Abstract
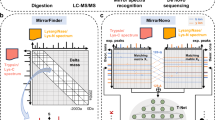
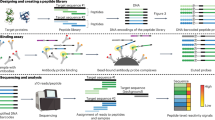
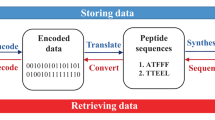
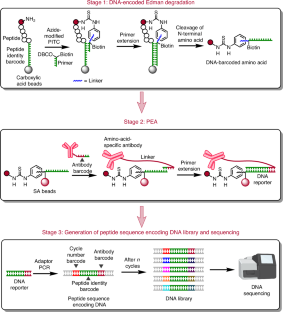
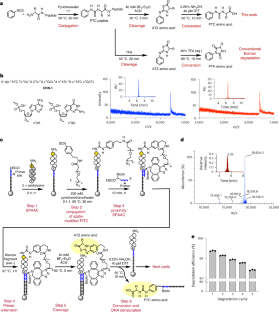
Despite advances in mass spectrometry and emerging single-molecule approaches, sequencing peptides at the single-molecule level remains a central challenge in proteomics. Here we present a ‘reverse translation’ strategy that enables single-molecule peptide sequencing with single-amino-acid resolution. In this approach, peptides undergo a modified Edman degradation that iteratively releases N-terminal amino acids tagged with peptide-specific DNA barcodes. Antibody-mediated proximity extension assays identify these barcoded amino acids and generate PCR-amplifiable DNA reporters that record the identity, position and originating peptide of each amino acid. The resulting DNA library is directly read by high-throughput sequencing, converting peptide sequences into digital DNA outputs. Using this approach, we demonstrate true single-molecule peptide sequencing, achieving full sequence coverage in millions of reads and accurate differentiation of both native and post-translationally modified peptides. These results establish a framework that redefines protein sequencing as a DNA sequencing problem and lays the foundation for high-throughput, de novo single-molecule protein sequencing.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
Data availability
The data supporting the findings of this study are available within the article and its Supplementary Information. The next-generation sequencing reads for ensemble and single-molecule peptide sequencing were deposited to the National Center for Biotechnology Information Sequence Read Archive under BioProjects PRJNA1420480 and PRJNA1423337, respectively. Source data are provided with this paper.
Code availability
The C++ and Python scripts for data processing and visualization of single-molecule peptide sequencing are available from GitHub (https://github.com/whulwzheng-source/smPeptideSeq).
Change history
25 March 2026
In the version of this article initially published, the Peer review information section was incomplete and is now amended in the HTML and PDF versions of the article.
References
Pollen, A. A. et al. Low-coverage single-cell mRNA sequencing reveals cellular heterogeneity and activated signaling pathways in developing cerebral cortex. Nat. Biotechnol. 32, 1053–1058 (2014).
Wang, N. et al. Single-cell microRNA-mRNA co-sequencing reveals non-genetic heterogeneity and mechanisms of microRNA regulation. Nat. Commun. 10, 95 (2019).
Kolodziejczyk, A. A., Kim, J. K., Svensson, V., Marioni, J. C. & Teichmann, S. A. The technology and biology of single-cell RNA sequencing. Mol. Cell 58, 610–620 (2015).
Sonneveld, S., Verhagen, B. M. P. & Tanenbaum, M. E. Heterogeneity in mRNA Translation. Trends Cell Biol. 30, 606–618 (2020).
Smith, L. M. et al. The Human Proteoform Project: defining the human proteome. Sci. Adv. 7, eabk0734 (2021).
Tajik, M., Baharfar, M. & Donald, W. A. Single-cell mass spectrometry. Trends Biotechnol. 40, 1374–1392 (2022).
Ye, Z. et al. One-Tip enables comprehensive proteome coverage in minimal cells and single zygotes. Nat. Commun. 15, 2474 (2024).
Guzman, U. H. et al. Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition. Nat. Biotechnol. 1–12 https://doi.org/10.1038/s41587-023-02099-7 (2024).
Huffman, R. G. et al. Prioritized mass spectrometry increases the depth, sensitivity and data completeness of single-cell proteomics. Nat. Methods 20, 714–722 (2023).
MacCoss, M. J. et al. Sampling the proteome by emerging single-molecule and mass spectrometry methods. Nat. Methods 20, 339–346 (2023).
Alfaro, J. A. et al. The emerging landscape of single-molecule protein sequencing technologies. Nat. Methods 18, 604–617 (2021).
Restrepo-Pérez, L., Joo, C. & Dekker, C. Paving the way to single-molecule protein sequencing. Nat. Nanotechnol. 13, 786–796 (2018).
Reed, B. D. et al. Real-time dynamic single-molecule protein sequencing on an integrated semiconductor device. Science 378, 186–192 (2022).
Martin-Baniandres, P. et al. Enzyme-less nanopore detection of post-translational modifications within long polypeptides. Nat. Nanotechnol. 18, 1335–1340 (2023).
Yu, L. et al. Unidirectional single-file transport of full-length proteins through a nanopore. Nat. Biotechnol. 41, 1130–1139 (2023).
Brinkerhoff, H., Kang, A. S. W., Liu, J., Aksimentiev, A. & Dekker, C. Multiple rereads of single proteins at single-amino acid resolution using nanopores. Science 374, 1509–1513 (2021).
Nova, I. C. et al. Detection of phosphorylation post-translational modifications along single peptides with nanopores. Nat. Biotechnol. 42, 710–714 (2023).
Motone, K. et al. Multi-pass, single-molecule nanopore reading of long protein strands. Nature 633, 662–669 (2024).
Ouldali, H. et al. Electrical recognition of the twenty proteinogenic amino acids using an aerolysin nanopore. Nat. Biotechnol. 38, 176–181 (2020).
Wang, K. et al. Unambiguous discrimination of all 20 proteinogenic amino acids and their modifications by nanopore. Nat. Methods 21, 92–101 (2024).
Zhang, M. et al. Real-time detection of 20 amino acids and discrimination of pathologically relevant peptides with functionalized nanopore. Nat. Methods 21, 609–618 (2024).
Swaminathan, J. et al. Highly parallel single-molecule identification of proteins in zeptomole-scale mixtures. Nat. Biotechnol. 36, 1076–1082 (2018).
Boutureira, O. & Bernardes, G. J. L. Advances in chemical protein modification. Chem. Rev. 115, 2174–2195 (2015).
Szijj, P. A., Kostadinova, K. A., Spears, R. J. & Chudasama, V. Tyrosine bioconjugation—an emergent alternative. Org. Biomol. Chem. 18, 9018–9028 (2020).
Choi, W. S. et al. Structural basis for the recognition of N-end rule substrates by the UBR box of ubiquitin ligases. Nat. Struct. Mol. Biol. 17, 1175–1181 (2010).
Stein, B. J., Grant, R. A., Sauer, R. T. & Baker, T. A. Structural basis of an N-degron adaptor with more stringent specificity. Structure 24, 232–242 (2016).
Edman, P., Högfeldt, E., Sillén, L. G. & Kinell, P.-O. Method for determination of the amino acid sequence in peptides. Acta Chem. Scand. 4, 283–293 (1950).
McGregor, L. M., Gorin, D. J., Dumelin, C. E. & Liu, D. R. Interaction-dependent PCR: identification of ligand−target pairs from libraries of ligands and libraries of targets in a single solution-phase experiment. J. Am. Chem. Soc. 132, 15522–15524 (2010).
Zhang, P. et al. Highly sensitive serum protein analysis using magnetic bead-based proximity extension assay. Anal. Chem. 94, 12481–12489 (2022).
Zoltewicz, J. A., Clark, D. F., Sharpless, T. W. & Grahe, G. Kinetics and mechanism of the acid-catalyzed hydrolysis of some purine nucleosides. J. Am. Chem. Soc. 92, 1741–1750 (1970).
Hong, J. M. et al. ProtSeq: toward high-throughput, single-molecule protein sequencing via amino acid conversion into DNA barcodes. iScience 25, 103586 (2022).
Iida, T., Santa, T., Toriba, A. & Imai, K. Semi-automatic amino acid sequencing and D/L-configuration determination of peptides with detection of liberated N-terminal phenylthiocarbamoylamino acids. Analyst 123, 2829–2834 (1998).
Matsunaga, H. et al. Proton: a major factor for the racemization and the dehydration at the cyclization/cleavage stage in the Edman sequencing method. Anal. Chem. 68, 2850–2856 (1996).
Mitchell, M. J., Hirschowitz, W., Rastinejad, F. & Lu, P. Boron trifluoride–methanol complex as a non-depurinating detritylating agent in DNA synthesis. Nucleic Acids Res. 18, 5321 (1990).
Seela, F., Menkhoff, S. & Behrendt, S. Furanoside–pyranoside isomerization of tubercidin and its 2′-deoxy derivatives: influence of nucleobase and sugar structure on the proton-catalysed reaction. J. Chem. Soc. Perkin Trans. 2, 525–530 (1986).
Potowski, M. et al. Chemically stabilized DNA barcodes for DNA-encoded chemistry. Angew. Chem. Int. Ed. 60, 19744–19749 (2021).
Farnsworth, V. & Steinberg, K. The generation of phenylthiocarbamyl or anilinothiazolinone amino acids from the postcleavage products of the Edman degradation. Anal. Biochem. 215, 200–210 (1993).
Matsudaira, P. (ed.) A Practical Guide to Protein and Peptide Purification for Microsequencing (Academic Press, 1993).
Rydberg, P., Lüning, B., Wachtmeister, C. A., Eriksson, L. & Törnqvist, M. Applicability of a modified Edman procedure for measurement of protein adducts: mechanisms of formation and degradation of phenylthiohydantoins. Chem. Res. Toxicol. 15, 570–581 (2002).
Flood, D. T. et al. Expanding reactivity in DNA-encoded library synthesis via reversible binding of DNA to an inert quaternary ammonium support. J. Am. Chem. Soc. 141, 9998–10006 (2019).
MacConnell, A. B., McEnaney, P. J., Cavett, V. J. & Paegel, B. M. DNA-encoded solid-phase synthesis: encoding language design and complex oligomer library synthesis. ACS Comb. Sci. 17, 518–534 (2015).
Halpin, D. R., Lee, J. A., Wrenn, S. J. & Harbury, P. B. DNA display III. Solid-phase organic synthesis on unprotected DNA. PLoS Biol. 2, e175 (2004).
Wang, Z., Li, D., Tian, X. & Zhang, C. A copper-free and enzyme-free click chemistry-mediated single quantum dot nanosensor for accurate detection of microRNAs in cancer cells and tissues. Chem. Sci. 12, 10426–10435 (2021).
Nakano, S. & Sugimoto, N. The structural stability and catalytic activity of DNA and RNA oligonucleotides in the presence of organic solvents. Biophys. Rev. 8, 11–23 (2016).
Niall, H. D. Automated Edman degradation: the protein sequenator. Methods Enzymol. 27, 942–1010 (1973).
Laursen, R. A. Solid-phase Edman degradation. Eur. J. Biochem. 20, 89–102 (1971).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Budnik, B., Levy, E., Harmange, G. & Slavov, N. SCoPE-MS: mass spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation. Genome Biol. 19, 161 (2018).
Kokoris, M. et al. Sequencing by expansion (SBX)—a novel, high-throughput single-molecule sequencing technology. Preprint at bioRxiv https://doi.org/10.1101/2025.02.19.639056 (2025).
Bloom, S. et al. Decarboxylative alkylation for site-selective bioconjugation of native proteins via oxidation potentials. Nat. Chem. 10, 205–211 (2018).
Zhang, L. et al. Photoredox-catalyzed decarboxylative C-terminal differentiation for bulk- and single-molecule proteomics. ACS Chem. Biol. 16, 2595–2603 (2021).
Rehm, F. B. H. et al. Enzymatic C-terminal protein engineering with amines. J. Am. Chem. Soc. 143, 19498–19504 (2021).
Antos, J. M. et al. Site-specific N- and C-terminal labeling of a single polypeptide using sortases of different specificity. J. Am. Chem. Soc. 131, 10800–10801 (2009).
Boga, S. B. et al. Site-selective synthesis of insulin azides and bioconjugates. Bioconjug. Chem. 30, 1127–1132 (2019).
Edman, P. & Begg, G. A protein sequenator. Eur. J. Biochem. 1, 80–91 (1967).
Swaminathan, J., Boulgakov, A. A. & Marcotte, E. M. A theoretical justification for single molecule peptide sequencing. PLoS Comput. Biol. 11, e1004080 (2015).
Yang, K.-A. et al. Recognition and sensing of low-epitope targets via ternary complexes with oligonucleotides and synthetic receptors. Nat. Chem. 6, 1003–1008 (2014).
Boder, E. T., Midelfort, K. S. & Wittrup, K. D. Directed evolution of antibody fragments with monovalent femtomolar antigen-binding affinity. Proc. Natl Acad. Sci. USA 97, 10701–10705 (2000).
Javanpour, A. A. & Liu, C. C. Evolving small-molecule biosensors with improved performance and reprogrammed ligand preference using OrthoRep. ACS Synth. Biol. 10, 2705–2714 (2021).
Lee, G. R. et al. Small-molecule binding and sensing with a designed protein family. Preprint at bioRxiv https://doi.org/10.1101/2023.11.01.565201 (2023).
An, L. et al. Binding and sensing diverse small molecules using shape-complementary pseudocycles. Science 385, 276–282 (2024).
Acknowledgements
This work was supported by the Helmsley Trust, Wellcome LEAP SAVE program. We thank A. Hugenmatter at Stanford Innovative Medicines Accelerator for the helpful discussion. We thank J. Lowitz at Antibody Solutions for his assistance with custom antibody generation. We thank T. McLaughlin at Stanford University MS for her assistance with developing LC–MS methods for oligonucleotides. This work was supported by the Vincent Coates Foundation MS Laboratory, Stanford University MS (RRID:SCR_017801) using the Bruker Microflex MALDI-TOF MS instrument (RRID:SCR_018696) and Thermo Exploris 240 LC–MS system (RRID:SCR_022216) that was purchased with funding from Stanford C-ShaRP (RRID:SCR_022986).
Author information
Authors and Affiliations
Contributions
L.Z. and H.T.S. conceptualized the study. L.Z. and Y.S. performed the experiments and analyzed the experimental data. L.Z. and L.A.H. analyzed the single-molecule peptide sequencing data. H.T.S. supervised the research. L.Z. and H.T.S. wrote the original draft. L.Z., Y.S., M.E. and H.T.S. reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
L.Z., Y.S. and H.T.S. are listed as coinventors on a pending patent application related to this work filed at the US Patent and Trademark Office (no. PCT/US2024/017167). L.A.H. and M.E. declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks Chirlmin Joo, Stefan Howorka and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Materials, procedure for the preparation of oligonucleotides and organic compounds, Supplementary Text, Figs. 1–18 and Tables 1, 3–7, MS spectra of oligonucleotides, MS spectra of antibody–oligonucleotide conjugates, nuclear magnetic resonance spectra of organic compounds and uncropped gel images.
Supplementary Table 2 (download XLSX )
List of oligonucleotides.
Source data
Source Data Fig. 3 (download PDF )
Unprocessed gel.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zheng, L., Sun, Y., Hein, L.A. et al. Single-molecule peptide sequencing through reverse translation of peptides into DNA. Nat Biotechnol (2026). https://doi.org/10.1038/s41587-026-03061-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-026-03061-z