Abstract
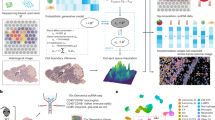
Imaging-based spatially resolved transcriptomics can localize transcripts within tissue sections in three dimensions. However, cell segmentation, which assigns transcripts to cells, is usually performed in two dimensions and spatial doublets in the vertical dimension result in segmented cells containing transcripts originating from multiple cell types. Here we present a computational tool called ovrlpy that identifies overlapping cells, tissue folds and inaccurate cell segmentation by analyzing transcript localization in three dimensions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
This study used publicly available SRT datasets. 10x Genomics Xenium datasets were downloaded from https://www.10xgenomics.com/datasets/fresh-frozen-mouse-brain-replicates-1-standard and https://www.10xgenomics.com/datasets/xenium-prime-ffpe-neonatal-mouse. Vizgen MERSCOPE datasets were downloaded from https://info.vizgen.com/mouse-brain-map and https://info.vizgen.com/mouse-liver-data. The MERFISH hypothalamus dataset was downloaded; metadata is available from the original publication at ref. 29, segmented cells are available via Dryad at https://doi.org/10.5061/dryad.8t8s248 (ref. 30), transcript information and segmentation masks are available via Zenodo at https://zenodo.org/records/3478502 (ref. 31) and the single-cell RNA sequencing reference dataset from the GEO (GSE113576). The brain reference dataset used for generating signatures was downloaded from the Allen Brain Map at https://portal.brain-map.org/atlases-and-data/rnaseq/mouse-whole-cortex-and-hippocampus-10x. The liver reference signatures were generated from the Mouse StSt dataset of the Liver cell atlas at https://www.livercellatlas.org/. Data generated as part of this study and necessary to reproduce results are available via Zenodo at https://zenodo.org/records/14226546 (ref. 32). Source data are provided with this paper.
Code availability
The ovrlpy package is available as free and open-source software via GitHub with a permissive MIT license at https://github.com/HiDiHlabs/ovrl.py. The package can also be downloaded from PyPI at https://pypi.org/project/ovrlpy/ and Bioconda at https://anaconda.org/bioconda/ovrlpy. We provide a repository with Jupyter Notebooks for reproducing all results and figures of this study via GitHub at https://github.com/HiDiHlabs/ovrlpy-publication.
References
Vandereyken, K., Sifrim, A., Thienpont, B. & Voet, T. Methods and applications for single-cell and spatial multi-omics. Nat. Rev. Genet. 24, 494–515 (2023).
Chen, H., Li, D. & Bar-Joseph, Z. SCS: cell segmentation for high-resolution spatial transcriptomics. Nat. Methods 20, 1237–1243 (2023).
Fang, S. et al. Computational approaches and challenges in spatial transcriptomics. Genom. Proteom. Bioinform. 21, 24–47 (2023).
Fu, X. et al. BIDCell: biologically-informed self-supervised learning for segmentation of subcellular spatial transcriptomics data. Nat. Commun. 15, 509 (2024).
Shu, J., Fu, H., Qiu, G., Kaye, P. & Ilyas, M. Segmenting overlapping cell nuclei in digital histopathology images. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2013, 5445–5448 (2013).
Palokangas, S., Selinummi, J. & Yli-Harja, O. Segmentation of folds in tissue section images. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2007, 5642–5645 (2007).
Taqi, S. A., Sami, S. A., Sami, L. B. & Zaki, S. A. A review of artifacts in histopathology. J. Oral Maxillofac. Pathol. 22, 279 (2018).
Moses, L. & Pachter, L. Museum of spatial transcriptomics. Nat. Methods 19, 534–546 (2022).
Yao, Z. et al. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317–332 (2023).
Allen, N. J. & Lyons, D. A. Glia as architects of central nervous system formation and function. Science 362, 181–185 (2018).
Megı́as, M., Emri, Z., Freund, T. F. & Gulyás, A. I. Total number and distribution of inhibitory and excitatory synapses on hippocampal CA1 pyramidal cells. Neuroscience 102, 527–540 (2001).
Park, J. et al. Cell segmentation-free inference of cell types from in situ transcriptomics data. Nat. Commun. 12, 3545 (2021).
Müller-Bötticher, N., Tiesmeyer, S., Eils, R. & Ishaque, N. Sainsc: a computational tool for segmentation-free analysis of in situ capture data. Small Methods 9, 2401123 (2025).
Petukhov, V. et al. Cell segmentation in imaging-based spatial transcriptomics. Nat. Biotechnol. 40, 345–354 (2022).
Jones, D. C. et al. Cell simulation as cell segmentation. Nat. Methods 22, 1331–1342 (2025).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291 (2019).
Marco Salas, S. et al. Optimizing Xenium in situ data utility by quality assessment and best-practice analysis workflows. Nat. Methods 22, 813–823 (2025).
Mitchel, J., Gao, T., Petukhov, V., Cole, E. & Kharchenko, P. V. Impact and correction of segmentation errors in spatial transcriptomics. Nat. Genet. https://doi.org/10.1038/s41588-025-02497-4 (2026).
Fresh Frozen Mouse Brain Replicates, In Situ Gene Expression Dataset Analyzed using Xenium Onboard Analysis 1.0.2 (10x Genomics, 2023).
Pokrajac, N. T., Tokarew, N. J. A., Gurdita, A., Ortin-Martinez, A. & Wallace, V. A. Meningeal macrophages inhibit chemokine signaling in pre-tumor cells to suppress mouse medulloblastoma initiation. Dev. Cell 58, 2015–2031 (2023).
Remsik, J. et al. Characterization, isolation, and in vitro culture of leptomeningeal fibroblasts. J. Neuroimmunol. 361, 577727 (2021).
Tiklová, K. et al. Single cell transcriptomics identifies stem cell-derived graft composition in a model of Parkinson’s disease. Nat. Commun. 11, 2434 (2020).
Pietilä, R. et al. Molecular anatomy of adult mouse leptomeninges. Neuron 111, 3745–3764.e7 (2023).
Howe, J. R. et al. Control of innate olfactory valence by segregated cortical amygdala circuits. Preprint at bioRxiv https://doi.org/10.1101/2024.06.26.600895 (2024).
Mah, C. K. et al. Bento: a toolkit for subcellular analysis of spatial transcriptomics data. Genome Biol. 25, 82 (2024).
Vizgen Data Release V1.0 (Vizgen, 2021).
Vizgen Post-processing Tool (Vizgen, 2025).
Whole Mouse Pup (FFPE) with 5 K Mouse Pan Tissue and Pathways Panel and Snap25a/Snap25b Add-on, In Situ Gene Expression Dataset Analyzed using Xenium Onboard Analysis 3.0.0 (10x Genomics, 2024).
Moffitt, J. R. et al. Molecular, spatial, and functional single-cell profiling of the hypothalamic preoptic region. Science 362, eaau5324 (2018).
Moffitt, J. R. et al. Data from: molecular, spatial and functional single-cell profiling of the hypothalamic preoptic region. Dryad https://doi.org/10.5061/dryad.8t8s248 (2018).
Park, J. et al. Supplemental data for: segmentation-free inference of cell types from in situ transcriptomics data. Zenodo https://doi.org/10.5281/zenodo.3478502 (2019).
Tiesmeyer, S. & Müller-Bötticher, N. Data to re-create ‘2D, or not 2D? Investigating vertical signal integrity of tissue slices’. Zenodo https://doi.org/10.5281/zenodo.16736589 (2025).
Acknowledgements
The ovrlpy project was conceptualized during the de.NBI BioHackathon SpaceHack project in Lutherstadt-Wittenberg (December 2022); we thank the organizers of and participants in the de.NBI BioHackathon SpaceHack project. We thank T. Rheude for establishing an analogy of 2D analysis of tissue sections to the flat earth conspiracy. This research has received funding from the Federal Ministry of Education and Research of Germany in the framework of SAGE (project number 031L0265, S.T. and N.M.-B.) and CNAScope (grant no. 01KD2443, A.M.), the German Research Foundation (DFG TRR 412, Project-ID 35081457, N.I. and F.J.T.), ELIXIR Spatial2Galaxy (N.I.) and the project grant (no. 2024-02533, S.M.-S. and M.N.) from the Swedish Research Council. We thank the GESTALT community for positive engagement and ideas. We thank the following people for useful discussion and sharing data on tissue preparation and other artifacts: H. C. Etchevers, Aix Marseille University, for the Visium mouse heart samples with folds, smears and slippage; T. Conrad, Berlin Institute of Health at Charité, F. Baumgartner and U. Keller, Charité—Universitätsmedizin Berlin, for the Xenium human uveal melanoma sample with detachment; O. Raineteau, G. Marcy and C. Dégletagne, Cancer Research Center of Lyon, for the Xenium mouse brain samples with tissue folds; and M. Grillo and C. M. Langseth, Stockholm University, for the ISS and Xenium multiple sclerosis samples with signal smearing.
Author information
Authors and Affiliations
Contributions
N.I. conceived of and designed the study. S.T. and N.M.-B. with some assistance from A.M. implemented the ovrlpy package. S.T., N.M.-B., A.M., L.M., S.M.-S., L.B.K. and N.I. performed the data analysis. S.T., N.M.-B., B.L. and N.I. interpreted the brain sample analyses. S.T., P.K., P.H., A.G., C.K. and N.I. interpreted the liver sample analysis. S.T., N.M.-B. and N.I. wrote the paper. All authors proofread and corrected the paper. All authors contributed to the article and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
S.M.-S. is a co-founder of Spatialist, a data analysis company focused on spatial omics. P.H.’s laboratory has received research grants (funding to the institution) from MSD. P.H. has received lecture fees from AstraZeneca, Falk and Orphalan, as well as travel support from Falk, Ipsen and Boehringer Ingelheim. A.G.’s laboratory has received research grants (funding to the institution) from Agomab. F.T.’s laboratory has received research grants (funding to the institution) from Agomab, AstraZeneca, MSD and Gilead. F.T. has received honoraria for consulting or lectures from Gilead, AbbVie, Falk, AstraZeneca, Boehringer Ingelheim, Madrigal, MSD, GSK, Ipsen, Mirum, Novartis, Novo Nordisk and Sanofi. F.J.T. consults for Immunai, Singularity Bio, CytoReason and Omniscope, and holds ownership interests in Dermagnostix and Cellarity. N.I.’s laboratory has received research grants (funding to the institution) from Owkin. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Lambda Moses and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Supervised annotation of virtually subsliced cell segments implicates widespread presence of vertical doublets in the Xenium mouse brain dataset.
a, Map of cell segment centroids marked as singlets or vertical doublets depending on their consistency of the MapMyCells assigned class across two virtual vertical subslices. Vertical doublets appear widespread over the tissue, but are more concentrated on the lower, ventral part of the sample. b, Cell type-specific observed vertical doublets (row-normalized).
Extended Data Fig. 2 Ovrlpy-identified vertical doublets are consistent with supervised segmentation-based doublets.
a, Vertical signal integrity (VSI) map of the Xenium mouse brain dataset, indicating 7 of the top vertical doublets identified by ovrlpy. b, UMAP and RGB embedding of the gene expression sampled at local maxima. Different cell types appear as separate clusters in the UMAP as well as the RGB embedding. c–i, Close-up of highlighted regions in panel a. The color-embedded transcripts highlight the different cell types and structures involved in spatial vertical doublets (column 1-3, RGB color-embedded as shown in panel b. VSI map of the highlighted region (column 4). Marker transcripts are based on the Xenium gene panel design (column 5, 6). Cell types are based on MapMyCells annotation of the top and bottom virtual subslice of nuclear transcripts (column 7, 8). Scale bar: 15 μm. Panel c additionally shows DAPI images of multiple focal planes (spaced 3 µm apart) highlighting the different layers in the fold. Distribution of VSI in the (j) entire dataset and (k) the tissue fold in the ventral region (highlighted in panel a).
Extended Data Fig. 3 Vizgen’s 3D segmentation fails to identify most cell overlaps detected by ovrlpy in the MERSCOPE mouse brain dataset.
a, UMAP and RGB embedding of the gene expression sampled at local maxima. b, Gene expression map of the MERSCOPE mouse brain dataset colored using ovrlpy RGB embeddings at local maxima sampling locations. c, Vertical signal integrity (VSI) map of the whole brain slice. The ten most prominent VSI local minima identified by ovrlpy are highlighted. d–m, Visualizations of vertical doublets. First three columns show transcript maps of top and bottom virtual subslices, and from the sides. Transcripts are colored by ovrlpy RGB color. Last column shows the VSI map in the corresponding region with overlayed segmentation masks (merged across all segment layers per cell). Scale bar: 15 μm.
Extended Data Fig. 4 Nuclear stain supports ovrlpy-detected vertical doublets.
a, UMAP and RGB embedding of the gene expression sampled at local maxima. b, Gene expression map of the MERSCOPE mouse brain dataset colored using ovrlpy RGB embeddings at local maxima sampling locations. c, Vertical signal integrity (VSI) map of the whole brain slice. Four selected out of the ten most prominent VSI local minima identified by ovrlpy are highlighted and visualized in subsequent panels. d–g, Visualizations of vertical doublets. First 2 columns show transcript maps of top and bottom virtual subslices (colored by ovrlpy RGB embedding). Third column shows the VSI map in the corresponding region. The other columns correspond to the DAPI-stained images of multiple imaging focal planes (1.5 µm distance between adjacent planes). The arrows indicate nuclei participating in the overlap and are colored according to the RGB embedding in the first two columns. Scale bar: 5 µm.
Extended Data Fig. 5 Ovrlpy’s unsupervised vertical doublet filtering enhances cell-type separation in gene expression space.
a–d, UMAP embeddings of gene expression data from Xenium mouse brain cell segments colored by (a) mean vertical signal integrity (VSI), (b) singlets and vertical doublets (VSI threshold of 0.7; 162,033 cell segments of which 36,846 are vertical doublets), (c) MapMyCells cell type annotations, (d) MapMyCells cell type annotations (UMAP embedding after excluding vertical doublets). Same as a-d for e–h, the Xenium mouse brain with only nuclear transcripts (162,018 nuclei of which 36,831 are vertical doublets), and i–l, the nuclear segmented MERSCOPE mouse brain (83,505 nuclei of which 15,780 are vertical doublets).
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Results and Figs. 1–15.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tiesmeyer, S., Müller-Bötticher, N., Malt, A. et al. Identifying 3D signal overlaps in spatial transcriptomics data with ovrlpy. Nat Biotechnol (2026). https://doi.org/10.1038/s41587-026-03004-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-026-03004-8