaker Funerary Practices (Sidestone Press, 2020).
Dyselinck, T. et al. Gent-Hogeweg, Vlakdekkende Opgraving. Mariakerke BAAC-Rapport A-11.0045 (BAAC, 2013).
Vander Linden, M. What linked the Bell Beakers in third millennium BC Europe? Antiquity 81, 343–352 (2007).
Lanting, J. N. De NO-Nederlandse/NW-Duitse Klokbekergroep: culturele achtergrond, typologie van het aardewerk, datering, verspreiding en grafritueel. Palaeohistoria 49–50, 11–326 (2008).
Posth, C. et al. Palaeogenomics of Upper Palaeolithic to Neolithic European hunter-gatherers. Nature 615, 117–126 (2023).
Immel, A. et al. Genome-wide study of a Neolithic Wartberg grave community reveals distinct HLA variation and hunter-gatherer ancestry. Commun. Biol. 4, 113 (2021).
Veselka, B. et al. Assembling ancestors: the manipulation of Neolithic and Gallo-Roman skeletal remains at Pommerœul, Belgium. Antiquity 98, 1576–1591 (2024).
Patterson, N. et al. Large-scale migration into Britain during the Middle to Late Bronze Age. Nature 601, 588–594 (2021).
ten Anscher, T. J., Knippenberg, S., van der Linde, C. M., Roessingh, W. & Willemse, N. W. Doorbraken aan de Rijn. Een Swifterbant-Gehucht, een Hazendonk-Nederzetting en Erven en Graven uit de Bronstijd in Medel-De Roeskamp RAAP-Rapport 6519, Archol Rapport 742, ADC Rapport A-16.0207 (BAAC, 2023).
Arzelier, A. et al. Neolithic genomic data from southern France showcase intensified interactions with hunter-gatherer communities. iScience 25, 105387 (2022).
Seguin-Orlando, A. et al. Heterogeneous hunter-gatherer and steppe-related ancestries in Late Neolithic and Bell Beaker genomes from present-day France. Curr. Biol. 31, 1072–1083 (2021).
Rivollat, M. et al. Ancient genome-wide DNA from France highlights the complexity of interactions between Mesolithic hunter-gatherers and Neolithic farmers. Sci. Adv. 6, aaz5344 (2020).
Mathieson, I. et al. The genomic history of southeastern Europe. Nature 555, 197–203 (2018).
Olalde, I. et al. Derived immune and ancestral pigmentation alleles in a 7,000-year-old Mesolithic European. Nature 507, 225–228 (2014).
Gelabert, P. et al. Social and genetic diversity in first farmers of central Europe. Nat. Hum. Behav. https://doi.org/10.1038/s41562-024-02034-z (2024).
Ringbauer, H. et al. Accurate detection of identity-by-descent segments in human ancient DNA. Nat. Genet. 56, 143–151 (2023).
Bourgeois, Q. P. J. et al. Spatiotemporal reconstruction of Corded Ware and Bell Beaker burial rituals reveals complex dynamics divergent from steppe ancestry. Sci. Adv. 11, eadx22622262 (2025).
Lazaridis, I. et al. The genetic origin of the Indo-Europeans. Nature https://doi.org/10.1038/s41586-024-08531-5 (2025).
Linderholm, A. et al. Corded Ware cultural complexity uncovered using genomic and isotopic analysis from south-eastern Poland. Sci. Rep. 10, 6885 (2020).
Allentoft, M. E. et al. Population genomics of post-glacial western Eurasia. Nature 625, 301–311 (2024).
Brunel, S. et al. Ancient genomes from present-day France unveil 7, 000 years of its demographic history. Proc. Natl Acad. Sci USA https://doi.org/10.1073/pnas.1918034117 (2020).
Parasayan, O. et al. Late Neolithic collective burial reveals admixture dynamics during the third millennium BCE and the shaping of the European genome. Sci. Adv. 10, eadl2468 (2024).
Heyd, V. in Yamnaya Interactions. Proceedings of the International Workshop held in Helsinki, 25–26 April 2019. The Yamnaya Impact on Prehistoric Europe (eds Heyd, V. et al.) 383–414 (Archaeolingua Alapítvány, (2021).
Jeunesse, C. The dogma of the Iberian origin of the Bell Beaker: attempting its deconstruction. J. Neolit. Archaeol. 16, 158–166 (2015).
Price, T. D., Knipper, C., Grupe, G. & Smrcka, V. Strontium isotopes and prehistoric human migration: The Bell Beaker period in Central Europe. Eur. J. Archaeol. 7, 9–40 (2004).
Parker Pearson, M. et al. The Beaker People: Isotopes, Mobility and Diet in Prehistoric Britain Vol. 7 (Oxbow Books, 2019).
Zvelebil, M. in Archaeogenetics: DNA and the Population Prehistory of Europe (eds Renfrew, C. & Boyle, K.) 57–79 (McDonald Institute Monographs, 2000).
Armit, I. & Reich, D. The return of the Beaker folk? Rethinking migration and population change in British prehistory. Antiquity 95, 1464–1477 (2021).
Cleal, R. M. J. & Pollard, J. in Is There a British Chalcolithic? People, Place and Polity in the Later 3rd Millennium (eds Allen, M. J. et al.) Vol. 4, 317–332 (Oxbow Books, 2012).
Booth, T. J., Brück, J., Brace, S. & Barnes, I. Tales from the supplementary information: ancestry change in Chalcolithic–Early Bronze Age Britain was gradual with varied kinship organization. Camb. Archaeol. J. https://doi.org/10.1017/s0959774321000019 (2021).
Gibson, A. Beakers in Britain. The Beaker package reviewed. Préhist. Méd. 8, 32 (2020).
Adler, C. J., Haak, W., Donlon, D. & Cooper, A. Survival and recovery of DNA from ancient teeth and bones. J. Archaeol. Sci. 38, 956–964 (2011).
Pinhasi, R., Fernandes, D. M., Sirak, K. & Cheronet, O. Isolating the human cochlea to generate bone powder for ancient DNA analysis. Nat. Protoc. 14, 1194–1205 (2019).
Rohland, N., Glocke, I., Aximu-Petri, A. & Meyer, M. Extraction of highly degraded DNA from ancient bones, teeth and sediments for high-throughput sequencing. Nat. Protoc. 13, 2447–2461 (2018).
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl Acad. Sci. USA 110, 15758–15763 (2013).
Korlević, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. BioTechniques 59, 87–93 (2015).
Dulias, K. et al. Ancient DNA at the edge of the world: continental immigration and the persistence of Neolithic male lineages in Bronze Age Orkney. Proc. Natl Acad. Sci. USA 119, e2108001119 (2022).
Briggs, A. W. et al. Removal of deaminated cytosines and detection of in vivo methylation in ancient DNA. Nucleic Acids Res. 38, e87 (2009).
Rohland, N., Harney, E., Mallick, S., Nordenfelt, S. & Reich, D. Partial uracil–DNA–glycosylase treatment for screening of ancient DNA. Philos. Trans. R. Soc. Lond. B 370, 20130624 (2015).
Gansauge, M. T., Aximu-Petri, A., Nagel, S. & Meyer, M. Manual and automated preparation of single-stranded DNA libraries for the sequencing of DNA from ancient biological remains and other sources of highly degraded DNA. Nat. Protoc. 15, 2279–2300 (2020).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Rohland, N. et al. Three assays for in-solution enrichment of ancient human DNA at more than a million SNPs. Genome Res. 32, 2068–2078 (2022).
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res. 40, e3 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Fu, Q. et al. A revised timescale for human evolution based on ancient mitochondrial genomes. Curr. Biol. 23, 553–559 (2013).
Korneliussen, T. S., Albrechtsen, A. & Nielsen, R. ANGSD: analysis of next generation sequencing data. BMC Bioinform. 15, 356 (2014).
Patterson, N. et al. Ancient admixture in human history. Genetics 192, 1065–1093 (2012).
Biagini, S. A. et al. People from Ibiza: an unexpected isolate in the Western Mediterranean. Eur. J. Hum. Genetics 27, 941–951 (2019).
Schönherr, S., Weissensteiner, H., Kronenberg, F. & Forer, L. Haplogrep 3—an interactive haplogroup classification and analysis platform. Nucleic Acids Res. 51, W263–W268 (2023).
Lazaridis, I. et al. The genetic history of the Southern Arc: a bridge between West Asia and Europe. Science 377, eabdm4247 (2022).
Fowler, C. et al. A high-resolution picture of kinship practices in an Early Neolithic tomb. Nature 601, 584–587 (2021).
Rubinacci, S., Ribeiro, D. M., Hofmeister, R. J. & Delaneau, O. Efficient phasing and imputation of low-coverage sequencing data using large reference panels. Nat. Genet. 53, 120–126 (2021).
The 1000 Genomes Project Consortium. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65 (2012).
Rivollat, M. et al. Extensive pedigrees reveal the social organization of a Neolithic community. Nature https://doi.org/10.1038/s41586-023-06350- (2023).
Ringbauer, H., Novembre, J. & Steinrücken, M. Parental relatedness through time revealed by runs of homozygosity in ancient DNA. Nat. Commun. 12, 5425 (2021).
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006).
Fournier, R., Fulton, A. P. & Reich, D. A SNP panel for co-analysis of capture and shotgun ancient DNA data. Preprint at bioRxiv https://doi.org/10.1101/2025.07.30.667733 (2025).
Narasimhan, V. M. et al. The formation of human populations in South and Central Asia. Science 365, eaat7487 (2019).
Fenner, J. N. Cross-cultural estimation of the human generation interval for use in genetics-based population divergence studies. Am. J. Phys. Anthropol. 128, 415–423 (2005).
Reimer, P. J. et al. The IntCal20 Northern Hemisphere radiocarbon age calibration curve (0–55 cal kBP). Radiocarbon 62, 725–757 (2020).
Philippsen, B. The freshwater reservoir effect in radiocarbon dating. Herit. Sci. 1, 24 (2013).
van der Plicht, J. & Streurman, H. J. A new model for radiocarbon dating of marine shells from the Netherlands. Radiocarbon 67, 378–411 (2025).
Kamjan, S., Gillis, R. E., Cakirlar, C. & Raemaekers, D. C. M. Specialized cattle farming in the Neolithic Rhine-Meuse Delta: results from zooarchaeological and stable isotope (δ18O, δ13C, δ15N) analyses. PLoS ONE 15, e0240464 (2020).
Massicotte, P. & South, A. rnaturalearth: World Map Data from Natural Earth (2024); docs.ropensci.org/rnaturalearth.
Acknowledgements
We thank N. Adamski, N. Broomandkhoshbacht, E. Curtis, I. Greenslade, K. Stewardson and F. Zalzala for laboratory work; the staff at the Provinciaal Depot voor Archeologie van Noord-Holland (M. Veen and R. van Eerden), Provinciaal Archeologisch depot Zuid-Holland (I. Riemersma and M. Phlippeau), Archeologisch Depot Gelderland (S. Weiss-König), Provinciaal Archeologisch Depot Flevoland (Tineke Roovers), and National Museum of Antiquities Leiden (L. Amkreutz) for granting permission to sample ancient remains and assistance in sampling; T. Grange and E.-M. Geigl for facilitating access to Bréviandes genomic data. The aDNA data generation and analysis was supported by the National Institutes of Health (R01-HG012287); the John Templeton Foundation (grant 61220); a private gift from J.-F. Clin; the Allen Discovery Center program, a Paul G. Allen Frontiers Group advised program of the Paul G. Allen Family Foundation; the Howard Hughes Medical Institute (D.R.); grant RYC2019-027909-I and project PID2022-140886NA-I00 funded by MCIN/AEI/10.13039/501100011033, “ESF Investing in your future” and FEDER, UE (I.O.); the Basque Government under “Grupos Consolidados” grant no. IT1633-22 (I.O.); NWO-VIDI grant “The Talking Dead” VI.VIDI.191.149 (Q.B.); the Max Planck Society, by the French Research Foundation and German Research Foundation (to M.R., W.H. and M.-F.D., project INTERACT, ANR-17-FRAL-0010 and DFG-HA-5407/4-1, 2018-21); and the Leverhulme Trust (A.F., F.G., C.J.E., M.B.R., M.P.). The accepted version of this article (before the editing, proofreading and formatting changes following the paper being accepted) is subject to the Howard Hughes Medical Institute (HHMI) Open Access to Publications policy; HHMI laboratory heads have previously granted a non-exclusive CC BY 4.0 licence to the public and a sublicensable licence to HHMI in their research articles. Pursuant to those licences, the accepted manuscript can be made freely available under a CC BY 4.0 licence immediately after publication.
Author information
Authors and Affiliations
Contributions
I.O., E.A., Q.B., H.F. and D.R. wrote the manuscript with the input of all of the other authors. M.B.R., R.P., W.H., M.P. and D.R. supervised parts of the study. I.O., I.L., N.P. and M.R. analysed genetic data. E.A., Q.B. and H.F. edited archaeological information. E.A, L.A., S.B., M.-F.D., A.F., D.F., F.G., J.F.K., L.M.K., C.v.d.L., J.v.d.L., K.L., L.L.K., R.L., R.M., H.M., P.N., D.C.M.R., M.R., L.S., J.R.S., T.t.A., M.T. and C.J.E. sampled anthropological remains and/or contributed to the creation of the archaeological supplement. G.S., M.M., A.M. and S.M. processed bioinformatic data. K.C., O.C., T.F., L.I., J.O., I.P., L.Q., J.N.W., C.J.E. and N.R. carried out wet laboratory work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Catherine Frieman, Eva-Maria Geigl, Choongwon Jeong, Oguzhan Parasayan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 2 Hunter-gatherer ancestry proportions across time in the Lower Rhine-Meuse area.
Ancestry proportions were estimated using qpAdm (Supplementary Table 6).
Extended Data Fig. 3 Genetic connections of Lower Rhine-Meuse CW/Vlaardingen individuals (n = 3).
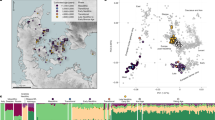
a, IBD sharing of Lower Rhine-Meuse CW/Vlaardingen individuals. Sites are represented by circles with size proportional to the number of individuals amenable to IBD calling. Grey circles indicate archaeological sites between 3000-2000 bce with no IBD connections to CW/Vlaardingen individuals. The map was drawn using public-domain Natural Earth data with the rnaturalearth package in R87. b, Decay of IBD sharing with geographic distance for Lower Rhine-Meuse CW/Vlaardingen individuals. Pairs were considered to share IBD if they share at least one segment of >12 cM. Dotted lines represent IBD connections involving at least one individual without steppe-related ancestry.
Extended Data Fig. 4 Admixture time estimates using DATES81.
Boxes represent the chronological range for each population. Confidence intervals represent admixture date ranges, using 28 years per generation and the average date of the chronological range. We tested Balkan_N + WHG admixture for groups in blue, and MN_Wartberg+Germany_CordedWare admixture for groups in orange. In bold, groups from the Lower Rhine-Meuse region.
Supplementary information
Supplementary Information (download DOCX )
Archaeological background, archaeological site information and methods descriptions, including Supplementary Information 1–4, Supplementary Figs. 1–7, additional Supplementary Tables and Supplementary References.
Supplementary Tables (download XLSX )
Supplementary Tables 1–16.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Olalde, I., Altena, E., Bourgeois, Q. et al. Lasting Lower Rhine–Meuse forager ancestry shaped Bell Beaker expansion. Nature (2026). https://doi.org/10.1038/s41586-026-10111-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41586-026-10111-8