Abstract
Adeno-associated viruses (AAVs) are preferred gene therapy vectors because of their versatility, durability and safety profile. Here, we demonstrate extensive chimerism, manifesting as pervasive barcode swapping, among complex recombinant AAV (rAAV) libraries that are packaged as a pool. The observed chimerism is length and homology dependent but capsid independent, in some cases affecting the majority of packaged rAAV genomes. These results have implications for the design and deployment of functional rAAV libraries.
Similar content being viewed by others
Main
Multiplexed functional genomic screens often use linked components within a cargo, for example, single-cell CRISPR screens1,2 (single guide RNA (sgRNA) and barcode) or massively parallel reporter assays (MPRAs; regulatory element and barcode)3. In these, any level of decoupling of expected pairings ultimately degrades signal quality. A prominent example is the frequent recombination seen in lentiviral cargo packaging4,5. Such recombination, which is length and homology dependent and results from lowly processive reverse transcription and template switching during replication6, stymied early single-cell functional genomics efforts using this delivery strategy7.
By contrast, pervasive chimeric rearrangements of recombinant adeno-associated virus (rAAV) genetic material during packaging have not been described to our knowledge. Yet, recent studies of long-read sequenced AAV-packaged DNA have revealed unexpected DNA arrangements8,9. In parallel, high levels of noise and limited dynamic range are commonly observed in barcoded MPRA experiments using rAAVs10,11,12,13,14. These hint at possible unknown complexities during rAAV packaging.
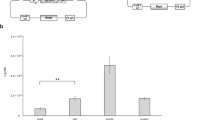
To explicitly test for chimera formation during rAAV packaging, we performed barcode swap experiments. Specifically, we constructed a series of rAAV libraries harboring a large number of uniquely associated pairs of barcodes separated by different inserts and flanked by AAV2 inverted terminal repeats (ITRs) (Fig. 1a; all plasmids and oligos listed in Supplementary Data 1). The six libraries harbored inserts of three different lengths (short: ~225 bp, mid-sized: ~800 bp, long: ~2.1 kb) each with two homology classes (homologous: identical sequences, nonhomologous: size-matched to homologous counterparts and bearing tagmented, narrowly size-selected Escherichia coli genomic DNA; Fig. 1a and Extended Data Figs. 1 and 2). Each insert further had a short internal sequence index for downstream demultiplexing (Extended Data Fig. 1d). The resulting libraries were complex (>5 million barcode pairs in parental library p146, bottlenecked to 15,000–45,000 pairs in libraries p149–p154). Inserts were size-adjusted outside the BC1–BC2 intervening sequence with filler sequences to fix the total ITR-to-ITR length to ~2.3 kb for inserts of different sizes (Extended Data Fig. 1c; see library p153× below). These libraries were then packaged separately into AAV capsids (all with PHP.eB, some with AAV2 serotypes, 14 packaging conditions total; Supplementary Data 2 and Methods).
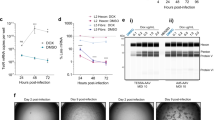
a, A complex doubly barcoded cloning dock with associated dictionary of valid BC1–BC2 pairs was constructed and served as the starting point to clone libraries of inserts of varying lengths and homology class within AAV2 ITRs (six separate libraries: [short ~0.2 kb, mid ~0.8 kb, long ~2.1 kb] × [homologous, nonhomologous]). The seven different libraries considered are schematized (Extended Data Figs. 1 and 2). b, Each cloned barcoded library was separately: (1) digested to liberate the barcoded insert and (2) AAV-packaged. Both plasmid-derived insert and AAV DNA were submitted for direct long-read sequencing. Resulting long reads were scanned for barcodes and the fraction of discordant BC1–BC2 pairs, as compared to the bona fide parental dictionary, was determined. c, Quantification of the fraction of discordant barcode pairs as a function of the full-length BC-to-BC average size. Left, plasmid DNA; middle, AAV-packaged DNA; right, zoomed-in view of y axis range 0–0.1. Each point corresponds to swap quantification for a library for both plasmid-derived (square) and AAV-derived (circle) material (n = 1 replicate per library). For each data point, we analyzed full-length BC-to-BC reads passing quality control filters and having separately valid BC1 and BC2 (Supplementary Figs. 3a and 4a). Quantifications derived from library p153× corresponding to the pool of multiple-sized inserts in a single AAV-packaged sample are marked by an ×. Swaps are significantly higher (one-sided bootstrap FDR < 10−5) for the homologous versus their respective size-matched nonhomologous libraries. d, The impact of starting plasmid dose on BC1–BC2 chimerism was assessed by packaging library p151:mid-hom with different starting amount per 15-cm plate of producing cells, ranging from 15 μg (100% dose) to 15 ng (0.1% dose). The total input of transfected DNA was kept fixed by adding ITR-free plasmid (Supplementary Fig. 6). Vertical error bars correspond to the 20th–80th percentiles from bootstrap resampling to document read counting noise (center: quantification from all reads). Horizontal error bars to the 10th–90th percentile of the BC-to-BC length distribution from plasmid digest inserts. Data from each library or condition is from one packaging replicate, with separate packaging for distinct libraries (Supplementary Data 2). A second biological packaging replicate is presented in Fig. 2b, with quantitative reproducibility, for libraries p151 and p152.
To assess for chimeras, defined as discordant barcode pairs within a single read, we performed long-read sequencing of the barcoded inserts (Fig. 1b) using PCR-free library preparation, from both sized-selected digested plasmids (‘zero-swap’ controls) and AAV-packaged DNA, on the Oxford Nanopore Technology (ONT) platform (through Plasmidsaurus). Focusing on full-length BC-to-BC reads bearing all BC adjacent signposts in their proper positions and exact but separate BC1 and BC2 matches (Extended Data Fig. 1d and Supplementary Fig. 1a,b), we measured what fraction of reads showed discordant BC1–BC2 pairs in each sample (Fig. 1c and Supplementary Figs. 1e and 2). As expected, the parental backbone p146 BC1–BC2 plasmid-derived inserts library exhibited near-complete concordance with the reference BC1–BC2 dictionary (0.1% discordant pairs; Fig. 1c, left), underscoring the robustness of our bioinformatic pipeline. Furthermore, the ‘no-swap’ controls (that is, inserts digested and size-selected from plasmid libraries p149–p154) showed low but slightly increased discordance (<2.5%; Fig. 1c, right inset), suggesting nonzero chimerism generated in the cloning process (the highest discordance library being in the short homologous p149, hinting at possible recombination during Gibson assembly).
However, homologous insert AAV-packaged libraries exhibited dramatically increased rates of discordance, ranging from ~20% for the short inserts to >60% for the long inserts (Fig. 1c, middle). By contrast, nonhomologous insert libraries displayed largely concordant barcode pairs in AAV-packaged DNA independent of length (≤6% discordance, significantly lower than size-matched homologous libraries; false discovery rate (FDR) < 10−5 according to bootstrap analysis). We note that, because of the library construction process (tagmentation followed by PCR), even the nonhomologous inserts shared short regions of homology corresponding to the Tn5 adaptors at both ends (33 + 34 = 67 bp; Extended Data Fig. 1d), which could contribute to the low but nonzero rates of chimerism in these libraries. Together, these results indicate extensive molecular chimerism forming during rAAV packaging, in some cases representing the majority of species, and suggest that the rate of chimera formation is dependent on the length of intervening homologous sequence.
In addition to length and homology, what other parameters modulate the level of chimerism? In line with studies focusing on maintaining linkage for capsid sequence engineering15,16,17,18, we reasoned that chimerism could be a function of the number of different rAAV plasmids received per cell during packaging. To test this hypothesis, we packaged library p151:mid-hom at four different doses spanning a 1,000-fold input range (15 ng to 15 μg per 15-cm plate of cells, total DNA fixed using ITR-free carrier plasmid; Fig. 1d and Supplementary Fig. 3). As a control, we repackaged p152:mid-nonhom at the 100% dose in parallel. Long-read sequencing (ONT, Plasmidsaurus) of AAV-packaged DNA revealed extensive chimerism in p151:mid-hom and a robust trend toward decreasing discordant BC1–BC2 fraction at lower input doses. Notably, the trend exhibited a shallow logarithmic scaling, with ~9% fewer swaps per tenfold decrease in input. We note that, even at the lowest dose, at most ~102 plasmids may be delivered per cell (15 ng = 2.4 billion × 6-kb plasmids; with 20 million cells, this equates to ~120 plasmids per cell) and the actual internalized copy number was previously estimated by qPCR to be ~10 per cell at such a dose16. Hence, in line with previous bespoke estimates of cotransfection with pairs of GFP or mCherry plasmids at a similarly low dose15, reaching a truly limiting regime likely will require substantially lower input plasmid concentration. Given that the titer at the lowest input dose remained substantial (Supplementary Data 2), this dosage series suggests a partial mitigation strategy for applications that are not critically dependent on high viral titers. Regardless, the decrease in chimerism as a function of input dose supports the view that these BC1–BC2 swapping events occur in cells during the packaging process.
To assess whether the observed chimerism was exclusive to serotype PHP.eB, library p153× was packaged with AAV2, revealing a similar level of barcode swaps (Extended Data Fig. 3) at both 100% and 10% input dose conditions. As before, sparser packaging did decrease chimerism (bootstrap FDR < 0.005 in all instances). These results suggest that AAV chimeras form in a variety of capsids and packaging conditions.
What alternative explanations would account for the observed chimerism? The AAV sequencing library preparation from Plasmidsaurus, based on a recent protocol19, relies on annealing of the ssAAV DNA followed by end repair and adaptor ligation. End repair can, in principle, induce a single polymerization event, such that incomplete products with an insert-internal 3′ end could prime homologous counterparts, leading to technically induced chimeras. For this mechanism to manifest, these subgenomic components with an internal 3′ end would need to be packaged to be coextracted with other encapsidated DNA molecules. This is, however, unlikely, as AAVs are packaged from their 3′ ITR20,21, such that subgenomes truncated at the 3′ end would not be preferentially packaged. Furthermore, given the observed BC1–BC2 swapping proportions, truncated or incomplete AAV genomes22 would need to make up a substantial proportion of the packaged material commensurate with the observed barcode swapping frequency. These are, however, a rare class of intermediates compared to snapback products23 (partial genomes flanked by ITRs on both ends). Lastly, AAV dimers generated from ITR priming, which serve as an internal control for this effect, are rarely observed in ONT libraries generated using this method (categorization of subgenomic fragments in Extended Data Fig. 4d–f).
To obtain further support for our interpretation, we performed several controls to confirm that the observed chimerism was not an artifact of the specific ONT long-read preparation procedure. First, as an alternative to the direct annealing-based library preparation from AAV-packaged DNA, we generated double-stranded DNA libraries of barcoded inserts by PCR templated from both plasmids and purified AAV-packaged genomes. Of note, PCR libraries avoid a possible confounder of libraries composed of large segments of nonhomology (p152:mid-nonhom and p154:long-nonhom), which might have hindered the annealing step in the direct AAV ONT preparation. Plasmid templates were diluted to have a similar number of PCR cycles total compared to the AAV genomes templates (n = 16–20 cycles for AAV samples, n = 18 cycles from plasmid). Long-read sequencing (Plasmidsaurus) and bioinformatic processing of the PCR-generated libraries as before revealed near-quantitative agreement in the level of swaps compared to the PCR-free samples (Fig. 2a and Supplementary Fig. 4). The main difference was a modest increase in swaps for plasmid samples originating from homologous insert (blue squares in Fig. 2a, from 1–2% (direct) to 3–5% (PCR) discordant pairs), in line with the low level of chimerism known to originate from PCR23. This PCR chimerism was nevertheless about one order of magnitude lower than that observed in AAV-derived samples. Second, we also used an orthogonal long-read sequencing platform to document BC swaps. Slightly modifying a recent library preparation protocol20 based on a single Bst2 polymerase extension step tailored to minimize intermolecule annealing, we characterized encapsidated DNA libraries p151:mid-hom and p152:mid-nonhom from the same packaged samples as for the direct ONT. Despite a rapid snap cooling step, analysis of consensus-circular PacBio sequences revealed populations consistent with both ssAAV annealing, in addition to molecules originating from the expected Bst2 extension (‘scAAV-like’, Extended Data Fig. 5a–g). This categorization was further supported by inspection of the identities of BC1s and BC2s across the reads (Extended Data Fig. 5h–l): different BC1s/BC2s on forward/reverse CCS reads for the intermolecular ssAAV annealing and identical BC1s/BC2s for intramolecular extension (scAAV-like). As expected biochemically, the proportion of annealed molecules was higher for the fully homologous insert (p151:mid-hom) compared to the nonhomologous library (note that p152:mid-nonhom still has homology regions flanking the BC1–BC2 section; Fig. 1a). Stratifying bioinformatic analysis of BC1–BC2 concordance across the different molecular species, however, showed that the fraction of swaps was insensitive to the specific steps in the library preparation and aligned with the ONT results, with or without PCR (Fig. 2b). Third, a hand-mixing experiment in our companion work (figure 3f,g in ref. 14) revealed that the ONT procedure is likely not the originator of swaps; in a low-complexity sample composed of separately packaged plasmids pooled before ONT processing, chimerism was nearly inexistent (<2.5% swaps) in contrast to plasmids pooled before packaging (>35% swaps). Fourth, enhancer reporters with two separate barcoded RNA per constructs24 were captured in single-cell RNA sequencing from mouse brains in our companion work. Reporter barcodes molecules captured confirmed pervasive lack of codetection compared to expected pairing mapped from the originating plasmid reporter library (swapping incidentally confirmed by ONT sequencing; Supplementary Fig. 5). This constitutes a linkage measurement without any long-read quantification (supplementary figure 7g in ref. 14). Collectively, these data strongly support our interpretation that the observed chimerism is not introduced through technical aspects of the long-read library preparation.
a, Comparison of PCR versus direct long-read library preparation for ONT sequencing of paired barcode libraries. Left, schematic of the workflow. For plasmid DNA, we compare digested BC1–BC2 inserts versus PCR products. For AAV-packaged DNA, we either extracted DNA and performed PCR before long-read sample preparation or submitted the AAV particles directly (library preparation using the annealing strategy19). All ONT sequencing was performed with Plasmidsaurus. The graph shows the fraction of discordant barcode pairs from PCR-derived libraries (x axis) versus direct (same data as Fig. 1c; y axis), as also shown in Supplementary Fig. 4b. AAV samples only include the PHP.eB serotype with standard packaging conditions. Error bars correspond to the 20th–80th percentiles from bootstrap resampling to document read counting noise (smaller than symbol size for PCR libraries along x axis because of high sequencing coverage for these libraries; center: quantification from all reads). The AAV data (circles) are from one packaging replicate per library, with ONT preparation either through direct (annealing-based) or PCR-based approaches. The plasmid-derived data (squares) are from a single replicate (one preparation per library). b, Comparison of the fraction of discordant BC1–BC2 pairs from AAV-packaged DNA from the same libraries (p151:mid-hom in blue and p152:mid-nonhom in orange) across ONT (both direct and PCR-based) and PacBio libraries. Quantification on the PacBio reads was stratified by class of molecules (produced from intramolecular extension or through annealing), as also shown in Extended Data Fig. 5.
Nevertheless, four technical points deserve note. First, one of our libraries, p153:long-hom, while intended to exclusively harbor long homologous inserts, also contained a sizeable proportion of short and mid-sized homologous inserts in the AAV-packaged ONT data because of cloning history (short: 8–49%, mid-sized: 10–23%, across the four samples), which could be identified because of their internal insert index and shorter total read lengths (Supplementary Fig. 2e). Quantifications from the resulting multisized library, denoted as p153×:long-hom, are marked with × in Fig. 1c (all points in Extended Data Fig. 3). Shorter insert elements in p153×:long-hom were present at low proportion in the starting plasmid library (not detected in whole-plasmid sequencing verification and barely visible on gel but clearly visible upon PCR amplification; Supplementary Fig. 2g–i) but were likely selectively enriched because of their shorter sizes during rAAV packaging. All other libraries overwhelmingly comprised the expected inserts (>99% apart from generally low level of empty parental carryover; mid-sized and long nonhomologous displayed higher proportion of parental p146 sequences, at 19% and 66% respectively, possibly enriched by the annealing step of the ONT library preparation, as discussed below). To provide additional evidence for the phenomenon of chimerism on more constructs with long inserts, we quantified discordance on another orthogonally prepared, high-complexity library of dual-barcoded reporters25 with a rigorously mapped barcode-to-enhancer association dictionary (Supplementary Fig. 5a–d). These indeed showed a level of swapping dependent on insert size (9% and 45% discordance for the 220-bp and 1,450-bp homologous inserts respectively; Supplementary Fig. 5e). The different quantitative magnitude of the observed chimerism for different intervening sequences suggest subtleties likely related to the underlying causative mechanism. Prior work with overlapping AAV dual vectors indeed confirms different ‘recombination’ propensities across varied sequences26,27.
Second, AAV-packaged long inserts (p153×:long-hom and p154:long-nonhom) showed a high proportion of reads with shorter than expected BC-to-BC inserts (Supplementary Fig. 2d; 14–67% in p153×, 85% in p154). These off-products (not included in the quantification shown in Fig. 1c) were associated with even lower rates of barcode pair concordance, irrespective of the homologous or nonhomologous nature of their inserts (Supplementary Fig. 2e). Inserts from these shorter off-products displayed a high proportion of complex composite multisegment alignments (Supplementary Data 3; Fisher’s exact P < 10−4 in all instances), in line with previous evidence of complex structural variants in AAV-packaged DNA8,9.
Third, some of the reads, despite having the full BC-to-BC length, did not span the full ITR-to-ITR length. This phenomenon was seen not only in the AAV-packaged samples but also in the size-selected digested plasmid inserts (Supplementary Figs. 1f and 2e), which were overwhelmingly ~2.2 kb upon submission for long-read sequencing. Therefore, we tentatively attribute these to downstream technical aspects of the ONT sequencing. Consistently, these incomplete ITR-to-ITR reads (but still with full BC-to-BC lengths) had similar rates of barcode swapping (Fisher’s exact P > 0.35 in all instances; Supplementary Data 4).
Fourth, given that only a small proportion of all long reads satisfied our rigorous quality control steps to quantify BC1–BC2 discordance, we explored whether error correction of barcodes was a viable way to recoup the ~35% of reads lost because of requiring perfect matches (separately) to BC1s and BC2s in the starting dictionary. We found that only a minor proportion (<10% of total reads) of the nonmatched barcodes were within one Hamming distance of bona fide barcodes (the complexity of our libraries prevented us from extending to larger Hamming distances; >90% of reads accounted for after allowing for different error modes, Supplementary Fig. 3d). BC1–BC2 discordance on error-corrected barcodes was even higher than BC1–BC2 discordance with perfect matches. Given this minor proportion and to avoid the risk of introducing any cryptic biases, we continued to require perfect matches and did not include error-corrected barcodes in our quantification. Future iterations with longer barcode designs would enable more robust error correction. We did, however, confirm that sequencing errors did not lead to apparent but spurious BC1–BC2 swaps in our experiment (Supplementary Note 1). Beyond nonmatched barcodes, we more thoroughly documented the classes of subgenomic particles in our data. We found that a large proportion of the reads that failed to pass our quality control filters (for example, 29–81% for ONT and 22–25% for PacBio with zero of four mapped signposts) were partial fragments predominantly mapping near the 3′-most ITR within our constructs (Extended Data Fig. 4). These fragments contained no barcodes and, therefore, could not be used for chimerism quantification. Notably, a subgenomic population, predominantly identified in our PacBio data, consisted of ‘snapback’ molecules28,29 originating at the Tn5 mosaic ends (from the Nextera handles used to generate the barcoded inserts; Extended Data Fig. 5c,j). Comparing the repeated BC2s concordance in those fragments also indicated homology-dependent swaps and at a higher level than for BC1–BC2 pairs in the complete genomic reads (Extended Data Fig. 5j).
A critical aspect of in vivo functional genomics is high-fidelity delivery of functional payloads to cells and the scale of these experiments has been growing30. rAAV vectors have known limitations in terms of the size of their DNA payload; however, to our knowledge, high-frequency rearrangement of genetic content upon packaging has not been reported. Following early discovery from serial subgenomic infections31, recombination is a recognized driving force for AAV evolution32,33. As a notable example, the AAV-6 serotype is believed to be the recombination product of AAV1 and AAV2 (ref. 34). Analogous observations were made in the context of oversized cargo production for gene therapy, in which homologous subfragments delivered through distinct rAAVs are fused to heterodimers in host cells upon high-multiplicity-of-infection dual transfer35,36,37,38,39,40,41,42, putatively through homologous recombination. However, these observations have not been connected to the distinct context of pervasive chimerism among library entities during AAV packaging. This is unlike the well-established case of lentiviral vectors, which recombine because of template switching, thereby unlinking their genetic components6,7,43,44,45,46 in a length-dependent and homology-dependent manner. Our results document widespread chimerism in AAV as well, with a key distinction being that, for lentivirus, template switching occurs after transduction in infected cells, whereas, with AAV, chimerism occurs during packaging. This chimerism is likely a substantial contributor to the noise observed in multiplexed barcoded AAV packaging by ourselves and others10,11,13,47,48,49 (evidence of consequences for the interpretation of in vivo experiments provided in ref. 14).
While the precise mechanism of AAV chimera formation remains unresolved and beyond the scope of this brief communication, several features of the phenomenon—most notably its dependence on cargo length, sequence homology and input dose—provide important constraints. Our leading hypothesis is that chimeras arise from homology-dependent template switching associated with DNA repair pathways activated under stress because of AAV replication. AAV production is known to induce host DNA damage responses50,51 and polymerases implicated in AAV replication (Pol δ, η and κ) also function in DNA repair51,52. Replication stress during AAV genome amplification could, therefore, promote template switching following fork stalling or collapse, a documented outcome of stressed replication forks53.
Other mechanisms are also plausible. Chimeras could form through template switching of partially replicated genomes during rolling hairpin amplification54, analogous to events observed with poorly processive polymerases in PCR24,55,56. Alternatively, open double-stranded AAV intermediates may mimic double-strand breaks and undergo resection followed by repair through single-strand annealing or homologous recombination57. In addition, recombination between cotransfected plasmids cannot be formally excluded; however, the weak scaling of chimera frequency with input DNA dose (Fig. 1d) argues against this explanation, as plasmid–plasmid recombination is unlikely to occur at appreciable frequencies at very low DNA concentrations58.
Another intriguing possibility is that chimerism follows from the packaging of multiple rAAV genomes in the same capsid. Such multiply packaged rAAV capsids are possible for ITR-to-ITR lengths < 3 kb (such as our BC1–BC2 constructs)59. However, our doubly barcoded enhancer reporters are >3 kb (ITR inclusive, larger than half of the rAAV limit; Supplementary Fig. 5a) and we still observe a similar level of chimerism for these constructs (Supplementary Fig. 5b), suggesting that the phenomenon is not strongly related to multiply packaged capsids.
Mechanistic diversity of the AAV packaging process is well documented, be it with regard to genome configuration (ssAAV versus scAAV), serotypes, Rep proteins60 or helper proteins (for example, Ad versus HSV-1)54. A notable example is serotype AAV5, which is a phylogenetic outlier and displays substantial differences in packaging and encapsidation61,62,63,64. That said, here, we confirm pervasive swaps with both PHP.eB and AAV2 serotypes (Extended Data Fig. 3), which derive from distinct AAV clades64. Quantitatively evaluating how a wider range of serotypes and packaging conditions modulate the chimerism phenomenon should be straightforward using our BC1–BC2 reporters with minor modifications.
Until the definitive mechanism is identified, chimerism can still be partially be mitigated7,45,46 by limiting the distance between complex elements (for example, 5′ instead of 3′ barcodes in MPRA assays, as applied in lentiMPRA65,66), decreasing the extent of entirely homologous regions, lowering cotransfection dose in packaging cells when possible (at the expense of titer yield) or dispensing of the need for a barcode altogether where possible (for example, direct capture of sgRNAs67). Future technical improvements in packaging cofactors or investigation of involvement of endogenous DNA repair pathways could also find general solutions to this problem. Our results illuminate an issue in pooled rAAV production of complex libraries, which will inform experimental design decisions to improve data quality in multiplex projects involving this important gene delivery vehicle.
Methods
Cloning complex libraries of doubly barcoded AAV constructs with various inserts
To clone a set of doubly barcoded AAV constructs, we digested plasmid AiP11839 (Addgene, 163509) with PacI and BbsI (New England Biolabs) and size-selected the 2.9 kb backbone containing the AAV2 ITRs on agarose gel. A GFP stuffer constructs with complex barcodes was created by two steps of PCRs (step 1: primers o924 + o925 amplifying GFP from p027, step 2: o926 + o927_v2) using Kappa Robust and standard cycling conditions. Random nucleotides were included in the primers to append random DNA barcodes (15 nt in o926, left BC1 and 16 nt in o927_v2, right BC2; Extended Data Fig. 1d) The second PCR was tracked by qPCR and stopped at the inflection point to maintain complexity and limit jackpotting. The barcoded GFP stuffer was then size-selected on agarose and inserted by Gibson assembly (4-μl reaction) in the AiP11839 PacI + BbsI-digested backbone above. The resulting library was cleaned up (reaction taken to 50 μl with 10 mM Tris 8 buffer and Zymo Clean and Concentrator with 3:1 binding buffer, eluted in 6 μl of water) and 3 μl was electroporated in 25 μl of C3020 cells (New England Biolabs). The resulting complex library (>5 million barcode transformants estimated by plating 0.03%) was grown overnight at 37 °C and plasmid-purified (Qiagen miniprep), generating the parental barcoded plasmid p146 (Extended Data Fig. 1b). This barcoded ‘dock’ contained two SapI sites internal to the BC1 and BC2 to allow insertion of various libraries of inserts. Just outside the SapI sites, a Nextera read1 adaptor served as homology for Gibson assembly on the left BC1 side (Extended Data Fig. 1d) and another constant-homology arm was used on the right BC2 side. Before replacing GFP with different classes of inserts, we generated additional dock plasmids to accommodate final constructs of fixed lengths by adding constant inserts outside of the barcoded stuffer. Briefly, p146 was sequentially digested with BglII and PmlI (New England Biolabs), the resulting linear product size-selected on agarose. Filler sequences of 1,291 bp and 1,898 bp were generated by PCR amplification from plasmid AiP11839 (constant_1291bp with primers o929 + o930, constant_1898bp with primers o928 + o930) using standard conditions. Following size selection on agarose, these were inserted by Gibson assembly and electroporated as described above, maintaining a high complexity of represented barcodes in the libraries, resulting in barcoded plasmids p147 (with constant_1291bp) and p148 (with constant_1898bp). All parental libraries were confirmed by whole-plasmid sequencing (Plasmidsaurus).
Parental barcoded AAV libraries with GFP stuffer flanked by SapI sites: p146, no additional insert; p147, with additional constant 1,291-bp insert outside of barcodes; p148, with additional constant 1,898-bp insert outside of barcodes.
Six libraries were then constructed to vary the insert length and class (homologous, meaning all components of the library are identical, or nonhomologous, meaning that all members of the library are different). To maintain a roughly constant length of 2.3 kb between the AAV ITRs, short inserts were integrated in p148, mid-sized inserts were integrated in p147 and long inserts were integrated in p146 (Extended Data Fig. 1b,c). All parental plasmids were digested with SapI (New England Biolabs) to release the GFP stuffer and the resulting barcoded backbones were size-selected on agarose for downstream steps, described below.
Homologous (fixed) inserts were taken from sections of the AiP11839 cargo (Extended Data Fig. 1a) and generated by two steps of PCR with primers. The first step appended handles (Nextera R1 on left, partial Nextera R2 on right) to the constant region: homologous_127bp with primers o931 + o934, homologous_739bp with o931 + o933 and homologous_2034bp with o931 + o932. These handles then served to prime a secondary PCR, which also appended a unique library index inside the construct for later demultiplexing. This secondary PCR was the same as that used for the construction of the nonhomologous libraries, corresponding to primers Nextera R1 (o759) in the forward direction and an indexed Nextera R2 with the constant right homology arm in the reverse direction (homologous_127bp with o937, homologous_739bp with o938 and homologous_2034bp with o939). The indexed constant inserts were then size-selected on agarose and inserted in their respective (for fixed ITR-to-ITR length) SapI-digested barcoded parental backbone (Extended Data Fig. 1b) with Gibson assembly. Following cleanup and electroporation, libraries were bottlenecked by serial dilution before outgrowth to a target complexity of ~20,000 transformants.
To generate nonhomologous insert libraries, we relied on tagmentation and PCR amplification of bacterial genomic DNA. Briefly, 1 μl at 10 ng μl−1 of gDNA extracted from E. coli cells (also containing plasmid p146, as described below) was tagmented with dually loaded Tn5 (Illumina, Nextera Tagment DNA enzyme, 15027916) at two doses: 0.4 μl of Tn5 enzyme 1 and 0.4 μl of a 20-fold dilution of the Tn5 enzyme together with 3.6 μl of water and 5 μl of 2× tagmentation buffer (Illumina, 15027866). Following cleanup (Zymo Clean and Concentrator, 3:1 binding buffer), 1 μl of the 10-µl Tris 8 10 mM elution (1 ng) was taken as input for 12 cycles of PCR (Kappa Robust) from the Nextera handles with indexed primer (same primers series as homologous inserts above, forward o759, reverse: short with o941, mid-sized with o942 and long with o943) to mark the libraries with internal insert indices for downstream demultiplexing. The resulting smear was size-selected to a narrow range in size on polyacrylamide gel for the short insert and on agarose for the mid-sized and long inserts. In all cases, to size-match nonhomologous fragments as carefully as possible, the homologous inserts of corresponding length were run on side lanes on the gels and the small corresponding range of the amplified tagmented gDNA was cut out and purified. The size-selected fragments were secondarily amplified with the same primers to generate more material for cloning, size-selected again and inserted by Gibson assembly in their respective SapI-digested barcoded parental backbone as for the homologous fragments. Following cleanup and electroporation, libraries were again bottlenecked to a target complexity of ~20,000 transformants.
Thus, all in all, we obtained the following six libraries fixed ITR-to-ITR length with the following insert characteristics and estimated complexity from transformant counts. All homologous libraries were confirmed by whole-plasmid sequencing (Plasmidsaurus) and nonhomologous libraries were spot-checked with Sanger sequencing of colonies (Genewiz).
Final dual-barcoded AAV libraries with various inserts (listed insert lengths do not include the Tn5 Nextera handles, included between all barcode pairs: 33 + 34 bp total): p149, short (127 bp) homologous insert; p150, short (~100 to 150 bp) nonhomologous inserts; p151, mid-sized (739 bp) homologous insert; p152, mid-sized (~650 to 850 bp) nonhomologous inserts; p153, long (2,034 bp) homologous insert; p154, long (~1,900 to 2,100 bp) nonhomologous inserts.
We note that the bacterial pellet used for genomic DNA extraction to tagment for nonhomologous library insert generation was outgrown from a colony on a p146 transformation plate (but grown on LB without ampicillin). As such, inserts from these libraries contained at substantial proportion sequences from plasmid p146 (proportion of mapped fragments: 65% for p150, 19% for p152 and 15% for p154; inserts mapping to p146 also mapped to AiP11839 given similarity). While inserts from these nonhomologous libraries are still very diverse, given the limited size of the plasmid and substantial proportions of the libraries, there are still nonzero pockets of homology for certain members of the libraries. To quantify this, we performed local pairwise alignment (pairwiseAlignment from R package Biostrings, version 2.62.0, option type = ‘local’) of randomly selected insert sequences from the size-selected digested plasmid long reads (pairs of reads with distinct barcodes). Setting a threshold score per library as the maximum of 1,000 alignments from a pair of insert sequences and another one-shuffled pair (FDR < 0.001), we quantified that about 1% of inserts had detectable homology (0.8% p150, 1% p152 and 1.8% p154), indeed close with theoretical expectation, assuming even random fragmentation with the proportion of reads within each library coming from the plasmid (size 5 kb) and the size of inserts (p150: 0.65 × 0.65 × (100 bp/5,000 bp) = 0.8%, p152 = 0.19 × 0.19 × (750 bp/5,000 bp) = 0.5%, p154 = 0.15 × 0.15 × (4,000 bp/5,000 bp) = 1.8%). Hence, despite the tagmentation material in the starting library being a mixture of genome and plasmid, the final libraries were effectively nearly completely nonhomologous.
Notably, a significantly enriched proportion of the rare swapped-barcode reads from the nonhomologous libraries originated from members of the library with regions of homology. For instance, among the 22/26 full BC-to-BC length barcode-swapped reads from library p150 for which the two corresponding preswap inserts could be mapped in the size-selected digested plasmid data, 3/22 had extensive homology (pairwise local alignment score > 50), which was over tenfold higher than randomly selected pairs of inserts from the same library (Fisher’s exact P < 0.005).
Generation of a valid BC1–BC2 pair dictionary from parental plasmid library p146
To generate the dictionary of valid barcode 1 and barcode 2 pairs, we amplified by PCR (primers o945 + o946v2 containing P5 and P7 Illumina adaptors) the GFP stuffer insert flanked by the two barcodes using 5 ng of starting template (50-μl reaction, Kapa Robust, standard conditions, ten cycles) and followed by 1× AMPure beads cleanup. The resulting library was sequenced using custom primers as a fraction of a NextSeq2000 P2 100-cycle run (read1: 42 cycles with o947, index1: 20 cycles with o761 Nextera_read1 (into SapI restriction site and GFP, not used), index2: 20 cycles with o948 (into GFP, not used), read2: 16 cycles with o762 Nextera_index2). Sequencing data were demultiplexed from other samples on the basis of the first ten indices of read1 (GATCCGTCGA) using bcl2fastq with base mask i10y*,y*,y*,y*, yielding 195.6 million reads to associate barcodes from p146. BC1 (5′/left of insert) was on cycles 1–16 within read2, whereas BC2 (3′/right of insert) was on cycles 21 to 36 within read1.
We then applied stringent representation and uniqueness criteria to identify unique valid pairs for our downstream swapping assessment. Read counts corresponding to identical barcodes 1 and 2 were first piled up. First, representation of barcodes (separately, summing reads from pairs with three or more counts) was inspected (Extended Data Fig. 2c), revealing a trimodal distribution: low-count barcodes (≤5 reads) corresponding to likely sequencing or PCR errors (BC1, n = 1,010,349, 2.5% of reads; BC2: n = 1,162,035, 2.9% of reads), intermediate-count barcodes making up the bulk of the coverage (BC1, n = 6,345,673 barcodes, 92.9% of reads; BC2, n = 6,596,695 barcodes, 93.5% of reads) and high-count barcodes (BC1, n = 827, 4.6% of reads; BC2, n = 530 barcodes, 3.7% of reads), possibly emerging from of clonal expansion during transformation outgrowth and/or PCR jackpotting. In the case of BC2, a single sequence (ATAACGACTTGTGAGC) was drastically overrepresented (2.5% of reads). This barcode and all other barcodes within a Levenshtein distance of ≤2 to it (n = 402 barcode, 0.25% of reads) were not considered in downstream analysis to avoid spurious nonunique pairs. We note that our barcode space was not saturated at that level of tolerance to mismatches (average of three BCs within our BC2 reads within Levenshtein distance of 2 to repeated one-shufflings of the ATAACGACTTGTGAGC sequence). Furthermore, barcodes with ten or more consecutive Gs or containing truncated BC (with the detected post-BC sequences: ATTAAAC for BC1, TAGCGCG for BC2) were removed, corresponding to a minute proportion of the library (BC1, n = 9,533, 1.1% of reads; BC2, n = 3,910, 0.06% of reads). To avoid mismatches from high-representation barcodes being retained as spurious mid-count barcodes, the list of mid-count barcodes was pruned by removing those within a Levenshtein distance of 2 from the high-count barcodes (number of mid-count barcodes removed in pruning process: BC1, n = 16,386; BC2, n = 4,747). All remaining pruned mid-count barcodes were retained downstream (BC1, n = 6,321,002; BC2, n = 6,588,391). Lastly, the high-count barcodes were further error-corrected by generating an undirected graph connecting barcodes within a Levenshtein distance of 2 or less. The most highly represented barcodes from each connected component were considered valid. Rare clusters composed of many well-represented barcodes (fold change between maximum and minimum < 10 within connected component) were discarded as possibly ambiguous, leading to n = 609 and n = 420 error-corrected high-count BC1 and BC2 sets, respectively. All in all, these filtering steps generated a list of well-represented high-quality barcodes (BC1, n = 6,321,611, BC2: n = 6,588,811).
From these well-represented BC1 and BC2 sets, we finally filtered unique pairings between the two. Specifically, all paired barcodes with ≥2 reads were filtered for members present in both separately valid sets. Then, the proportion of reads to each barcode within a pair (for example, the number of reads to BC1 from a given pair over number of reads containing the same BC1 across all pairs in the library) was computed. Only pairs for which >99% of reads mapped to a unique pair were retained. This led to a final set of n = 5,586,772 valid barcode pairs used for downstream assessment. Only exact matches to BC1 and BC2 constituents of these final pairs were used for filtering long-read data and setting the denominator in our barcode swap quantification. The retention proportion at filtering steps is presented in Extended Data Fig. 2b.
We note that the distinction between mid-count and high-count representation above was largely immaterial, as the overwhelming majority of detected BCs in our long-read libraries were from the mid-count set (n = 12 of 4,811 full BC-to-BC reads from AAV used to quantify swaps in Fig. 1 from the high-count barcode set), in line with their dominant representation, with no significant correlation with concordant or discordant pairing and barcode representation class (for the two libraries with detected high-count barcodes in the final read list, Fisher’s exact test: p149, P = 0.71; p151, P = 0.14).
AAV packaging
Complex libraries were packaged into PHP.eB or AAV2 capsids using the crude prep method previously described68. Maxiprep libraries cloned between AAV2 ITRs were transfected with PEI Max 40K (Polysciences, 24765-1) into one 15-cm plate of HEK-293T cells (American Type Culture Collection, CRL-11268), along with helper plasmid pHelper (Cell BioLabs) and either pUCmini-iCAP-PHP.eB69 (Addgene, 103005) or pAAV2/2 (Addgene, 104963). The final transfection mix contained 150 μg of PEI Max 40K, 30 μg of pHelper DNA, 15 μg of rep/cap plasmid DNA and 15 μg of library DNA per plate (‘standard conditions’). In the case of ‘sparse conditions’, we transfected with fewer library molecules per cell by using 10% (1.5 μg) library DNA along with 90% (13.5 μg) non-ITR-bearing empty expression vector plasmid DNA as a carrier (AiP12481, pCDNA3.1-CMV-empty-IRES2-mTFP1-BGHpA). Following transfection at 24 h, the medium was changed to low-serum conditions (1% FBS) and then, after 5 days, cells and supernatant were harvested into 50-ml conical tubes and AAV particles were released by three freeze–thaw cycles. The cell lysates were then treated with Benzonase to degrade free DNA (2 μl of Benzonase, 30 min at 37 °C, MilliporeSigma, E8263-25KU) and then cell debris was cleared with a low-speed spin (1,500g 10 min). The supernatant containing virus was concentrated over a Centricon column (100-kDa molecular weight cutoff; MilliporeSigma, Z648043) to a final volume of ∼100 μl, containing ~1 × 1012–3 × 1012 vector genomes. Crude AAVs were used for direct sequencing (Plasmidsaurus AAV sequencing service).
Sample preparation and sample submission for long-read sequencing
We long-read sequenced both direct and PCR-based libraries. For PCR-free sequencing of plasmid DNA, we digested the starting dually barcoded AAV plasmid libraries p146 and p149–p154 with NotI-HF and MluI-HF (New England Biolabs, using 2 μg of starting material, 37 °C, 1 h). The released the barcoded inserts (881 bp for p146, ~2.3 kb for p149–p154) were then size-selected on agarose (Zymoclean gel purification), pooled and submitted to Plasmidsaurus for a custom long-read project (project 8Y6YSQ.1; target: 3 million reads, recovery: 2.6 million reads). PCR-free libraries of AAV-packaged DNA were sequenced using the AAV service from Plasmidsaurus (project RP88L6).
To generate libraries of barcoded insert by PCR from the AAV-packaged DNA, we first extracted DNA by performing proteinase K treatment (3 μl of crude AAV prep, 6 μl of 10 mM Tris 8, 1 μl of proteinase K (Thermo Scientific, EO0491), 60 min at 50 °C, 5 min at 70 °C and then placed on ice), followed by phenol–chloroform extraction (adding 190 μl of 10 mM Tris 8, adding 200 μl of phenol–chloroform–isoamyl alcohol (Invitrogen, 15593-031), vortexing for 30 s, spinning at 16,000g at room temperature for 5 min and taking aqueous layer) and isopropanol precipitation (adding 1 μl of glycoblue, 50 μl of sodium acetate 3 M and 250 μl of isopropanol 100%, vortexing for 45 min at −80 °C and 45 min at 21,000g at 4 °C, washing with 80% ethanol, air-drying the pellet and resuspending in 10 μl of 10 mM Tris 8). Next, 2 μl of the precipitated DNA was taken as template for PCR with primers oJBL949 + oJBL950 (Kapa Robust HotStart Ready mix (Roche), annealing temperature: 60 °C, elongation time: 2 min 30 s) and tracked by qPCR (SYBr green). PCR libraries were generated from all samples packaged with the PHP.eB serotype in standard (nonsparse) packaging conditions (that is, samples 1–6; Supplementary Fig. 2f). Two separate reactions per sample (reaction 1: volume 20 μl, 16 cycles; reaction 2: volume 50 μl, 17–20 cycles) were pooled before submission. Of note, we tested the importance of adding a DNase treatment step by performing qPCR with primers targeting the insert between the AAV ITRs (oJBL905 + oJBL906) versus a sequence on the ampicillin cassette in the backbone (oJBL097 + oJBL098) comparing with and without DNase treatment before proteinase K treatment but saw no difference (>30-fold lower backbone material relative to cargo), likely because of the Benzonase treatment already having degraded the nonencapsidated DNA. PCR libraries from plasmids were prepared from 1 ng of template (with quantification possibly confounded by genomic DNA carryover) and reactions were stopped at 18 cycles (starting input material calibrated to still be in exponential phase before the inflection point). Plasmid-templated samples also included the parental p146 (which was pooled at 0.1-fold stoichiometry of other larger inserts to mitigate ONT size biases). In both plasmid-templated and AAV-templated conditions, PCR reactions were cleaned up with 0.4× AMPure clean, leaving predominantly >2-kb products (that is, largely excluding p153×-short and p153×-mid samples; Supplementary Fig. 4g). Two samples corresponding to pooled products (AAV pool sample 1, plasmid pool sample 2) were submitted to Plasmidsaurus for long-read sequencing (custom project DXYB6C, target reads: 1 million per sample).
Detailed methods on the computational pipeline to process the long-read data are provided in the Supplementary Methods.
PacBio AAV library preparation and sequencing
To generate libraries derived from AAV encapsidated DNA for PacBio sequencing, we adapted the protocol from Zhang et al.21 to reduce the number of gel extractions to improve yield and possibly reduce biases in subgenomic fragments. First, encapsidated DNA was purified using the Purelink Viral RNA/DNA mini kit (Invitrogen). Briefly, 50 μl of crude viral preparation (approximately 7 × 1011 viral genomes) was treated with 20 U of DNase 1 (Thermo, PI89836) at 37 °C for 30 min in a 200-μl total reaction. The DNase-treated encapsidated DNA was then treated for 15 min at 56 °C with 25 μl of proteinase K (Thermo, 4333793) in a total reaction volume of 425 μl (reaction including 5.6 μg of carrier RNA). The DNA was then purified using the column following the instructions and eluted in 50 μl. Polymerase Bst2 (New England Biolabs) was then used to perform double-stranding primed by the 3′ end of the AAV ITR. Specifically, 20 μl of purified DNA was mixed with 10 μl of 10x buffer, 6 μl of 100 mM MgSO4 and 45 μl of water, heated for 5 min at 95 °C and then placed on ice for 10 min. Then, 14 μl of 10 mM dNTPs were added, together with 4 μl of Bst2 polymerase and 1 μl of BSA (10 mg ml−1), and the mixture was incubated at 50 °C for 60 min. For PacBio library preparation, the SMRTbell prep kit 3.0 was used. Specifically, the Bst2 reaction was cleaned up with SMRTbell cleanup beads at 1.3× and eluted in 46 μl. After extraction and double-stranding, concentration was assessed by Qbit and integrity spot-checked with the Agilent TapeStation. End repair and poly(A) tailing was performed by adding 14 μl of the repair master mix (8 μl of repair buffer, 4 μl of end repair mix, 2 μl of DNA repair mix) and treated at 37 °C for 30 min and 65 °C for 5 min, followed by a hold at 4 °C. Next, 4 μl of SMRTbell barcoded adaptprs 3.0 were then added to the reaction from the previous step, together with 31 μl of ligation master mix (30 μl of ligation mix, 1 μl of ligation enhancer) and incubated for 30 min at 20 °C, followed by a hold at 4 °C. The resulting ligated samples were cleaned with 1.3× SMRTbell cleanup beads and eluted in 40 μl. The ligated DNA was then treated with nuclease by adding 10 μl of master mix (5 μl of nuclease buffer, 5 μl of nuclease mix) and incubated for 15 min at 37 °C, followed by a hold at 4 °C. The sample was then purified with 1.3× SMRTbell cleanup beads and eluted in 12 μl. Libraries were pooled and loaded on the PacBio Vega at 0.25 ng μl−1 (loading at a higher concentration would have increased the number of reads). The sequencing was run with application type ‘viral sequencing/AAV’, leading to modified adaptor calling (to allow for the capture of scAAV-like molecules).
Detailed methods on the computational pipeline to process the long-read data are provided in the Supplementary Methods. The associated pseudocode is provided in Supplementary Note 2.
Input plasmid dosage experiment
To assess the importance of input rAAV plasmid dosage on the chimerism phenomenon, we packaged the library p151:mid-hom as described above, but at four different doses per 15-cm plate of producing cells: 15 μg, 1.5 μg, 150 ng and 15 ng. The amount of DNA transfected was kept constant by compensating with the same ITR-free carrier plasmid (AiP12481, pCDNA3.1-CMV-empty-IRES2-mTFP1-BGHpA) as before. As a control, p152:mid-nonhom was also packaged a second time at the 15-μg dose. Analysis of the ONT data proceeded as described above with the same quality control filters. Metrics for these data are presented in Supplementary Fig. 3.
Statistical testing (bootstrap FDR)
To provide estimates of significance from counting noise (some AAV samples had relatively few reads passing quality control filters; Supplementary Fig. 2f), bootstrap resampling was performed to generate ensemble estimates of concordant barcode pairs. To compare two samples (for example, p149:AAV versus p150:AAV in Fig. 1c), n = 105 bootstrap resamplings were performed. In this case, the bootstrap FDR was taken as the fraction of resamplings in which sample p149:AAV had a higher bootstrap concordant pair fraction than the p150:AAV resampling, etc.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All scripts used to analyze the data, plasmid maps and amplicon files (for p146 BC pair dictionary generation and AAV-scQer constructs) are available from Zenodo (https://doi.org/10.5281/zenodo.14515776)70. Raw short-read (p146 barcode dictionary generation, AAV-scQer oBC-CRE-mBC dictionary generation) and long-read (ONT and PacBio for barcode swap assessment) sequencing data are available from the Gene Expression Omnibus (GEO) under accession code GSE284548. A list of processed files available on GEO with a brief description is provided in the Supplementary Information. Source data are provided with this paper.
Code availability
Custom code used in analyzing the data in this study is available from Zenodo (https://doi.org/10.5281/zenodo.14515776)70.
References
Adamson, B. et al. A multiplexed single-cell CRISPR screening platform enables systematic dissection of the unfolded protein response. Cell 167, 1867–1882 (2016).
Dixit, A. et al. Perturb-Seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 (2016).
Patwardhan, R. P. et al. High-resolution analysis of DNA regulatory elements by synthetic saturation mutagenesis. Nat. Biotechnol. 27, 1173–1175 (2009).
Yu, H., Jetzt, A. E., Ron, Y., Preston, B. D. & Dougherty, J. P. The nature of human immunodeficiency virus type 1 strand transfers. J. Biol. Chem. 273, 28384–28391 (1998).
Rhodes, T., Wargo, H. & Hu, W.-S. High rates of human immunodeficiency virus type 1 recombination: near-random segregation of markers one kilobase apart in one round of viral replication. J. Virol. 77, 11193–11200 (2003).
Jetzt, A. E. et al. High rate of recombination throughout the human immunodeficiency virus type 1 genome. J Virol 74, 1234–1240 (2000).
Hill, A. J. et al. On the design of CRISPR-based single-cell molecular screens. Nat. Methods 15, 271–274 (2018).
Tran, N. T. et al. AAV-genome population sequencing of vectors packaging CRISPR components reveals design-influenced heterogeneity. Mol. Ther. Methods Clin. Dev. 18, 639–651 (2020).
Tai, P. W. L. et al. Adeno-associated virus genome population sequencing achieves full vector genome resolution and reveals human–vector chimeras. Mol. Ther. Methods Clin. Dev. 9, 130–141 (2018).
Shen, S. Q. et al. Massively parallel cis-regulatory analysis in the mammalian central nervous system. Genome Res. 26, 238–255 (2016).
Hrvatin, S. et al. A scalable platform for the development of cell-type-specific viral drivers. eLife 8, e48089 (2019).
McAfee, J. C. et al. Systematic investigation of allelic regulatory activity of schizophrenia-associated common variants. Cell Genom. 3, 100404 (2023).
Nguyen, T. A. et al. High-throughput functional comparison of promoter and enhancer activities. Genome Res. 26, 1023–1033 (2016).
Hunker, A. C. et al. Technical and biological sources of noise confound multiplexed enhancer AAV screening. Nat. Commun. https://doi.org/10.1038/s41467-026-72147-8 (2026).
Schmit, P. F. et al. Cross-packaging and capsid mosaic formation in multiplexed AAV libraries. Mol. Ther. Methods Clin. Dev. 17, 107–121 (2020).
Nonnenmacher, M., van Bakel, H., Hajjar, R. J. & Weber, T. High capsid-genome correlation facilitates creation of AAV libraries for directed evolution. Mol. Ther. 23, 675–682 (2015).
Tabebordbar, M. et al. Directed evolution of a family of AAV capsid variants enabling potent muscle-directed gene delivery across species. Cell 184, 4919–4938 (2021).
Deverman, B. E. et al. Cre-dependent selection yields AAV variants for widespread gene transfer to the adult brain. Nat. Biotechnol. 34, 204–209 (2016).
Namkung, S. et al. Direct ITR-to-ITR nanopore sequencing of AAV vector genomes. Hum. Gene Ther. 33, 1187–1196 (2022).
King, J. A., Dubielzig, R., Grimm, D. & Kleinschmidt, J. A. DNA helicase-mediated packaging of adeno-associated virus type 2 genomes into preformed capsids. EMBO J. 20, 3282–3291 (2001).
Zhang, J. et al. Thorough molecular configuration analysis of noncanonical AAV genomes in AAV vector preparations. Mol. Ther. Methods Clin. Dev. 32, 101215 (2024).
Zhang, J. et al. Subgenomic particles in rAAV vectors result from DNA lesion/break and non-homologous end joining of vector genomes. Mol. Ther. Nucleic Acids 29, 852–861 (2022).
McColl-Carboni, A. et al. Analytical characterization of full, intermediate, and empty AAV capsids. Gene Ther. 31, 285–294 (2024).
Hegde, M., Strand, C., Hanna, R. E. & Doench, J. G. Uncoupling of sgRNAs from their associated barcodes during PCR amplification of combinatorial CRISPR screens. PLoS ONE 13, e0197547 (2018).
Lalanne, J.-B. et al. Multiplex profiling of developmental cis-regulatory elements with quantitative single-cell expression reporters. Nat. Methods 21, 983–993 (2024).
Ghosh, A., Yue, Y. & Duan, D. Efficient transgene reconstitution with hybrid dual AAV vectors carrying the minimized bridging sequences. Hum. Gene Ther. 22, 77–83 (2011).
Ghosh, A., Yue, Y. & Duan, D. Viral serotype and the transgene sequence influence overlapping adeno-associated viral (AAV) vector-mediated gene transfer in skeletal muscle. J. Gene Med. 8, 298–305 (2006).
Xie, J. et al. Short DNA hairpins compromise recombinant adeno-associated virus genome homogeneity. Mol. Ther. 25, 1363–1374 (2017).
Xie, J. et al. Effective and accurate gene silencing by a recombinant AAV-compatible microRNA scaffold. Mol. Ther. 28, 422–430 (2020).
Eisenstein, M. Biotechs take multiplexing into animal models to accelerate drug discovery. Nat. Biotechnol. 42, 1162–1164 (2024).
Senapathy, P. & Carter, B. J. Molecular cloning of adeno-associated virus variant genomes and generation of infectious virus by recombination in mammalian cells. J. Biol. Chem. 259, 4661–4666 (1984).
Gao, G. et al. Adeno-associated viruses undergo substantial evolution in primates during natural infections. Proc. Natl Acad. Sci. USA 100, 6081–6086 (2003).
Shackelton, L. A., Hoelzer, K., Parrish, C. R. & Holmes, E. C. Comparative analysis reveals frequent recombination in the parvoviruses. J. Gen. Virol. 88, 3294–3301 (2007).
Xiao, W. et al. Gene therapy vectors based on adeno-associated virus type 1. J. Virol. 73, 3994–4003 (1999).
Sondergaard, P. C. et al. AAV.Dysferlin overlap vectors restore function in dysferlinopathy animal models. Ann. Clin. Transl. Neurol. 2, 256–270 (2015).
Grose, W. E. et al. Homologous recombination mediates functional recovery of dysferlin deficiency following AAV5 gene transfer. PLoS ONE 7, e39233 (2012).
Duan, D., Yue, Y. & Engelhardt, J. F. Expanding AAV packaging capacity with trans-splicing or overlapping vectors: a quantitative comparison. Mol. Ther. 4, 383–391 (2001).
Wu, Z., Yang, H. & Colosi, P. Effect of genome size on AAV vector packaging. Mol. Ther. 18, 80–86 (2010).
Yang, J. et al. Concatamerization of adeno-associated virus circular genomes occurs through intermolecular recombination. J. Virol. 73, 9468–9477 (1999).
Dong, B., Nakai, H. & Xiao, W. Characterization of genome integrity for oversized recombinant AAV vector. Mol. Ther. 18, 87–92 (2010).
Sun, L., Li, J. & Xiao, X. Overcoming adeno-associated virus vector size limitation through viral DNA heterodimerization. Nat. Med. 6, 599–602 (2000).
Nakai, H., Storm, T. A. & Kay, M. A. Increasing the size of rAAV-mediated expression cassettes in vivo by intermolecular joining of two complementary vectors. Nat. Biotechnol. 18, 527–532 (2000).
Sack, L. M., Davoli, T., Xu, Q., Li, M. Z. & Elledge, S. J. Sources of error in mammalian genetic screens. G3 (Bethesda) 6, 2781–2790 (2016).
Schlub, T. E., Smyth, R. P., Grimm, A. J., Mak, J. & Davenport, M. P. Accurately measuring recombination between closely related HIV-1 genomes. PLoS Comput. Biol. 6, e1000766 (2010).
Feldman, D., Singh, A., Garrity, A. J. & Blainey, P. C. Lentiviral co-packaging mitigates the effects of intermolecular recombination and multiple integrations in pooled genetic screens. Preprint at bioRxiv https://doi.org/10.1101/262121 (2018)
Adamson, B., Norman, T. M., Jost, M. & Weissman, J. S. Approaches to maximize sgRNA-barcode coupling in Perturb-seq screens. Preprint at bioRxiv https://doi.org/10.1101/298349 (2018).
Öztürk, B. E. et al. scAAVengr, a transcriptome-based pipeline for quantitative ranking of engineered AAVs with single-cell resolution. eLife 10, e64175 (2021).
Brown, D. et al. Deep parallel characterization of AAV tropism and AAV-mediated transcriptional changes single-cell RNA sequencing. Front. Immunol. 12, 730825 (2021).
Coughlin, G. M. et al. Spatial genomics of AAVs reveals mechanism of transcriptional crosstalk that enables targeted delivery of large genetic cargo. Nat. Biotechnol. 44, 133–145 (2026).
Schwartz, R. A., Carson, C. T., Schuberth, C. & Weitzman, M. D. Adeno-associated virus replication induces a DNA damage response coordinated by DNA-dependent protein kinase. J. Virol. 83, 6269–6278 (2009).
Ning, K. et al. Adeno-associated virus monoinfection induces a DNA damage response and DNA repair that contributes to viral DNA replication. mBio 14, e0352822 (2023).
Nash, K., Chen, W., McDonald, W. F., Zhou, X. & Muzyczka, N. Purification of host cell enzymes involved in adeno-associated virus DNA replication. J. Virol. 81, 5777–5787 (2007).
Branzei, D. & Foiani, M. Template switching: from replication fork repair to genome rearrangements. Cell 131, 1228–1230 (2007).
Lkharrazi, A. et al. AAV2 can replicate its DNA by a rolling hairpin or rolling circle mechanism, depending on the helper virus. J. Virol. 98, e0128224 (2024).
Qiu, X. et al. Evaluation of PCR-generated chimeras, mutations and heteroduplexes with 16S rRNA gene-based cloning. Appl. Environ. Microbiol. 67, 880–887 (2001).
Smyth, R. P. et al. Reducing chimera formation during PCR amplification to ensure accurate genotyping. Gene 469, 45–51 (2010).
McClements, M. E. & MacLaren, R. E. Adeno-associated virus (AAV) dual vector strategies for gene therapy encoding large transgenes. Yale J. Biol. Med. 90, 611–623 (2017).
Folger, K. R., Wong, E. A., Wahl, G. & Capecchi, M. R. Patterns of integration of DNA microinjected into cultured mammalian cells: evidence for homologous recombination between injected plasmid DNA molecules. Mol. Cell. Biol. 2, 1372–1387 (1982).
Dong, J. Y., Fan, P. D. & Frizzell, R. A. Quantitative analysis of the packaging capacity of recombinant adeno-associated virus. Hum. Gene Ther. 7, 2101–2112 (1996).
Mietzsch, M. et al. Improved genome packaging efficiency of adeno-associated virus vectors using Rep hybrids. J. Virol. 95, e0077321 (2021).
Chiorini, J. A., Afione, S. & Kotin, R. M. Adeno-associated virus (AAV) type 5 Rep protein cleaves a unique terminal resolution site compared with other AAV serotypes. J. Virol. 73, 4293–4298 (1999).
Earley, L. F. et al. Adeno-associated virus (AAV) assembly-activating protein is not an essential requirement for capsid assembly of AAV serotypes 4, 5, and 11. J. Virol. 91, e01980-16 (2017).
Fasina, O. & Pintel, D. J. The adeno-associated virus type 5 small rep proteins expressed via internal translation initiation are functional. J. Virol. 87, 296–303 (2013).
Gao, G. et al. Clades of adeno-associated viruses are widely disseminated in human tissues. J. Virol. 78, 6381–6388 (2004).
Klein, J. C. et al. A systematic evaluation of the design and context dependencies of massively parallel reporter assays. Nat. Methods 17, 1083–1091 (2020).
Gordon, M. G. et al. lentiMPRA and MPRAflow for high-throughput functional characterization of gene regulatory elements. Nat. Protoc. 15, 2387–2412 (2020).
Santinha, A. J. et al. Transcriptional linkage analysis with in vivo AAV-Perturb-seq. Nature 622, 367–375 (2023).
Mich, J. K. et al. Functional enhancer elements drive subclass-selective expression from mouse to primate neocortex. Cell Rep. 34, 108754 (2021).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Lalanne, J.-B., & Huynh, C. AAV chimerism, computational scripts, plasmid/amplicon maps, and alignment files. Zenodo https://doi.org/10.5281/zenodo.18841634 (2026).
Acknowledgements
We thank N. Donadio, R. Kutsal, S. Khem and S. Yao for AAV packaging support at the Allen Institute, N. Kamps-Hughes (Plasmidsaurus) for technical comments and the entire J.S. lab for discussion. We acknowledge L. F. Earley, M. Kosicki and D. Durocher for insightful input regarding possible underlying mechanisms. We thank M. Gasperini for critical reading of the manuscript. This work was supported by the National Institutes of Health (NIH; R01HG010632 to J.S.). J.-B.L. was supported by the Damon Runyon Foundation (DRG-2435-21) and by a Next Generation Scientist award from the Cancer Research Society of Canada (grant no. 1155581). T.A.M. was supported by a Banting Postdoctoral Fellowship from the Natural Sciences and Engineering Research Council of Canada. H.K. is a Washington Research Foundation Postdoctoral Fellow. J.S. is an Investigator of the Howard Hughes Medical Institute. The Allen Institute authors wish to thank the Paul G. Allen Family Foundation and NIH BRAIN Initiative Armamentarium Grant UF1MH128339 (to J.T.T. and B.P.L.) for their support.
Author information
Authors and Affiliations
Contributions
J.-B.L., J.K.M. and A.C.H. conceptualized the barcode swap experiment with input from T.A.M. J.-B.L. and C.H. cloned the libraries. C.H. prepared the PacBio libraries. H.K. provided expert assistance with the PacBio libraries and instrument. J.-B.L. analyzed the data. J.-B.L., J.K.M. and J.S. wrote the manuscript. B.P.L., J.T.T. and J.S. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
J.S. is a scientific advisory board member, consultant and/or cofounder of Prime Medicine, Guardant Health, Camp4 Therapeutics, Phase Genomics, Adaptive Biotechnologies, Sixth Street Capital, Pacific Biosciences, Somite AI and 10x Genomics. J.K.M. and B.P.L. are founders of EpiCure Therapeutics. The other authors declare no competing interests.
Peer review
Peer review information
Nature Biotechnology thanks John Doench, Florian Schmidt and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Barcode pairs AAV constructs and cloning strategy.
(a) Schematic of components of plasmid AiP11839 (Addgene#163509) between the AAV2 ITRs. Segments under the map highlight constant regions (cloned by PCR) used both as homologous library inserts (blue) and filler sequences to fix the ITR-to-ITR length (white). These segments are shown panel c below. Positions of PacI and BbsI restriction sites are marked by green carets, NotI and MluI sites by pale blue carets. (b) Molecular cloning scheme used. First complex BC1-BC2 parental dock with GFP stuffer p146 was cloned, and served as template for cloning secondary docks p147 and p148 with filler sequences. Complex library p146 was used as template for barcode dictionary generation. All parental docks were digested with SapI to liberate the GFP, which was replaced by respective internally indexed inserts to generate the final series p149-p154 (which were all bottlenecked to a target of 20k transformants). (c) At scale schematics of ITR-to-ITR components of cloned parental and insert-containing libraries. Position of SapI restriction sites are marked by pale red carets, BglII and PmlI sites by grey carets. (d) Sequences surrounding the two barcodes highlighting alignment signposts (yellow) used to create local position reference frames around barcodes in the long-read data. The ‘Insert index’ is shown as Ns, but is fixed and different for each type of insert (not degenerate). Constant Nextera handles between the barcodes (which constitute short homologous regions even in the non-homologous libraries) are shown.
Extended Data Fig. 2 Barcode pairs dictionary generation with paired-end short read sequencing.
(a) Schematic of procedure to generate the BC1-BC2 pairs dictionary. Parental plasmid dock p146 was used as template for PCR (primers o945 + o946v2) to append Illumina P5 and P7 handles, and sequenced on NS2000 with paired-end sequencing to retrieve barcode pair representation. (b) Top: Table of number of barcodes and reads for the different categories shown in panel c. Bottom: Table of number of barcode pairs, reads, and proportion displaying retention at every filtering step (i.e., both barcodes present in the well-represented set and uniquely paired [>99% reads] with a single other barcode). (c) Read count distribution by summing only on respective barcodes (not pairs) for BC1 (left) and BC2 (right). Dashed lines indicate cut-offs used for the mid- and high-counts classes of BCs. (d) Two-dimensional distributions showing the proportion of reads to the BC arising from the pair on y-axis (BC1 left panel, BC2 right panel) vs. the total read count to the given pair (x-axis). Retained pairs are within the green boundary (>5 reads and >99% unique proportion). (e) Similar to panel d, but now showing unique pairing proportions for BC1 (y-axis) vs. BC2 (x-axis) as a two-dimensional histogram. Retained pairs are in the top right corner within the green boundaries (i.e., BC1-BC2 pairs showing >99% uniqueness of read mapping to both barcodes). Marginalized histograms for BC1 and BC2 are shown on right and top respectively.
Extended Data Fig. 3 AAV packaging chimerism occurs for different serotypes.
Quantification of the fraction of discordant barcode pairs as a function of the full-length BC-to-BC average size. Library p153× (homologous harbouring short, mid, and long inserts) was packaged with AAV2 (losange) and PHP.eB (circle). Further, each serotype was packaged at 100% (standard, dark blue) and 10% (sparse, green) total dose. With both PHP.eB and AAV2, and for the three sizes of inserts, sparse packaging reduces swapping significantly (one-sided bootstrap FDR: *<0.005, **<0.0005, ***<10−5) albeit incompletely.
Extended Data Fig. 4 Sub-genomic fragments analysis on long-read libraries.
(a) Cumulative distribution of read size stratified by the number of BC flanking signposts (0 to 4, see Extended Data Figure. 1d) mapped on the read (left panel, shades of red color encoding shown in panel g, black line full unstratified read length distribution) and by the number of mapped ITRs (right panel, shades of blue color encoding shown in panel h, black line full distribution) for library p149:short-hom. Dotted vertical lines mark the expected length (ITR-to-ITR) of full ssAAV reads and scAAV-like reads. (b) IGV browser visualization of the pile-up for the alignment (minimap2) of reads to the p149 plasmid sequence. Annotation following Extended Data Figure 1 is shown below the track. (c) Same as (a), but with p150:short-non-hom. Panels (d-f): same as (a-b) but with plasmids p151:mid-hom and p152:mid-non-hom. Different rows correspond to different long-read sequencing preparations. The PacBio run was prepared on the same packaged AAV particles as the ONT rep1 sample. (g) Table showing read counts per sample stratified based on the number of identified BC-flanking signposts. (h) Table showing read counts per sample stratified based on the number of mapped ITRs.
Extended Data Fig. 5 Quality control and details of PacBio sequencing of BC1-BC2 constructs.
(a) Schematic of the molecular and processing steps for the two main distinct classes of molecular species observed in the data. Bst2 extension leads to scAAV-like species with one strand replicated by priming from the free 3’ end of the ITR. A single SMRT-bell is ligated at the free double stranded DNA end. The second class results from annealing of the two strands of ssAAV species, and SMRT-bells are ligated at both ends. (b) Cumulative distribution of read lengths, showing discrete populations at the expected sizes for ssAAV (anneal class) and scAAV-like (Bst2 extension class). Snapback molecules putatively due to the Nextera Tn5 mosaic end hairpin are also indicated. A population of more heterogenous snapback events not fully characterized in p152:mid-non-hom is indicated by a *. (c) Table quantifying the reads in the different size categories shown in the cumulative distribution of panel b. (d) Pile-up IGV visualization of the alignment (minimap2) of the p151:mid-hom reads from the ssAAV (left) and scAAV-like (right) size range, supporting our interpretation. The excess signal in the central ITR for the scAAV-like pile-up comes from the difficulty of minimap2 to deal with long reverse complementary sequences (randomly assigned read to one of the two ITR-to-ITR intervals). (e) Same as panel d, but for the putative snapback fragments. The snapback redirection event corresponds to the position of the Tn5 mosaic end hairpin, see panel m. (f-g) Read attrition for the different filtering steps applied for the ssAAV reads (f) and scAAV-like reads (g) species respectively. Notably, only the reads within the size ranges indicated in panel b are used as starting points for this analysis (p151:mid-hom ssAAV 2350-2650 bp, scAAV-like 4750-5000 bp; p152:mid-non-hom ssAAV 2300-2700 bp, scAAV-like 4750-5200 bp). (h-i) Comparisons of concordance of BC1-BC1 and BC2-BC2 (not BC1-BC2 swaps) from different parts of the reads/different orientation to support interpretation of molecular species for ssAAV reads (h) and scAAV-like reads (i). ssAAV are expected to have predominantly different BC1s and BC2s on forward and reverse reads if originating from annealing of distinct molecules. scAAV-like particles should have identifiable BC1/BC2s on both left and right halves of their reads, with matched respective BC1 & BC2s if originating from a single Bst2 extension event. Quantifications align with these expectations. Read counts are stratified by CCS orientation. (k-l) Quantification of the fraction of discordant BC1-BC2 pairs indicative of chimerism for ssAAV (k) and scAAV-like reads (l). These quantifications are reproduced in Fig. 2b. (m) Schematic of the snapback molecule with zoomed in view of the Nextera R1 and R2 handles flanking the insert. The Tn5 mosaic end putatively leading to the snapback-causing hairpin is marked in red. The two instances of BC2 in these snapback reads were compared (after placing on the same strand). Table on the right shows attrition QC table for these species (using as starting point: reads within range 3800-3900 bp for p151:mid-hom and 3800-4000 bp for p152:mid-non-hom).
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods, Notes 1 and 2 and Figs. 1–7.
Supplementary Data 1 (download XLSX )
List of plasmids and oligos used.
Supplementary Data 2 (download XLSX )
AAV packaging metadata.
Supplementary Data 3 (download XLSX )
Insert alignment metadata and read counts stratified by full/nonfull BC-to-BC length for ONT-direct (rep. 1) and ONT-PCR libraires.
Supplementary Data 4 (download XLSX )
Table of read counts of barcode swap stratified by full/partial ITR-to-ITR length (ONT-direct rep. 1 and ONT-PCR libraires).
Supplementary Data 5 (download XLSX )
Source data for Supplementary Fig. 1.
Supplementary Data 6 (download XLSX )
Source data for Supplementary Fig. 2.
Supplementary Data 7 (download XLSX )
Source data for Supplementary Fig. 3.
Supplementary Data 8 (download XLSX )
Source data for Supplementary Fig. 4.
Supplementary Data 9 (download XLSX )
Source data for Supplementary Fig. 5.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 5 (download XLSX )
Statistical source data.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lalanne, JB., Huynh, C., Mich, J.K. et al. Pool-packaged AAV libraries exhibit extensive length-dependent and homology-dependent chimerism. Nat Biotechnol (2026). https://doi.org/10.1038/s41587-026-03097-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41587-026-03097-1