Abstract
Designing lipid nanoparticle (LNP) delivery systems with specific targeting, potency and minimal side effects is crucial for their clinical use. However, traditional characterization methods, such as dynamic light scattering, cannot accurately quantify physicochemical properties of LNPs and how these are influenced by the lipid composition and mixing method. Here, we structurally characterize polydisperse LNP formulations by applying emerging solution-based biophysical methods that have higher resolution and provide biophysical data beyond size and polydispersity. These techniques include sedimentation velocity analytical ultracentrifugation, field-flow fractionation followed by multiangle light scattering and size-exclusion chromatography in line with synchrotron small-angle X-ray scattering. We show that LNPs have intrinsic polydispersity in size, RNA loading and shape, which depend on both the formulation technique and the lipid composition. Lastly, we predict LNP transfection in vitro and in vivo by examining the relationship between mRNA translation and physicochemical characteristics. Solution-based biophysical methods will be essential for determining LNP structure–function relationships, facilitating the creation of new design rules for LNPs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
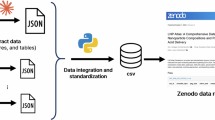
All data supporting the findings of this study are available within the paper and Supplementary Information. The SV-AUC, FFF–MALS and SEC–SAXS raw and analyzed data are available from Zenodo (https://doi.org/10.5281/zenodo.17042311)60. Source data are provided with this paper.
Code availability
No original code was generated for this study.
Change history
22 January 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41587-026-03014-6
References
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines—a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Hajj, K. A. et al. A potent branched-tail lipid nanoparticle enables multiplexed mRNA delivery and gene editing in vivo. Nano Lett. 20, 5167–5175 (2020).
Finn, J. D. et al. A single administration of CRISPR/Cas9 lipid nanoparticles achieves robust and persistent in vivo genome editing. Cell Rep/ 22, 2455–2468 (2018).
Walsh, G. & Walsh, E. Biopharmaceutical benchmarks 2022. Nat. Biotechnol. 40, 1722–1760 (2022).
Hou, X., Zaks, T., Langer, R. & Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 6, 1078–1094 (2021).
Hammel, M. et al. Correlating the structure and gene silencing activity of oligonucleotide-loaded lipid nanoparticles using small-angle X-ray scattering. ACS Nano 17, 11454–11465 (2023).
Chen, D. et al. Rapid discovery of potent siRNA-containing lipid nanoparticles enabled by controlled microfluidic formulation. J. Am. Chem. Soc. 134, 6948–6951 (2012).
Leung, A. K. K. et al. Lipid nanoparticles containing siRNA synthesized by microfluidic mixing exhibit an electron-dense nanostructured core. J. Phys. Chem. C 116, 18440–18450 (2012).
Shepherd, S. J. et al. Scalable mRNA and siRNA lipid nanoparticle production using a parallelized microfluidic device. Nano Lett. 21, 5671–5680 (2021).
Hallan, S. S., Sguizzato, M., Esposito, E. & Cortesi, R. Challenges in the physical characterization of lipid nanoparticles. Pharmaceutics 13, 1–31 (2021).
Henrickson, A. et al. Density matching multi-wavelength analytical ultracentrifugation to measure drug loading of lipid nanoparticle formulations. ACS Nano 15, 5068–5076 (2021).
Sych, T. et al. High-throughput measurement of the content and properties of nano-sized bioparticles with single-particle profiler. Nat. Biotechnol. 42, 587–590 (2024).
Li, S. et al. Payload distribution and capacity of mRNA lipid nanoparticles. Nat. Commun. 13, 5561 (2022).
Li, S. et al. Single-particle spectroscopic chromatography reveals heterogeneous RNA loading and size correlations in lipid nanoparticles. ACS Nano 18, 15729–15743 (2024).
Dao, H. M. et al. Characterization of mRNA lipid nanoparticles by electron density mapping reconstruction: X-ray scattering with density from solution scattering (DENSS) algorithm. Pharm. Res. 41, 501–512 (2024).
Akinc, A. et al. The Onpattro story and the clinical translation of nanomedicines containing nucleic acid-based drugs. Nat. Nanotechnol. 14, 1084–1087 (2019).
Hogan, M. J. & Pardi, N. mRNA vaccines in the COVID-19 pandemic and beyond. Annu. Rev. Med. 73, 17–39 (2022).
Love, K. T. et al. Lipid-like materials for low-dose, in vivo gene silencing. Proc. Natl Acad. Sci. USA 107, 1864–1869 (2010).
Guerrini, G., Mehn, D., Scaccabarozzi, D., Gioria, S. & Calzolai, L. Analytical ultracentrifugation to assess the quality of LNP-mRNA therapeutics. Int. J. Mol. Sci. 25, 5718 (2024).
Thaller, A. et al. SV-AUC as a stability-indicating method for the characterization of mRNA-LNPs. Eur. J. Pharm. Biopharm. 182, 152–156 (2023).
Schuck, P. & Rossmanith, P. Determination of the sedimentation coefficient distribution by least-squares boundary modeling. Biopolymers 54, 328–341 (2000).
Zhao, H., Sousa, A. A. & Schuck, P. Flotation coefficient distributions of lipid nanoparticles by sedimentation velocity analytical ultracentrifugation. ACS Nano 18, 18663–18672 (2024).
Jia, X. et al. Enabling online determination of the size-dependent RNA content of lipid nanoparticle-based RNA formulations. J. Chromatogr. B 1186, 123015 (2021).
Some, D., Amartely, H., Tsadok, A. & Lebendiker, M. Characterization of proteins by size-exclusion chromatography coupled to multi-angle light scattering. J. Vis. Exp. 148, e59615 (2019).
Gao, Z. et al. Development of an advanced separation and characterization platform for mRNA and lipid nanoparticles using multi-detector asymmetrical flow field-flow fractionation. Anal. Bioanal. Chem. 416, 5281–5293 (2024).
Graewert, M. A. et al. Quantitative size-resolved characterization of mRNA nanoparticles by in-line coupling of asymmetrical-flow field-flow fractionation with small angle X-ray scattering. Sci. Rep. 13, 15764 (2023).
Kulkarni, C. V., Yaghmur, A., Steinhart, M., Kriechbaum, M. & Rappolt, M. Effects of high pressure on internally self-assembled lipid nanoparticles: a synchrotron small-angle X-ray scattering (SAXS) study. Langmuir 32, 11907–11917 (2016).
Uebbing, L. et al. Investigation of pH-responsiveness inside lipid nanoparticles for parenteral mRNA application using small-angle X-ray scattering. Langmuir 36, 13331–13341 (2020).
Meisburger, S. P., Xu, D. & Ando, N. REGALS: a general method to deconvolve X-ray scattering data from evolving mixtures. IUCrJ 8, 225–237 (2021).
Hura, G. L. et al. Comprehensive macromolecular conformations mapped by quantitative SAXS analyses. Nat. Methods 10, 453–454 (2013).
Guimaraes, P. P. G. et al. Ionizable lipid nanoparticles encapsulating barcoded mRNA for accelerated in vivo delivery screening. J. Control. Release 316, 404–417 (2019).
Rhym, L. H., Manan, R. S., Koller, A., Stephanie, G. & Anderson, D. G. Peptide-encoding mRNA barcodes for the high-throughput in vivo screening of libraries of lipid nanoparticles for mRNA delivery. Nat. Biomed. Eng. 7, 901–910 (2023).
DeBerardinis, R. J. & Chandel, N. S. Fundamentals of cancer metabolism. Sci. Adv. 2, e1600200 (2016).
Banushi, B., Joseph, S. R., Lum, B., Lee, J. J. & Simpson, F. Endocytosis in cancer and cancer therapy. Nat. Rev. Cancer 23, 450–473 (2023).
Thanh Cong, V. et al. How can we use the endocytosis pathways to design nanoparticle drug-delivery vehicles to target cancer cells over healthy cells? Chem. Soc. Rev. 51, 7531–7559 (2022).
Sedic, M. et al. Safety evaluation of lipid nanoparticle–formulated modified mRNA in the Sprague–Dawley rat and cynomolgus monkey. Vet. Pathol. 55, 341–354 (2018).
Whitehead, K. A. et al. Degradable lipid nanoparticles with predictable in vivo siRNA delivery activity. Nat. Commun. 5, 4277 (2014).
Mrksich, K. et al. Influence of ionizable lipid tail length on lipid nanoparticle delivery of mRNA of varying length. J. Biomed. Mater. Res. A 112, 1494–1505 (2024).
Mrksich, K., Padilla, M. S. & Mitchell, M. J. Breaking the final barrier: evolution of cationic and ionizable lipid structure in lipid nanoparticles to escape the endosome. Adv. Drug Deliv. Rev. 214, 115446 (2024).
Lindsay, S., Hussain, M., Binici, B. & Perrie, Y. Exploring the challenges of lipid nanoparticle development: the in vitro–in vivo correlation gap. Vaccines 13, 339 (2025).
Shepherd, S. J. et al. Throughput-scalable manufacturing of SARS-CoV-2 mRNA lipid nanoparticle vaccines. Proc. Natl Acad. Sci. USA 120, e2303567120 (2023).
Witten, J. et al. Artificial intelligence-guided design of lipid nanoparticles for pulmonary gene therapy. Nat. Biotechnol. https://doi.org/10.1038/s41587-024-02490-y (2024).
Madathiparambil Visalakshan, R. et al. The influence of nanoparticle shape on protein corona formation. Small 16, 2000285 (2020).
Li, Z. et al. Shape effect of glyco-nanoparticles on macrophage cellular uptake and immune response. ACS Macro Lett. 5, 1059–1064 (2016).
Vácha, R., Martinez-Veracoechea, F. J. & Frenkel, D. Receptor-mediated endocytosis of nanoparticles of various shapes. Nano Lett. 11, 5391–5395 (2011).
Han, X. et al. Adjuvant lipidoid-substituted lipid nanoparticles augment the immunogenicity of SARS-CoV-2 mRNA vaccines. Nat. Nanotechnol. 18, 1105–1114 (2023).
Zhang, Y. et al. Not all cells are created equal—endosomal escape in fluorescent nanodiamonds in different cells. Nanoscale 13, 13294–13300 (2021).
Chatterjee, S., Kon, E., Sharma, P. & Peer, D. Endosomal escape: a bottleneck for LNP-mediated therapeutics. Proc. Natl Acad. Sci. USA 121, e2307800120 (2024).
Karikó, K., Muramatsu, H., Ludwig, J. & Weissman, D. Generating the optimal mRNA for therapy: HPLC purification eliminates immune activation and improves translation of nucleoside-modified, protein-encoding mRNA. Nucleic Acids Res. 39, e142 (2011).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and Lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
DiFabio, J. et al. The Life Science X-ray Scattering beamline at NSLS-II. In Proc. 12th International Conference on Synchrotron Radiation Instrumentation Vol. 1741 (eds Chubar, O. et al.) 030049 (AIP, 2016).
Yang, L. et al. Tools for supporting solution scattering during the COVID-19 pandemic. J. Synchrotron Rad. 28, 1237–1244 (2021).
Yang, L. et al. Solution scattering at the Life Science X-ray Scattering (LiX) beamline. J. Synchrotron Rad. 27, 804–812 (2020).
Hopkins, J. B., Gillilan, R. E. & Skou, S. BioXTAS RAW: improvements to a free open-source program for small-angle X-ray scattering data reduction and analysis. J. Appl. Crystallogr. 50, 1545 (2017).
Maeder, M. Evolving factor analysis for the resolution of overlapping chromatographic peaks. Anal. Chem. 59, 527–530 (1987).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Cryst. 25, 495–503 (1992).
Semenyuk, A. V. & Svergun, D. I. GNOM—a program package for small-angle scattering data processing. J. Appl. Cryst. 24, 537–540 (1991).
Johansen, D., Trewhella, J. & Goldenberg, D. P. Fractal dimension of an intrinsically disordered protein: small-angle X-ray scattering and computational study of the bacteriophage λ N protein. Protein Sci. 20, 1955–1970 (2011).
Grant, T. D. Ab initio electron density determination directly from solution scattering data. Nat. Methods 15, 191–193 (2018).
Padilla, M. & Gupta, K. SV-AUC, FFF–MALS, SEC–SAXS datasets for ‘Elucidating lipid nanoparticle properties and structure through biophysical analyses’. Zenodo https://doi.org/10.5281/zenodo.17042311 (2025).
Acknowledgements
The SV-AUC experiments were performed at the Johnson Foundation Biophysical and Structural Biology Core Facility (University of Pennsylvania). The LiX beamline is part of the Center for Biomolecular Structure (CBMS), which is primarily supported by the Department of Energy Office of Biological and Environmental Research (KP1605010). As part of NSLS-II, a national user facility at Brookhaven National Laboratory, work performed at the CBMS is supported in part by the US Department of Energy, Office of Science, Office of Basic Energy Sciences Program under contract number DE-SC0012704. Additionally, we thank E. Cento, Z. Chen, M. A. Eldabbas and E. Maddox of the Human Immunology Core and the Division of Transfusion Medicine and Therapeutic Pathology at the Perelman School of Medicine (University of Pennsylvania) for providing deidentified CD4+ and CD8+ T cells that were purified from healthy donor apheresis using StemCell RosetteSep kits. Cryo-EM imaging was provided by the Beckman Center for Cryo-EM at the University of Pennsylvania Perelman School of Medicine. M.J.M. acknowledges support from a Burroughs Wellcome Fund Career Award at the Scientific Interface, an American Cancer Society Research Scholar Grant (RSG-22-122-01-ET) and a US National Science Foundation CAREER Award (CBET-2145491).
Author information
Authors and Affiliations
Contributions
M.S.P., S.J.S., M.K., X.Z., M.C., J.B., K.G. and M.J.M. conceptualized and designed the experiments. M.S.P., S.J.S., A.H., M.K., X.Z., J.B., H.M.Y., A.R., R.A.J. and K.M. performed the experiments. M.S.P., S.J.S., M.K., X.Z., M.C., J.B. and K.G. analyzed the data. M.S.P., D.I., K.G. and M.J.M. wrote and edited the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Yizhou Dong, Guangjun Nie and Wei Tao for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–14, Tables 1–6 and References.
Supplementary Data 1 (download XLSX )
Source data for supplementary figures.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Padilla, M.S., Shepherd, S.J., Hanna, A.R. et al. Elucidating lipid nanoparticle properties and structure through biophysical analyses. Nat Biotechnol (2025). https://doi.org/10.1038/s41587-025-02855-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-025-02855-x
This article is cited by
-
Lipid Nanoparticle Development in Practice: Challenges and Collective Insights
Pharmaceutical Research (2026)