Abstract
Although virus ecogenomics has expanded access to and understanding of the virosphere, existing classification tools lack taxonomic resolution and are unable to scale to modern discovery-based datasets or classify previously unknown sequence space. Here we develop vConTACT3—a machine learning-based tool that improves scalability and accuracy of virus taxonomy. By optimizing gene-sharing thresholds and leveraging adaptive, realm-specific cut-offs, vConTACT3 expands classification to both eukaryote and prokaryote viruses for four of the six officially recognized realms, and establishes accurate hierarchical taxonomy from genus to order. Specifically, vConTACT3 achieves >95% agreement with official taxonomy for 35,545 and 13,524 public prokaryotic and eukaryotic virus genomes, respectively, to surpass vConTACT2 across most realms, while still uniquely classifying previously uncharacterized taxa, and doing so even faster. vConTACT3 application provides taxonomy assignments for tens of thousands of unclassified taxa rapidly, automatically and systematically; evaluates virus sequence space to reveal support for fewer taxonomic ranks than currently available and identifies taxonomically challenging areas across the virosphere.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Data used for benchmarks (parameter optimizations), construction of databases and fine-tuning the pipeline are available from NCBI Virus RefSeq (v.218). Data used to test scalability, assess fragmentation and evaluate labeling stability are available from IMG/VR v.4.1 (December 2022 release). Databases used by vConTACT3 (as well as source files) are available via Zenodo at https://doi.org/10.5281/zenodo.10035619 and https://doi.org/10.5281/zenodo.10935513 (refs. 67,68).
Code availability
vConTACT3 is available via Bitbucket at https://bitbucket.org/MAVERICLab/vcontact3 (ref. 69) as an installable Python package, as well as through Python package managers Anaconda (https://anaconda.org/) and Mamba (https://mamba.readthedocs.io/). Instructions for building an Apptainer container of vConTACT3 is available on Bitbucket, along with a definitions file. A comprehensive documentation site is available through https://vcontact3.readthedocs.io. Optimization benchmarks were performed using v.3.0.0b36 (‘beta’ v.36), fragmentation analyses using v.3.0.0b63 and label stability, v3.1.4. All other results (including tool comparisons) should be assumed as v.3.0.0.b36. Through all analyses, Python 3.10 was used. Data processing and analyses were conducted using numpy v.1.23.5, pandas v.2.1.1 and scipy v.1.10.1. Taxonomic parsing was handled by the ETE3 Toolkit, v.3.1.3. Statistical analyses were performed using scikit-learn v.1.2.2 and scikit-bio v.0.5.8. Data visualizations were created using matplotlib v.3.7.1, seaborn v.0.12.1 and UpSetPlot v.0.7.0. Networks were rendered through a combination of Cytoscape v.3.10.1, networkx v.3.1 and Python-igraph v.0.10.4. Gene predictions were done through pyprodigal v.2.3.0 and pyprodigal-gv v.0.3.1 and all sequence processing through biopython v.1.81. Protein clustering was done using MMSeqs2 v.14-7e284.
References
Roux, S. et al. Ecogenomics and potential biogeochemical impacts of globally abundant ocean viruses. Nature 537, 689–693 (2016).
Guidi, L. et al. Plankton networks driving carbon export in the oligotrophic ocean. Nature 532, 465–470 (2016).
Zimmerman, A. E. et al. Metabolic and biogeochemical consequences of viral infection in aquatic ecosystems. Nat. Rev. Microbiol. https://doi.org/10.1038/s41579-019-0270-x (2020).
Emerson, J. B. et al. Host-linked soil viral ecology along a permafrost thaw gradient. Nat. Microbiol. 3, 870–880 (2018).
Jansson, J. K. & Wu, R. Soil viral diversity, ecology and climate change. Nat. Rev. Microbiol. 21, 296–311 (2023).
Koskella, B. & Taylor, T. B. Multifaceted impacts of bacteriophages in the plant microbiome. Annu. Rev. Phytopathol. 56, 361–380 (2018).
Yan, M. et al. Interrogating the viral dark matter of the rumen ecosystem with a global virome database. Nat. Commun. 14, 5254 (2023).
Yan, M. & Yu, Z. Viruses contribute to microbial diversification in the rumen ecosystem and are associated with certain animal production traits. Microbiome 12, 82 (2024).
Shkoporov, A. N. & Hill, C. Bacteriophages of the human gut: the “known unknown” of the microbiome. Cell Host Microbe 25, 195–209 (2019).
Shkoporov, A. N., Turkington, C. J. & Hill, C. Mutualistic interplay between bacteriophages and bacteria in the human gut. Nat. Rev. Microbiol. 20, 737–749 (2022).
Walker, P. J. et al. Changes to virus taxonomy and the Statutes ratified by the International Committee on Taxonomy of Viruses (2020). Arch. Virol. 165, 2737–2748 (2020).
Walker, P. J. et al. Recent changes to virus taxonomy ratified by the International Committee on Taxonomy of Viruses (2022). Arch. Virol. 167, 2429–2440 (2022).
Zerbini, F. M. et al. Changes to virus taxonomy and the ICTV Statutes ratified by the International Committee on Taxonomy of Viruses (2023). Arch. Virol. 168, 175 (2023).
Gorbalenya, A. E. et al. The new scope of virus taxonomy: partitioning the virosphere into 15 hierarchical ranks. Nat. Microbiol 5, 668–674 (2020).
Camargo, A. P. et al. IMG/VR v4: an expanded database of uncultivated virus genomes within a framework of extensive functional, taxonomic, and ecological metadata. Nucleic Acids Res. 51, D733–D743 (2023).
Roux, S. et al. Minimum Information about an Uncultivated Virus Genome (MIUViG): a community consensus on standards and best practices for describing genome sequences from uncultivated viruses. Nat. Biotechnol. 37, 29–37 (2018).
Simmonds, P. et al. Consensus statement: virus taxonomy in the age of metagenomics. Nat. Rev. Microbiol. 15, 161–168 (2017).
Simmonds, P. et al. Four principles to establish a universal virus taxonomy. PLoS Biol. 21, e3001922 (2023).
Dutilh, B. E. et al. Perspective on taxonomic classification of uncultivated viruses. Curr. Opin. Virol. 51, 207–215 (2021).
Koonin, E. V., Senkevich, T. G. & Dolja, V. V. The ancient Virus World and evolution of cells. Biol. Direct 1, 29 (2006).
Holmes, E. C. What does virus evolution tell us about virus origins? J. Virol. 85, 5247–5251 (2011).
Koonin, E. V. & Dolja, V. V. Virus World as an evolutionary network of viruses and capsidless selfish elements. Microbiol. Mol. Biol. Rev. 78, 278–303 (2014).
Moraru, C. VirClust—a tool for hierarchical clustering, core protein detection and annotation of (prokaryotic) viruses. Viruses 15, 1007 (2023).
Aiewsakun, P. & Simmonds, P. The genomic underpinnings of eukaryotic virus taxonomy: creating a sequence-based framework for family-level virus classification. Microbiome 6, 38 (2018).
Pons, J. C. et al. VPF-Class: taxonomic assignment and host prediction of uncultivated viruses based on viral protein families. Bioinformatics https://doi.org/10.1093/bioinformatics/btab026 (2021).
Camargo, A. P. et al. Identification of mobile genetic elements with geNomad. Nat. Biotechnol. 42, 1303–1312 (2024).
Moraru, C., Varsani, A. & Kropinski, A. M. VIRIDIC—a novel tool to calculate the intergenomic similarities of prokaryote-infecting viruses. Viruses 12, 1268 (2020).
Bao, Y., Chetvernin, V. & Tatusova, T. Improvements to pairwise sequence comparison (PASC): a genome-based web tool for virus classification. Arch. Virol. 159, 3293–3304 (2014).
Tisza, M. J., Belford, A. K., Domínguez-Huerta, G., Bolduc, B. & Buck, C. B. Cenote-Taker 2 democratizes virus discovery and sequence annotation. Virus Evol. 7, veaa100 (2021).
Lima-Mendez, G., Van Helden, J., Toussaint, A. & Leplae, R. Reticulate representation of evolutionary and functional relationships between phage genomes. Mol. Biol. Evol. 25, 762–777 (2008).
Bolduc, B. et al. vConTACT: an iVirus tool to classify double-stranded DNA viruses that infect Archaea and Bacteria. PeerJ 5, e3243 (2017).
Bin Jang, H. et al. Taxonomic assignment of uncultivated prokaryotic virus genomes is enabled by gene-sharing networks. Nat. Biotechnol. 37, 632–639 (2019).
Barylski, J. et al. Analysis of Spounaviruses as a case study for the overdue reclassification of tailed phages. Syst. Biol. 69, 110–123 (2020).
Turner, D. et al. Abolishment of morphology-based taxa and change to binomial species names: 2022 taxonomy update of the ICTV bacterial viruses subcommittee. Arch. Virol. 168, 74 (2023).
Van Dongen, S. Graph clustering via a discrete uncoupling process. SIAM J. Matrix Anal. Appl. 30, 121–141 (2008).
Gorbalenya, A. E. & Lauber, C. Bioinformatics of virus taxonomy: foundations and tools for developing sequence-based hierarchical classification. Curr. Opin. Virol. 52, 48–56 (2022).
Wertheim, J. O., Steel, M. & Sanderson, M. J. Accuracy in near-perfect virus phylogenies. Syst. Biol. 71, 426–438 (2022).
Meier-Kolthoff, J. P. & Göker, M. VICTOR: genome-based phylogeny and classification of prokaryotic viruses. Bioinformatics 33, 3396–3404 (2017).
Gregory, A. C. et al. Genomic differentiation among wild cyanophages despite widespread horizontal gene transfer. BMC Genomics 17, 930 (2016).
Bobay, L. & Ochman, H. Biological species in the viral world. Proc. Natl Acad. Sci. USA 115, 6040–6045 (2018).
Ndovie, W. et al. Exploration of the genetic landscape of bacterial dsDNA viruses reveals an ANI gap amid extensive mosaicism. mSystems https://doi.org/10.1128/msystems.01661-24 (2025).
Cook, R. et al. INfrastructure for a PHAge REference Database: identification of large-scale biases in the current collection of cultured phage genomes. PHAGE 2, 214–223 (2021).
Nelson, D. Phage taxonomy: we agree to disagree. J. Bacteriol. 186, 7029–7031 (2004).
Krupovic, M., Quemin, E. R. J., Bamford, D. H., Forterre, P. & Prangishvili, D. Unification of the globally distributed spindle-shaped viruses of the Archaea. J. Virol. 88, 2354–2358 (2014).
Rokyta, D. R., Burch, C. L., Caudle, S. B. & Wichman, H. A. Horizontal gene transfer and the evolution of microvirid coliphage genomes. J. Bacteriol. 188, 1134–1142 (2006).
Dominguez-Huerta, G. et al. Diversity and ecological footprint of Global Ocean RNA viruses. Science 376, 1202–1208 (2022).
Gregory, A. C. et al. Marine DNA viral macro- and microdiversity from Pole to Pole. Cell 177, 1109–1123 (2019).
Gregory, A. C. et al. The gut virome database reveals age-dependent patterns of virome diversity in the human gut. Cell Host Microbe 28, 724–740 (2020).
Graham, E. B. et al. A global atlas of soil viruses reveals unexplored biodiversity and potential biogeochemical impacts. Nat. Microbiol. 9, 1873–1883 (2024).
Nayfach, S. et al. Metagenomic compendium of 189,680 DNA viruses from the human gut microbiome. Nat. Microbiol 6, 960–970 (2021).
Shi, M. et al. Redefining the invertebrate RNA virosphere. Nature 540, 539–543 (2016).
Schulz, F. et al. Giant virus diversity and host interactions through global metagenomics. Nature 578, 432–436 (2020).
Camarillo-Guerrero, L. F., Almeida, A., Rangel-Pineros, G., Finn, R. D. & Lawley, T. D. Massive expansion of human gut bacteriophage diversity. Cell 184, 1098–1109 (2021).
Rand, W. M. Objective criteria for the evaluation of clustering methods. J. Am. Stat. Assoc. 66, 846–850 (1971).
Strehl, A. & Ghosh, J. Cluster ensembles—a knowledge reuse framework for combining multiple partitions. J. Mach. Learn. Res. 3, 583–617 (2003).
Larralde, M. Pyrodigal: Python bindings and interface to Prodigal, an efficient method for gene prediction in prokaryotes. J. Open Source Softw. 7, 4296 (2022).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinf. 11, 119 (2010).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Staudt, C. L., Sazonovs, A. & Meyerhenke, H. NetworKit: a tool suite for large-scale complex network analysis. Netw. Sci. 4, 508–530 (2016).
Huerta-Cepas, J., Serra, F. & Bork, P. ETE 3: reconstruction, analysis, and visualization of phylogenomic data. Mol. Biol. Evol. 33, 1635–1638 (2016).
Steinegger, M. & Söding, J. MMseqs2 enables sensitive protein sequence searching for the analysis of massive data sets. Nat. Biotechnol. 35, 1026–1028 (2017).
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q. & Vinh, L. S. UFBoot2: improving the ultrafast bootstrap approximation. Mol. Biol. Evol. 35, 518–522 (2018).
Minh, B. Q. et al. IQ-TREE 2: New models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Letunic, I. & Bork, P. Interactive Tree of Life (iTOL) v6: recent updates to the phylogenetic tree display and annotation tool. Nucleic Acids Res. 52, W78–W82 (2024).
Millard, A. et al. taxmyPHAGE: Automated taxonomy of dsDNA phage genomes at the genus and species level. Phage (New Rochelle) 6, 5–11 (2025).
Bolduc, B. vConTACT3 database v.220. Zenodo https://doi.org/10.5281/zenodo.10035618 (2023).
Bolduc, B. vConTACT3 database v.223. Zenodo https://doi.org/10.5281/zenodo.10935512 (2024).
Bolduc, B. vConTACT3 database v.223 (software repository). Bitbucket https://bitbucket.org/MAVERICLab/vcontact3/src/master/ (2025).
Acknowledgements
This work was supported by the National Science Foundation under Grants No. DBI-2149505 (iVirus2) and DBI-2022070 (BII-Implementation: the EMERGE Institute). This material is based upon work supported by the U.S. Department of Energy, Office of Science, Office of Biological and Environmental Research, under Award Number DE-SC0023307. High-performance computating was provided by the Ohio Supercomputer Center. Additional support was provided by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany’s Excellence Strategy (EXC 2051) Project-ID 390713860, the European Research Council (ERC) Consolidator grant 865694: DiversiPHI, the Alexander von Humboldt Foundation in the context of an Alexander von Humboldt-Professorship founded by German Federal Ministry of Education and Research, and the European Union’s Horizon 2020 research and innovation program, under the Marie Skłodowska-Curie Actions Innovative Training Networks grant agreement no. 955974 (VIROINF). EMA gratefully acknowledges the support of the Biotechnology and Biological Sciences Research Council (BBSRC); this research was funded by the BBSRC Institute Strategic Programme Food Microbiome and Health BB/X011054/1 and its constituent projects BBS/E/QU/230001B and BBS/E/QU/230001D, as well as the BBSRC Institute Strategic Programme Microbes and Food Safety BB/X011011/1 and its constituent projects BBS/E/QU/230002A, BBS/E/QU/230002B and BBS/E/QU/230002C.
Author information
Authors and Affiliations
Contributions
B.B. and M.B.S. designed the study. B.B., O.Z. and M.B.S. wrote the manuscript with substantial contributions from all authors. D.T., H.B.J. and B.B. performed the phylogenetic analyses and B.E.D., D.T., H.B.J. and B.B. performed the statistical and network analyses. J.G., E.M.A. and B.B. evaluated the distance metrics. B.B. developed the code with contributions from J.G.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Alexander Gorbalenya, Arthur Gruber and Guanxiang Liang for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
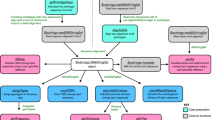
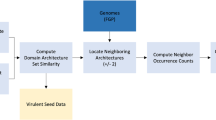
Extended Data Fig. 1 Detailed vConTACT3 workflow and outputs.
User genomes are provided to vConTACT3. ORFs are predicted with prodigal, and then sent to MMSeqs2 to be clustered at 5 identities (30%, 40%, 50%, 60%, 70%). These 5 clustering identities are used to build 5 separate protein cluster (PC) profiles, corresponding to each clustering identity. These 5 PC profiles are sent to the “Resolver,” which constructs a distance matrix (per profile) based on the selected distance metric (default “SqRoot”, see Methods). This distance matrix is subsequently converted into a network, which is annotated with any available genome information, such as realm-unique genes and proximity to reference sequences. The network is then filtered by the minimum number of shared genes allowed between genomes (see Main Text for details on recommended values), and then “repaired” if they are within the same connected component. Additionally, users can select a high-accuracy repair, which more carefully reviews dropped edges at the expense of computation time. The final stage of the resolver predicts virus realms associated with each genome using network edge-connected references and/or the presence of PCs exclusively identified and co-shared with references. Entirely novel genomes are assigned a default, user-selectable realm. The 5 filtered and repaired networks are then sent to the guilt-by-genome-association (GBGA) assigner. The GBGA aggregates realm predictions per-genome and per-network component (combining both genome and network information) and assigns a final realm for each genome. This realm prediction is used to select the optimal distance cutoff for each virus rank (genus, subfamily, family and order) determined by benchmarks (see Methods) and hierarchically cluster all genomes of that realm at that cutoff. These clusters are then matched against references (if co-clustered and/or available) or used as novel ranks to assign each virus rank order and below. (The upper ranks of phylum and class inherit reference-based assignments within the predicted realm). After assignments, GBGA output is then compared with reference sequences (if available) and performance metrics are calculated. Finally, PC profiles, performance metrics, and GBGA outputs are integrated into the exports/results component, which provides user-controlled outputs in Cytoscape format, a d3js interactive HTML network, UpSet plots, profiles, and Newick-formatted dendrograms.
Extended Data Fig. 2 vConTACT3-Extended-Data-Fig2-prokaryotes-duplo.jpg.
A 5×7 grid of line plots representing accuracies of Duplornaviria viruses infecting prokaryotes (Bacteria and Archaea). From left-to-right, plots increase in the minimum number of shared genes required to establish an edge/relationship between genomes and be considered as related. The top left plot is 1 minimum gene shared, the top right plot has 5 minimum genes shared, and moving from left to right increases minimum genes shared by 1. From the top-to-bottom direction, plots increase in minimum clustering identity used during MMSeqs2 clustering to establish protein clusters (PCs). Since PCs are used to determine the number of shared genes between genomes, increasing clustering identity increases the stringency required for two genes between two separate genomes to be considered shared, and thus, related. The top left plot is 30% clustering identity, the bottom left plot is 90% clustering identity, with each plot moving downward an increase in 10% clustering identity. Within each plot, accuracy (Y-axis) is a measure of agreement between NCBI taxonomy and vConTACT3 predictions. Pairwise distance cutoff (X-axis) represents the cutoff threshold used during hierarchical clustering to define clusters. The cutoff ranges between 0 - 0.99, with 0 representing completely identical PC profiles, and 0.99 representing nearly no shared genes. Line colors represent taxonomic rank, and dashed lines represent the type of distance metric (Jaccard, “SqRoot”, ‘VirClust” and “Shorter”) employed. See Methods for description of each distance metric.
Extended Data Fig. 3 Eukaryotic-infecting viruses of the Duplornaviria realm.
A 5×7 grid of line plots representing accuracies of Duplornaviria viruses infecting Eukaryota. Details are as Extended Data Fig. 2.
Extended Data Fig. 4 Prokaryotic-infecting viruses of the Adnaviria and Varidnaviria realms.
A 5×7 grid of line plots representing accuracies of Adnaviria and Varidnaviria viruses infecting prokaryotes (Bacteria and Archaea). Details are as Extended Data Fig. 2.
Extended Data Fig. 5 Eukaryotic-infecting viruses of the Adnaviria and Varidnaviria realms.
A 5×7 grid of line plots representing accuracies of Adnaviria and Varidnaviria viruses infecting Eukaryota. Details are as Extended Data Fig. 2.
Extended Data Fig. 6 Comparison in distance cutoffs of Duplodnaviria and Adnaviria & Varidnaviria between domains.
Accuracy plots showing the similarity in optimal cutoffs between realms, with prokaryote-infecting viruses appearing downshifted relative to Eukaryote-infecting viruses.
Extended Data Fig. 7 Prokaryotic-infecting viruses of the Monodnaviria realm.
A 5×5 grid of line plots representing accuracies of Monodnaviria viruses infecting prokaryotes (Bacteria and Archaea). Details are as Extended Data Fig. 2.
Extended Data Fig. 8 Eukaryotic-infecting viruses of the Monodnaviria realm.
A 5×5 grid of line plots representing accuracies of Monodnaviria viruses infecting Eukaryota. Details are as Extended Data Fig. 2.
Extended Data Fig. 9 Prediction labeling stability.
Line plots of adjusted rand index (ARI) and normalized mutual information (NMI) of a set of prediction labels. Colors indicate rank, dashed lines exclude singleton groups from the analysis, whereas solid lines include singletons. “Successive” plots are when ARI/NMI is calculated between fractions, “cumulative” is ARI/NMI calculated against the final (100%) fraction. High ARI and NMI indicate high agreement between genome predictions (that is labels) as more data is added, where an ARI and NMI of 1.00 indicate perfect agreement between labels between datasets.
Supplementary information
Supplementary Information (download PDF )
Supplemental Notes 1–3.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Bolduc, B., Zablocki, O., Turner, D. et al. Machine learning enables scalable and systematic hierarchical virus taxonomy. Nat Biotechnol (2025). https://doi.org/10.1038/s41587-025-02946-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-025-02946-9
This article is cited by
-
Diversity and ecological roles of hidden viral players in groundwater microbiomes
Nature Communications (2026)