Abstract
Fine-mapping refines genotype–phenotype association signals to identify causal variants underlying complex traits. However, current methods typically focus on individual genomic loci and do not account for the global genetic architecture. Here we demonstrate the advantages of performing genome-wide fine-mapping (GWFM) with functional annotations and develop methods to facilitate GWFM. In simulations and real data analyses, GWFM outperforms current methods across several metrics, including error control, mapping power, resolution, precision, replication rate and trans-ancestry phenotype prediction. Across 48 complex traits, we identify credible sets that collectively explain 18% of the SNP-based heritability \(({h}_{\mathrm{SNP}}^{2})\) on average, with 30% credible sets located outside genome-wide significant loci. Leveraging the genetic architecture estimated from GWFM, we predict that fine-mapping over 50% of \({h}_{\mathrm{SNP}}^{2}\) would require an average of 2 million samples. Finally, as proof-of-principle, we highlight a known causal variant at FTO influencing body mass index and identify new missense causal variants influencing schizophrenia and Crohn’s disease risk.
Similar content being viewed by others
Main
Despite the success of genome-wide association studies (GWAS) in identifying trait-associated variants1, the causal variants underlying complex traits remain unresolved due to extensive linkage disequilibrium (LD) between single nucleotide polymorphisms (SNPs)2 and the polygenic nature of these traits3,4. Statistical fine-mapping, often employing a Bayesian mixture model (BMM) that jointly fit several SNPs, offers a direct approach to identifying candidate causal variants5. However, current fine-mapping methods focus on genome-wide significant loci only (for example, 1–2-Mb windows centered on lead SNPs after LD clumping6,7,8,9,10,11,12) or consider one genomic region at a time (for example, an LD block13), in isolation from the rest of the genome.
Although used widely, region-specific analysis has several limitations. First, restricting fine-mapping to GWAS loci alone, which often explain only a small fraction of the total genetic variance14,15, omits meaningful signals that have not yet reached the stringent GWAS significance threshold. Second, the prior probability of association can be influential but is often conservatively predetermined (for example, as the inverse of the number of SNPs in the region6,7,16) due to challenges in estimating the genetic architecture precisely within a region. Third, fine-mapping can benefit from incorporating functional genomic annotations10,11,12,17, but region-specific methods often estimate functional priors separately before performing functionally informed fine-mapping10,11,12,17, rather than jointly modeling GWAS data and functional annotations. Finally, none of the current methods estimate the power of identifying the causal variants for a trait, which is critical to inform the experimental design of prospective studies (such a power analysis is available in GWAS18 but absent in fine-mapping).
These limitations can be addressed with GWFM analysis. Genome-wide BMMs (GBMMs), used extensively for predicting breeding values in agriculture19,20,21 and complex trait phenotypes in humans22,23,24, have emerged recently as a method for GWFM25,26. Unlike conventional GWAS and region-specific fine-mapping approaches, GBMMs jointly fit genome-wide SNPs in the model, simultaneously estimating the genetic architecture and functional priors through ‘information borrowing’24,25. For example, SNPs sharing the same functional annotation across the genome collectively prioritize that annotation based on their aggregated evidence of trait association, which, in turn, informs the estimation of individual SNP effects. This learning process is often performed iteratively using Markov chain Monte Carlo (MCMC) sampling, leading to posterior inference with superior asymptotic accuracy compared to variational inference27,28, although MCMC sampling can be computationally intensive with high-density SNPs. Moreover, GBMMs estimate polygenicity and the distribution of variant effect sizes3,20,22,23,24,29, enabling the prediction of power for prospective studies with larger sample sizes. However, relevant theory and methods have not yet been developed.
In this study, we comprehensively evaluated the performance of GWFM analysis using SBayesRC24, a state-of-the-art GBMM that enables efficient MCMC-based fitting of all common SNPs with functional annotations. Through extensive simulations under various genetic architectures, we compared methods using several metrics, including posterior inclusion probability (PIP) calibration, fine-mapping power, mapping precision, credible set size, replication rate in independent samples and out-of-sample prediction accuracy using fine-mapped variants. We developed an LD-based method to construct local credible sets (α-LCSs) for GBMM, each capturing a causal variant with posterior probability α, and estimated the proportion of SNP-based heritability \(({h}_{\mathrm{SNP}}^{2})\) explained by these LCSs. To quantify overall fine-mapping uncertainty, we provided a global credible set (GCS) (α-GCS) that captures α% of all causal variants for the trait. Leveraging the genetic architecture estimated from SBayesRC, we further developed a method to predict fine-mapping power and the proportion of \({h}_{\mathrm{SNP}}^{2}\) explained by identified variants in prospective studies, enabling estimation of the minimal sample size required to capture a desired proportion of causal variants or \({h}_{\mathrm{SNP}}^{2}\). Finally, we applied SBayesRC to 599 complex traits and diseases with 13 million SNPs and compared the fine-mapping results for 48 well-powered traits, using data primarily from the UK Biobank30 (UKB).
Results
Method overview
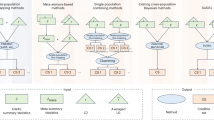
In contrast to existing fine-mapping methods that focus on individual loci identified from GWAS, GWFM combines genetic discovery with fine-mapping into a unified framework, enabling identification of causal variants across the genome. We selected SBayesRC as the method for GWFM (Fig. 1) due to its demonstrated superiority in polygenic prediction24 and its ability to effectively incorporate functional annotations. SBayesRC is a multicomponent mixture model that simultaneously analyzes all SNPs across approximately independent LD blocks13,31, using a hierarchical prior to borrow information from genome-wide functional annotations for fine-mapping within each block, where both annotation and SNP effects are estimated jointly from the data (Methods). To optimize fine-mapping performance, we estimated the number of mixture components heuristically, adapted a tempered Gibbs sampling algorithm32 to improve mixing properties and implemented a method33 to assess MCMC convergence using several independent chains (Methods). Further methodological differences between our approach and existing methods are discussed in Supplementary Note 1.
GBMM uses GWAS summary statistics and genome-wide LD reference to fine-map candidate causal variants for complex traits, incorporating functional annotations when available. Unlike region-specific fine-mapping approaches, GBMM maps causal signals across the entire genome and provides greater flexibility in modeling the distribution of causal effects by estimating priors using genome-wide SNPs (Supplementary Table 1). Different GBMMs share the same fitted model but differ in prior specification for SNP effects. As shown in the middle box, SBayesRC assumes a mixture prior consisting of a point mass at zero and multiple normal distributions with variance \({\gamma }_{k}{h}_{\mathrm{SNP}}^{2}\), where \({\gamma }_{k}=\left[0,{1}^{-5},{1}^{-4},{1}^{-3},{1}^{-2}\right]{\prime} .\)The mixing probability for SNP j in distribution k (\({\pi }_{{jk}}\)) is modeled as a linear combination of SNP annotations through a probit link. Blue: input data; green: SNP-specific parameters; red: global parameters shared across SNPs. The GBMM outputs include SNP PIPs, local and global credible sets, and prediction of fine-mapping power for future studies. Illustrations in left panel are created in BioRender; Bu, F. https://biorender.com/jtg1az6 (2026).
To capture the uncertainty in identifying causal variants driven by LD, we developed an LD-based method for constructing α-LCSs (Supplementary Fig. 1; Methods). Our approach forms an α-LCS for each candidate causal variant with a leading PIP by grouping it with SNPs in LD (r2 > 0.5) until their combined PIPs sum to α, followed by filtering based on the posterior \({h}_{\mathrm{SNP}}^{2}\) enrichment probability (PEP > 0.7) to ensure that the selected LCS explains more \({h}_{\mathrm{SNP}}^{2}\) than a random set of SNPs with the same size (Methods). Additional justification for the r2 and PEP thresholds is provided in Supplementary Note 2 and Supplementary Figs. 2 and 3. At the genome-wide level, we construct α-GCS that covers α% of all causal variants for the trait to quantify the overall uncertainty (Methods). Using the estimated genetic architecture, we further derive analytically tractable fine-mapping power prediction to estimate the minimal sample size required to achieve the desired power to identify all causal variants or those explaining a specific proportion of \({h}_{\mathrm{SNP}}^{2}\) (Methods). Our fine-mapping power prediction method is available as a publicly accessible online tool (https://gctbhub.cloud.edu.au/shiny/power/).
Calibration of fine-mapping methods under various genetic architectures
We performed extensive genome-wide simulations to evaluate SBayesRC for GWFM, comparing to FINEMAP7, SuSiE6, FINEMAP-inf8, SuSiE-inf34, PolyFun+SuSiE10 and another two GWFM methods, SBayesR22 and SBayesC (two-component SBayesR) (Supplementary Table 1). For a fair comparison, we defined fine-mapping regions consistently across all methods using independent LD blocks and applied the default parameter settings recommended for each method (Methods). Using 100,000 individuals with ~1 million HapMap3 SNPs from the UKB30, we simulated three genetic architectures: (1) a sparse architecture with 1% randomly selected causal SNPs explaining 50% of phenotypic variance, (2) a large-effects architecture where ten random causal variants contributed 10% of phenotypic variance and the rest 40%, and (3) an LD-and-minor allele frequency (MAF)-stratified (LDMS) architecture with causal variants sampled from high LD score and high MAF SNPs (Methods).
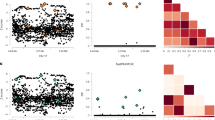
Results showed that GWFM methods generally performed better in terms of PIP calibration compared to region-specific methods, with PIPs from SBayesRC aligning closely with true probabilities of causality (measured by true discovery rate (TDR)) across all genetic architectures (Fig. 2a,d,g). In contrast, SuSiE and FINEMAP exhibited notable inflation of false discovery rate (FDR) in high-PIP SNPs (Fig. 2b,e,h), as the observed TDR was lower than the expected (FDR = 1 − TDR). While SuSiE-inf and FINEMAP-inf performed reasonably well under the sparse and large-effects architectures (despite some deflation in low-PIP SNPs), their PIPs failed to track TDR accurately under the LDMS architecture, where causal variants were not distributed randomly (Fig. 2c,f,i). Polyfun+SuSiE, when incorporating the same LD/MAF annotations as SBayesRC, showed improved performance over SuSiE and FINEMAP but still struggled with FDR control (Fig. 2i). The alternative GWFM methods, SBayesC and SBayesR, were inferior to SBayesRC under the large-effects and LDMS architectures (Supplementary Fig. 4), respectively, highlighting the importance of using several mixture components and integrating informative genomic annotations for robust calibration.
a–i, Shown are relationships between PIP and the TDR across 100 PIP bins. a, SBayesRC; sparse genetic architecture. b, SuSiE-inf and FINEMAP-inf; sparse genetic architecture. c, SuSiE and FINEMAMP; sparse genetic architecture. d, SBayesRC; large-effects genetic architecture. e, SuSiE-inf and FINEMAP-inf; large-effects genetic architecture. f, SuSiE and FINEMAMP; large-effects genetic architecture. g, SBayesRC; LDMS genetic architecture. h, SuSiE-inf and FINEMAP-inf; LDMS genetic architecture. i, SuSiE and FINEMAMP; LDMS genetic architecture.
Assessing mapping power, resolution and precision via simulations
We first compared the power of different fine-mapping strategies, GWFM and GWAS loci-based fine-mapping, by evaluating the proportion of causal variants captured by α-LCSs. For a fair comparison, GWAS loci-based fine-mapping used the same GWFM (from SBayesRC) but was restricted to 2-Mb regions around GWAS lead SNPs (P < 5 × 10−8). As expected, GWFM was significantly more powerful, with 46–61% improvement across genetic architectures (Fig. 3a–c). Moreover, GWFM also outperformed standard GWAS across a range of thresholds in terms of receiver-operator characteristic performance (Supplementary Fig. 5a–c) and retained higher power even under a stringent false positive rate control (Supplementary Fig. 5d–f), highlighting the advantage of fine-mapping the entire genome, including regions not reaching genome-wide significant in GWAS.
a–c, Plots comparing mapping power between GWFM and GWAS loci-based strategies at the same confidence level (α) for LCSs under sparse (a), large-effects (b) and LDMS (c) genetic architecture. For a fair comparison, both strategies used the same PIP estimates from SBayesRC, whereas GWAS loci-based fine-mapping constructed α-LCSs restricted to 2-Mb regions around GWAS lead SNPs (P < 5 × 10−8). ‘GWAS-loci-based’ refers to the application of GWFM only to loci with genome-wide significant SNPs. GWAS P values are based on two-sided t-test and adjustments are made for multiple comparisons. d–f, Plots comparing mapping power across different methods, including SBayesRC, SuSiE-inf and Polyfun + SuSiE under sparse (d), large-effects (e) and LDMS (f) genetic architecture. Power was quantified as the proportion of simulated causal variants included in the credible sets reported by each method. The reported power was calculated based on 100 simulation replicates. Data are presented as mean ± s.d. g–i, Plots comparing mapping resolution across SBayesRC, SuSiE-inf and Polyfun + SuSiE, under sparse (g), large-effects (h) and LDMS (i) genetic architecture, measured by the credible set size. The credible set size results were from 100 simulation replicates. Each box plot shows the spread of data; the line is the middle (median), the box covers the middle half (interquartile range (IQR)), the whiskers extend to 1.5 times the IQR, and dots show outliers. j–l, Plots comparing mapping precision across different methods, measured by the physical distance between the causal variants and the SNPs identified at PIP ≥ 0.9 in each method under sparse (j), large-effects (k) and LDMS (l) genetic architecture.
Next we compared the performance of different fine-mapping methods, focusing on SBayesRC and SuSiE-inf, as SuSiE-inf had the best PIP calibration among the region-specific methods. Our results showed that SBayesRC was significantly more powerful (Fig. 3d–f), with similar TDR (Supplementary Fig. 6) but smaller LCS sizes at the same α threshold, indicating superior mapping resolution (Fig. 3g–i). For α = 0.9, SBayesRC outperformed SuSiE-inf by up to 194% increase in power and 21% reduction in average LCS size across the three genetic architectures. Polyfun + SuSiE, which incorporated LD/MAF annotations through a stepwise approach, performed slightly better than SuSiE-inf under the LDMS architecture but remained inferior to SBayesRC. SBayesRC also achieved higher mapping precision, which was defined as the distance from an identified variant (PIP > 0.9) to the nearest causal variant. Under the sparse architecture, 98% of SNPs identified by SBayesRC were causal, and 99% of SNPs with PIP > 0.9 were within 18 kb of a causal variant, leading to a 2% increase in TDR and a 68% reduction in distance compared to other methods (Fig. 3j). The advantage of SBayesRC persisted under large-effects architecture (Fig. 3k) and became more evident under the LDMS architecture (Fig. 3l). When restricted to GWAS loci, GWFM achieved higher power and yielded smaller credible set sizes compared to other fine-mapping methods (Supplementary Fig. 7).
Furthermore, we used these simulations to evaluate GCS (Supplementary Fig. 8) and conducted sensitivity analyses to confirm the robustness of SBayesRC to missing annotations (Supplementary Fig. 9), unobserved causal variants (Supplementary Fig. 10) and the chosen value of LD matrix factorization parameter (Supplementary Fig. 11). To understand why GWFM had higher power, we investigated power gains in causal variants with different LD and MAF properties (Supplementary Fig. 12) and assessed SBayesRC within each LD block separately (Supplementary Fig. 13). These results are discussed in Supplementary Note 3. Overall, these simulation results suggested that SBayesRC is a reliable method for GWFM and can improve fine-mapping performance substantially.
Assessing replication rate, effect size estimation and prediction accuracy in real data
In real data analysis, the true causal variants are often unknown, we therefore assessed methods using metrics that do not require knowledge of causal variants. We first evaluated replication rate by calculating the proportion of variants with PIP > 0.9 in a GWAS sample that were also identified in an independent replication sample at a PIP threshold (Methods). Using 100,000 UKB individuals as a GWAS sample for height, red blood cell counts and high-density lipoprotein (HDL), SBayesRC achieved the highest replication rate in each PIP threshold, improving by 8% (compared to SuSiE-inf) to 31% (compared to FINEMAP) at PIP > 0.9 when replication n = 100,000 (Fig. 4a and Supplementary Fig. 14). When replication sample size was doubled, the improvement increased to 14% compared to SuSiE-inf and 36% compared to FINEMAP (Supplementary Fig. 14). The results were also robust under a reversed discovery-replication design and under a downsampling strategy (Supplementary Figs. 15 and 16 and Supplementary Note 4). These high replication rates suggest that variants identified by GWFM are likely to be true associations.
a, Replication rate of discovery using different methods at a given PIP threshold in the replication sample (x axis) for height in the UKB. b, Regression of the estimated marginal effect sizes in replication samples on the estimated joint effect sizes in discovery samples using different fine-mapping methods. Dashed lines show the regression slopes, where values closer to 1 indicate less bias, as the marginal effect estimated in the independent replication sample is an unbiased estimate to the true effect. Brown solid line: y = x. c, Comparison of cross-ancestry prediction accuracy using fine-mapped variants (PIP > 0.9) from SBayesRC and SuSiE-inf, based on the analysis using samples of European ancestry for GWAS and the other ancestries for validation across six UKB traits. MR, base metabolic rate; hBMD, heel bone mineral density; HT, height; RBC, red blood cell count. AFR, African ancestry; EAS, East Asian ancestry; SAS, South Asian ancestry. d, Relationship between cross-ancestry prediction transferability and PIP estimates in the European ancestry. The transferability was computed as non-EUR-R2/EUR-R2. The solid lines are regression lines across traits in each ancestry.
We next assessed bias in effect size estimates and trans-ancestry prediction using fine-mapped variants. Bias was evaluated through regressing their marginal effect sizes from the replication sample on the joint effect sizes estimated from the GWAS discovery sample, expecting a regression slope of one for unbiased estimation. In the UKB height analysis, SBayesRC produced the minimal bias, with a regression slope of 0.98, outperforming all the other methods (Fig. 4b). Given that common causal variants and their effect sizes are mostly shared across ancestries35,36, we then compared the trans-ancestry prediction using the fine-mapped variants. Specifically, we used fine-mapped variants and their posterior effect sizes (Methods) from UKB participants with European ancestry to predict phenotypes in African, East Asian and South Asian populations across six complex traits with at least 50 SNPs with PIP > 0.9. Compared to SuSiE-inf, SBayesRC improved the trans-ancestry prediction accuracy, with nearly a tenfold increase in mean relative prediction R2 across traits and ancestries (Fig. 4c). Similar improvements were observed when using 0.9-LCSs instead of individual SNPs (Supplementary Fig. 17), when comparing to Polyfun + SuSiE with the same functional annotations (Supplementary Fig. 18), or when restricting to GWAS loci (Supplementary Fig. 19). We further quantified the transferability of fine-mapped SNPs by computing the ratio of per-SNP prediction accuracy in a hold-out European ancestry sample versus a different ancestry. The result showed that relative prediction accuracy increased with PIP in the GWAS sample (Fig. 4d), consistent with a model of shared genetic effects across ancestries.
We validated these advantages of SBayesRC through simulations, confirming its superior power (Supplementary Fig. 20), replication rate (Supplementary Fig. 21a), effect size estimation (Supplementary Fig. 21b) and out-of-sample prediction using fine-mapped variants (Extended Data Fig. 1), with further discussion in Supplementary Note 5. These results indicated that SNPs identified by SBayesRC are more likely to be causal, as indicated by higher replication rates, whereas the improved prediction accuracy probably arises from both better causal variant identification and more accurate effect size estimation.
Prediction of fine-mapping power and variance explained for future studies
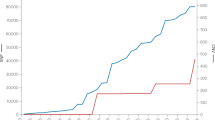
As a unique feature of the GWFM approach, the genetic architecture estimated from SBayesRC enables prediction of fine-mapping power and the proportion of \({h}_{\mathrm{SNP}}^{2}\) explained (proportion of SNP-based heritability explained (PHE)) by these variants in future studies (Methods). To evaluate our approach, we computed the predicted values of power and PHE across varying GWAS sample sizes and projected the outcome of GWFM with SBayesRC onto the prediction for a simulated trait (under sparse architecture), height30,37, HDL30,38, schizophrenia (SCZ)39,40 and Crohn’s disease (CD)30,41, which represented diverse genetic architectures, using two datasets with different sample sizes for each trait (Fig. 5a–c). Despite some variation across traits, the observed GWFM outcomes were broadly consistent with the theoretical predictions (Fig. 5d,e).
a, Estimates of \({h}_{\mathrm{SNP}}^{2}\) obtained using SBayesRC for height, CD, SCZ, HDL and a simulated trait with the sparse genetic architecture. b, Proportions of \({h}_{{\rm{SNP}}}^{2}\) explained by different mixture components in the SBayeRC model. c, Proportions of SNPs assigned to different mixture components \((\pi\)) based on their estimated effect sizes. The ‘zero,’ ‘very small,’ ‘small,’ ‘medium’ and ‘large’ effect categories correspond to the five variance components in SBayesRC, which explain 0, 0.001, 0.01, 0.1 and 1% \({h}_{\mathrm{SNP}}^{2}\), respectively. For the simulated traits, all 10,000 causal variants had small effects sampled from a single normal distribution, leading to all estimated effect sizes being assigned to the very small-effect category. Data shown in a and c are presented as the posterior mean ± standard deviation across 3,000 MCMC samples. d,e, Theoretical prediction of the power of identifying causal variants (d) and the proportion of \({h}_{{\rm{SNP}}}^{2}\) explained by the identified causal variants (e) across different GWAS sample sizes for these traits (on the liability scale for diseases). Dots show the empirical observations based on 0.9-LCSs identified from SBayesRC. The genetic architecture estimates from a and c were used as input data for the fine-mapping power prediction (d,e). To check consistency, two datasets with different GWAS sample sizes were analyzed for each trait, with genetic architecture estimates derived from the dataset with the smaller sample size.
Applying this framework to SCZ using the latest GWAS summary statistics from the Psychiatric Genomics Consortium39, we identified 13 SNPs and 156 LCSs, collectively explaining 3.9% of \({h}_{\mathrm{SNP}}^{2}\) on the liability scale42. These estimates aligned closely with our theoretical prediction, based on the 53,386 cases and 77,258 controls in their study39, which is equivalent to a sample size of 228,810 on the liability scale43 (Methods). For a prospective SCZ study using GWFM, we predict that ~180,000 cases (with equal controls and a population prevalence of 0.01) would be required to fine-map 1,000 common causal variants explaining ~20% of \({h}_{\mathrm{SNP}}^{2}\) (Fig. 5), whereas ~550,000 and 1.4 million cases would be needed to explain 50% and 80% of \({h}_{\mathrm{SNP}}^{2}\), respectively. For height, based on the UKB data (n = 350,000), we predicted that, with 5 million samples, ~10,000 variants would be identified with individual PIP > 0.9 or ~30,000 when considering LCSs, explaining up to 95% of \({h}_{{\rm{SNP}}}^{2}\) (Fig. 5). This prediction is consistent with a recent GWAS of 5 million people that reported 12,111 independently significant SNPs identified from conditional and joint analysis accounting for nearly all of the common \({h}_{{\rm{SNP}}}^{2}\) in height36.
Applying genome-wide fine-mapping to a range of complex traits
We applied GWFM with SBayesRC to 599 complex traits (597 UKB traits plus SCZ39 and CD41) and developed an online resource to query these fine-mapping results (Supplementary Table 3; ‘Data Availability’). The 597 UKB traits were selected based on z score > 4 and high confidence for heritability estimates using LD score regression44 (https://zenodo.org/records/7186871). We analyzed 13 million imputed SNPs with functional genomic annotations from Finucane et al.45, and focused on 48 well-powered traits, including SCZ39, CD41 and 46 UKB traits measured in the European ancestry individuals (Methods).
Across the 48 traits, we identified 1,820 SNPs with PIP > 0.9, with the number of fine-mapped SNPs correlated strongly with the number of GWAS loci identified by LD clumping (Supplementary Figs. 22 and 23). However, 1,158 of these SNPs were not GWAS lead SNPs, and 14.9% located outside GWAS loci, highlighting the importance of conducting genome-wide fine-mapping. Notably, 469 fine-mapped SNPs (25.8%) were associated with several traits, suggesting prevalent pleiotropy in the human genome. Moreover, the MAF of pleiotropic SNPs decreased as the number of affected traits increased (Supplementary Fig. 24), consistent with a model of negative selection46. Our results recapitulated previous findings and led to new discoveries, including new putative causal missense variants for SCZ and CD risk and recovery of variants from previous studies (Supplementary Figs. 25–27, Supplementary Tables 4 and 5 and Supplementary Note 6).
We identified 19,863 0.9-LCSs with a median size of five SNPs, of which 29.8% were located outside the genome-wide significant loci. While these LCSs captured only 0.9% of causal variants, they explained 17.7% of \({h}_{{\rm{SNP}}}^{2}\), with 2.7% PHE (relative proportion of PHE = 2.7 of 17.7 = 15.3%) attributed to LCSs in nonsignificant GWAS regions (Fig. 6a). Across trait categories, cognitive traits had the highest proportion of LCSs and PHE observed outside GWAS loci (Fig. 6b), consistent with their high polygenicity3. On average, the 0.1-GCS (expected to contain 10% of all causal variants) comprised ~1.5% of the SNPs, explaining 44.7% of the \({h}_{{\rm{SNP}}}^{2}\), with substantial differences across trait categories (Fig. 6c,d).
a, Plot showing the proportion of the identified LCSs within or outside the GWAS loci (left) and the proportion of PHE by the identified LCSs within or outside the GWAS loci (right). Results are from 19,863 identified LCSs for 48 complex traits. b, Plot showing the relationship between the proportion of LCSs outside GWAS loci and proportion of PHE across the 48 complex traits. c, Plot showing the proportion of identified GCS SNPs at different alpha threshold (proportion of causal variants captured) for the 48 complex traits (average sample size = 291,000). d, Plot showing the proportion of \({h}_{{\rm{SNP}}}^{2}\) explained by the GCS SNPs at different alpha threshold. e,f, Plots showing theoretical prediction of power (e) and the proportion \({h}_{{\rm{SNP}}}^{2}\) explained (f) by LCS SNPs at different GWAS sample sizes for the 48 complex traits, respectively. Colors indicate different trait categories. Results shown in c–f are from GWFM analysis for 48 complex traits. Each box plot in a and c–f shows the spread of data; the line is the middle (median), the box covers the middle half (IQR), the whiskers extend to 1.5 times the IQR and dots show outliers.
Leveraging the estimated genetic architecture for these 48 traits, we predicted the power of prospective fine-mapping studies. With 2 million individual samples, we predict an average fine-mapping power of 10.4% (Fig. 6e) and PHE of 57.9% (Fig. 6f). The predicted power and PHE varied substantially across trait categories: blood cell traits had both the highest power (12.8%) and PHE (69.4%), whereas cognitive traits had the lowest (5.2% power, 35.9% PHE). To achieve a PHE of 50%, blood cell traits required 1 million participants, while cognitive traits required 4 million. For PHE = 80%, the required sample sizes increased to 3 million for blood cell traits and 8 million for cognitive traits (Fig. 6f).
Incorporating functional annotations improves fine-mapping
SBayesRC incorporates functional annotations by learning their contributions directly from the data, rather than relying on pre-assigned weights. We observed several lines of evidence supporting the value of incorporating functional annotations in fine-mapping. For example, the 1,820 fine-mapped SNPs showed greater enrichment than GWAS significant SNPs in biologically important regions, such as coding sequences, transcription start sites, nonsynonymous variants and evolutionarily conserved regions, and were more depleted in repressed regions (Fig. 7a). Additional supporting results are presented in Supplementary Figs. 28–31 and discussed in Supplementary Note 7.
a, Plot showing enrichment of the genome-wide fine-mapped SNPs from SBayesRC and GWAS clumped SNPs in the 22 main functional categories defined in the LDSC baseline model. b, Plot showing the prioritized causal variant at the FTO locus for BMI. The top track shows the FTO locus plot of the standard GWAS for BMI, the second track shows the similar plot but with the PIP from SuSiE-inf for BMI and the third track shows the similar plot with PIP from SBayesRC. The dots with green circles are credible set SNPs identified by SBayesRC. The starred SNP is the known causal variant (rs1421085) for obesity at the FTO locus supported by previous functional studies. The GWAS P values were based on two-sided t-test and adjustments were made for multiple comparisons. Chr.: chromosome. c, Plot showing the per-SNP heritability enrichment for the causal variant (rs1421085), the GWAS lead variant (rs11642015) and the secondary signal (rs76488452) for BMI at the FTO locus. The annotations on the x axis were those present at least once in these three variants, excluding quantitative annotations and duplicated annotations with flanking windows. d, Plot showing the prioritized causal variant at the ACTR1B locus for SCZ. The top track shows the ACTR1B locus plot of the standard GWAS, and the second track shows the similar plot but with the PIP from SuSiE-inf, and the third track shows the similar plot with PIP from SBayesRC. The starred SNP is the fine-mapped variant (rs11692435) for SCZ at the ACTR1B locus. The GWAS P values were based on two-sided t-test and adjustments were made for multiple comparisons. e, Plot showing the per-SNP heritability enrichment for the causal variant (rs11692435) at the ACTR1B locus. Data shown in c and e are presented as the posterior mean ± standard deviation across 3,000 MCMC samples.
One notable example is rs1421085 at the FTO locus—a known causal variant influencing body mass index (BMI)47, which was fine-mapped in a credible set for body fat percentage, hip circumference and waist circumference. Although many SNPs at FTO reached genome-wide significance, both SBayesRC and SuSiE-inf identified a five-SNP 0.9-LCS including rs1421085, along with four additional SNPs in almost complete LD (minimum LD r2 = 0.997) and with strong annotation overlaps (Fig. 7b). Notably, rs1421085 was prioritized in SBayesRC (PIP = 0.47), probably due to its cross-species conservation annotations (Fig. 7c), whereas SuSiE-inf prioritized the GWAS lead SNP instead, underscoring the importance of incorporating functional annotations into fine-mapping. Furthermore, SBayesRC identified a secondary signal with another 0.9-LCS, which aligned with the result of SuSiE-inf but had not been reported previously. The lead SNP rs76488452 (PIP = 0.41) resided in a primate conserved region and was significant in conditional and joint analysis48 (P = 1.8 × 10−17) when conditioned on rs1421085, but was only nominally significantly in GWAS (P = 3.6 × 10−4), indicating a masked effect at the SNP49.
Another example is a missense variant (rs13107325) in SLC39A8—a gene implicated in the latest SCZ analysis for its function in regulating dendritic spine density50,51. This variant was identified through an aggregated effect from several key functional annotations (Supplementary Fig. 25a). We also observed evidence of a secondary signal at a distinct SNP (rs13107325, PIP = 0.82), which was annotated with primate conservation. This secondary signal was also shown in the SuSiE-inf analysis, albeit with a lower PIP value.
We also highlight a new variant missed by other fine-mapping methods. rs11692435—a nonsynonymous variant in ACTR1B—was prioritized by SBayesRC with a high PIP (0.96), probably due to its strong functional relevance (Fig. 7d,e). Both brain-specific proteome-wide association studies and protein quantitative trait locus (QTL) colocalization analyses have found association between ACTR1B and SCZ52,53, with rs11692435 identified as a single-cell expression QTL for ACTR1B in excitatory neurons54—a cell type consistently implicated in SCZ39,55. Despite its biological relevance, this variant, whose GWAS P value (3 × 10−8) was just above the genome-wide significance threshold, was missed by the fine-mapping analysis (using FINEMAP) in the original study39 and by our SuSiE-inf analysis (PIP = 0.19).
These results demonstrate that incorporating functional annotations can improve the identification of biologically relevant variants. Together, they highlight the power of SBayesRC in identifying putative causal variants and provide a valuable resource for downstream analysis and functional validation.
Discussion
In this study, we evaluated GWFM comprehensively using SBayesRC through extensive simulations and real data analyses. Compared with existing fine-mapping methods that analyze one genomic region at a time, SBayesRC demonstrated superior PIP calibration, leading to better FDR control and improved mapping power, resolution and precision (Fig. 3). In real trait analyses, SBayesRC achieved higher replication rates, lower estimation bias and greater prediction accuracy using fine-mapped SNPs in independent validation samples (Fig. 4), identifying putative causal variants missed by other methods (Fig. 7). To date, most fine-mapping studies have focused exclusively on genome-wide significant GWAS loci6,7,8,9,10,11,12. However, our analysis revealed, that averaging across traits, 30% of the LCSs (median size = 5) identified by SBayesRC were beyond genome-wide significant loci, contributing to 15% of the \({h}_{{\rm{SNP}}}^{2}\) captured by all LCSs (Fig. 6a), highlighting SBayesRC as a reliable and superior method for GWFM analysis.
The advantage of GWFM over GWAS loci-based fine-mapping arise from several key aspects. First, SBayesRC incorporates functional genomic information from genome-wide variants with shared annotations, improving SNP effect weighting and the ability to identify causal variants that would otherwise be overlooked in GWAS. Second, fine-mapping can identify causal variants masked by LD that do not reach genome-wide significance in GWAS49,56. Third, the P value threshold in GWAS is designed to control false positive rate under the null hypothesis2, whereas PIP directly controls the FDR within a Bayesian framework57. Despite these differences, GWFM often captures findings from the traditional two-step approach (GWAS followed by fine-mapping), but not vice versa. For example, our identified 0.9-LCSs by GWFM captured most GWAS clumped loci (for example, 91% for CD; Supplementary Fig. 32), indicating that our GWFM approach provides consistent and conceptually equivalent findings to the traditional two-step approach (GWAS followed by fine-mapping) and can be used as a complementary method. To further assess locus-level importance, we estimated the PHE by each LCS. Notably, LCSs with similarly high cumulative PIP values show substantially different PHE, which can be used to reflect GWAS signal strength. A Manhattan-like plot of the GWFM results showed that the magnitude of PHE for LCS was highly consistent with the −log10(P value) from GWAS for both CD and simulation (Extended Data Fig. 2). Our LCSs identified additional signals under the genome-wide significance, with ~90% of them containing true causal variants in simulation. Together, we recommend performing GWFM analysis directly, rather than restricting fine-mapping to genome-wide significant loci identified from GWAS.
The advantages of SBayesRC over the region-specific fine-mapping methods is because it fits all SNPs jointly across LD blocks, learns realistic effect-size distributions and priors informed by functional annotations, and uses MCMC sampling to more accurately approximate posterior distributions27. This unified genome-wide Bayesian framework enables robust estimation of genetic architecture parameters and improves inference in regions with complex LD structure, such as the major histocompatibility complex region10,39 (Supplementary Fig. 33). Additional methodological detailed comparisons are described in Supplementary Note 8.
In addition to the improved fine-mapping performance, we introduced new features for GWFM using MCMC sampling, including refining LCS by PEP (Supplementary Note 1 and Supplementary Figs. 34 and 35), constructing GCS, and estimating the proportion of polygenicity and \({h}_{{\rm{SNP}}}^{2}\) explained by LCSs and GCSs. Moreover, our GWFM can inform the sample size required to identify a certain proportion of causal variants or \({h}_{{\rm{SNP}}}^{2}\) explained in future studies, and implemented a robust posterior inference using scalable tempered Gibbs sampling algorithm32 and several independent chains to assess the convergence of each SNP’s PIP. A detailed discussion of these new features can be found in Supplementary Note 9.
We note several limitations of this work. First, method evaluations were performed under three simulated scenarios and do not cover all possible effect-size distributions. Second, as a summary-level method, SBayesRC ignores LD between LD blocks, depends on a well-matched LD reference (Supplementary Fig. 36), and uses an empirically defined LD threshold to construct LCS, which may affect performance in regions with complex LD. Third, our applications were restricted to GWAS data from European ancestry, primarily from UKB, and the functional annotations from BaselineLD (v.2.2)58. A more detailed discussion of limitations is provided in Supplementary Note 10. Despite these limitations, our study provides a powerful and robust GWFM framework for identifying causal variants, highlighting the advantages of this approach over existing region-specific fine-mapping methods. With its capacity to enhance mapping power in the current study and to predict mapping power for future studies, we anticipate that GWFM using a state-of-the-art GBMM will become the preferred method for fine-mapping complex traits.
Methods
Ethics approval
The University of Queensland Human Research Ethics Committee B (2011001173) provides approval for analysis of human genetic data used in this study on the high-performance cluster of the University of Queensland.
Low-rank GBMM
We used state-of-the-art GBMM that employed a low-rank model to improve computational efficiency and robustness24. As described below, the low-rank GBMM can be derived from the individual-level model. Consider a multiple linear regression of phenotypes on genotypes:
where y is an n × 1 vector of complex trait phenotypes and X is an n × m matrix of mean-centered genotypes, β is \(m\times 1\) vector of SNP effects on the trait, and e is \(n\times 1\) vector of residuals with \(\mathrm{var}({\mathbf{e}})=\mathbf{I}{\sigma }_{{\text{e}}}^{2}\). Let
where \({\pmb{\Lambda}}\) \({\rm{and}}\) \(\mathbf{U}\) are diagonal matrix of eigenvalues and matrix of eigenvectors for the LD correlation matrix \(\mathbf{R}=\mathbf{X}^{{{\prime} }}\mathbf{X}/n,\) respectively. It follows that \(\mathbf{K}^{{{\prime} }}\mathbf{K}=\mathbf{P}{n}^{-1}\), where P is the projection matrix of y on the column space of X and \(\mathbf{K}\mathbf{K}^{{\prime} }=\mathbf{I}{n}^{-1}\). Multiplying both sides of equation (2) by K gives
or
When only the top q principal components of R are used, the dimension of w is q × 1 and Q is q × m. By selecting q ≪ n, this model would have a substantially lower rank than equation (1), improving the computational efficiency for the estimation of \(\boldsymbol{\beta}\). By default, our method automatically determines q through pseudo cross-validation based on GWAS summary statistics (Supplementary Note 10 in ref. 24). With a recognition that \(\mathbf{ b={X}^{{{\prime} }}y}/n\) is the GWAS marginal effect estimates, w can be computed directly from the GWAS summary statistics
with Λ and U obtained from a LD reference sample, and Q can be computed as
Essentially, the low-rank model transforms the mutually correlated GWAS marginal effects (b) into a set of independent data points (w), while maintaining the ability to estimate individual SNP joint effects (β). In practice, we compute w and Q within quasi-independent LD blocks in the human genome. With this low-rank model, we can estimate β for all common variants jointly through considering β as random effects. In addition, this model allows a direction estimation of the residual variance, as var \(\left(\epsilon \right)=\mathbf{I}{\sigma }_{e}^{2}{n}^{-1}\), which can be used as a nuisance parameter to accommodate heterogeneity in the summary statistics and LD reference24.
SBayesC, SBayesR and SBayesRC
We considered three GBMMs with specifying different prior distribution of SNP effects. In SBayesC, the prior for the effect of SNP j is given by
where \({\sigma }_{\beta }^{2}\) is the common variance across all causal variants following a scaled inverse chi-squared prior distribution, π is the proportion of SNPs with nonzero effects following a uniform prior distribution24 and \(\phi\) is a point mass at zero. This is also the underlying model for most fine-mapping methods, assuming that only a fraction of SNPs are causal with nonzero effects.
SBayesR22 extends SBayesC by assuming a more flexible prior distribution for SNP effects, using a multicomponent Gaussian mixture:
where \({\sigma }_{g}^{2}\) is the total genetic variance, \(\boldsymbol{\gamma} ={[0,{10}^{-5},{10}^{-4},{10}^{-3},{10}^{-2}]}^{{\prime} }\) are prespecified scale factors, representing zero, very small, small, medium and large effect size categories, and \({\pi }_{k}\) is the prior probability of SNP j belonging to the kth distribution. This allows SBayesR to better model genetic architectures with a wide range of effect sizes compared to SBayesC.
To further account for functional annotations, SBayesRC24 assumes a SNP-specific prior probability of mixture-component membership, \({\pi }_{{jk}}\), which depends on the annotations at each SNP through a generalized linear model:
where f(∙) is the probit link function, μk is the intercept, Ajl is the value of annotation l on SNP j (either binary or continuous annotations) and αkl is the effect of annotation l on the prior probability of SNP j belonging to the kth distribution. To enable a data-driven estimation of annotation effects, we assume an independent and identical normal prior for each annotation effect, \({\alpha }_{{kl}} \sim N(0,{\sigma }_{{\alpha }_{k}}^{2})\), implying equal contribution of all functional annotations. These priors are then estimated in a trait-specific manner. Details of the method and the MCMC sampling scheme can be found in ref. 24.
In this study, we ran SBayesRC with four MCMC chains, each consisting of 3,000 iterations with the first 1,000 discarded as burn-in. The posterior effect size for each SNP was computed as the mean of SNP effect sizes across MCMC samples from the GWFM model. Analyzing 13 million SNPs with 96 annotations, SBayesRC required only 150 Gb of RAM and 13 h of computation using 24 CPU cores, which are commonly available in a standard computing cluster.
LD blocks
We partitioned the genome into 1,588 approximately independent LD blocks, following the methods of Li et al.13 and Berisa and Pickrell59. Specifically, LD blocks from ref. 13 were merged to ensure a minimum length of 1 Mb and major histocompatibility complex region was treated as a single LD block due to its complex LD patterns. As a result, our LD blocks ranged in size from 1 to 30 Mb, with a median length of 1.5 Mb.
Calculation of PIP
We assessed the strength of joint association of each SNP using PIP, that is, the probability of a SNP being included with a nonzero effect in the model, given the data (w). In SBayesRC, we computed PIP for SNP j as
where δj= 1 indicating a null effect and δj = 2,…, 5 indicating a nonzero component. \(\Pr \left({\delta }_{j}=1|\mathrm{Data}\right)\) is often computed by counting the frequency of \({\delta }_{j}=1\) across MCMC samples23,25. To improve precision, we use Rao-Blackwellized estimates60,61 and compute the posterior mean of [\(1-\Pr \left({\delta }_{j}=1|\mathbf{w},\boldsymbol\theta \right)]\) conditional on data and all the other parameters except \({\beta }_{j}\) \((\boldsymbol\theta )\) across T iterations. That is, suppose \({p}_{j}^{\left(t\right)}=1-\Pr\)\(\left({\delta }_{j}=1|\mathbf{w},{\boldsymbol\theta }^{(t)}\right)\), we have
where
with \(f(\mathbf{w}|{\delta }_{j}=k,{\boldsymbol\theta }^{(t)})\) being the likelihood given \({\delta }_{j}\) and the sampled values of other parameters. A detailed derivation can be found in Supplementary Note 11.
To further improve robustness, we run several MCMC chains simultaneously, randomly shuffling the sampling order of SNP effects in each iteration. For SNP j in chain s at iteration t, we have
Following Gelman and Rubin62, we calculated the potential scale reduction factor (PSRF) for each SNP PIP:
where \({W}_{j}={E}_{s}[{{\rm{Var}}}_{t}({p}_{j}^{(s,t)})]\) is the averaged within-chain variance across chains, and \({B}_{j}={{\rm{Var}}}_{s}[{E}_{t}({p}_{j}^{(s,t)})]\) is the variance of the means of the chains. By comparing the between-chain variance to the within-chain variance, PSRF assesses whether the MCMC chains have converged to a stationary distribution. Empirically, a PSRF value below 1.2 indicates adequate convergence62. By pooling samples across chains, our final PIP estimate is
We report the PSRF value to imply whether the SNP PIP is converged or not. SNPs with high PIP but large PSRF should be interpreted with caution, and longer chains are recommended if many variants have high PSRG values. Across 48 well-powered complex traits analyzed, none of fine-mapped individual SNPs (singleton LCSs) had PSRF < 1.2, while 99.4% of SNPs included in the LCSs had PSRF < 1.2 and 99.9% had PSRF < 2, suggesting adequate convergence in our analyses.
Heuristic estimation of mixture components
The standard SBayesRC requires prespecifying the number of mixture components before analysis, which can affect fine-mapping if an unnecessary small-effect component is included, leading to inflated PIP. To address this, we implemented a heuristic procedure for determining the number of mixture components in SBayesRC. Specifically, we run several models (varying from two to five mixture components) in parallel for 500 MCMC iterations, and assess model fit using posterior distribution of \({h}_{{\rm{SNP}}}^{2}\). We selected the model with the highest posterior mean, where the lower bound (posterior mean − s.e.) was greater than that of the second-best model. If no significant difference was found, the model with the fewest components was chosen.
Tempered Gibbs sampling
Joint analysis of all common variants presents a challenge for MCMC mixing when using the standard Gibbs sampling (GS) algorithm. For example, in a finite number of iterations, a common causal variant may fail to enter the model if the SNP in complete LD with it has already been selected. To address this issue, we incorporated a tempered GS (TGS) algorithm32 into our SBayesRC model. Essentially, TGS improves mixing by (1) strategically selecting SNPs for updating (‘informed choice’), focusing on those whose indicator variable \({\delta }_{j}\) is currently in the tail of the full conditional distribution and (2) allowing for longer jumps across local maxima by sampling \({\delta }_{j}\) from a tempered distribution. For computational efficiency, we applied TGS only to SNPs with nonzero effects sampled from the standard GS to assess whether alternative SNPs in high LD (r2 > 0.95) could provide a better fit. Further details on our TGS implementation and the corresponding pseudocode are provided in Supplementary Note 12.
Local credible sets
Following earlier work9, we defined LCS at confidence level α (α-LCS) as the minimum set of SNPs that contains at least one causal variant with probability α. To construct an α-LCS, SNPs were ranked by their PIPs. For a focal SNP that has the highest PIP and not yet assigned to an LCS, a candidate set was formed by including all remaining SNPs in high LD (r2 > 0.5) with the focal SNP. The α-LCS was then determined by summing PIPs in decreasing order until the total reached at least α. This process was iterated over SNPs until no possible LCS can be formed. For each α-LCS, we computed a posterior SNP heritability enrichment probability (PEP) that the LCS explains more \({h}_{{\rm{SNP}}}^{2}\) than a random set of SNPs with the same size. We reported all 0.9-LCSs with PEP > 0.7 for each LD block. A schematic of the procedure is shown in Supplementary Fig. 1, differences to existing approaches are discussed in Supplementary Note 1, and further justification of these thresholds based on simulations is provided in Supplementary Note 2.
Global credible sets
The α-GCS identifies the smallest set of SNPs that capture α% of the causal variants, quantifying the overall uncertainty in fine-mapping and reflecting the genetic architecture of the trait. The size of α-GCS decreases with increasing power and a smaller fraction of small-effect variants. We computed the α-GCS as the cumulative sum of decreasingly ranked PIPs until the total exceeded \({\alpha \times m}_{c}\), where mc was the estimated number of causal variants from GBMM for the trait. Since each SNP is causal with probability PIPj, the expected number of causal variants in a set Ω is E[Number of causal variants in Ω] \(=\mathop{\sum }\limits_{j{\in }\varOmega }{\mathrm{PIP}}_{j}.\) Since the α-GCS is constructed such that \({\sum }_{j\in \mathrm{GCS}}{\mathrm{PIP}}_{j}\ge {\alpha \times m}_{c}\), this directly implies that E[Number of causal variants in α-GCS] ≥ α × mc.
Estimation of power and variance explained given the data
For the identified SNPs using individual PIP or LCS, we estimated the true positive rate (TPR or power) of identifying the causal variants at a given threshold α,
where M is the total number of SNPs and π is the proportion of causal SNPs. A formal derivation is given in Supplementary Note 13.
The PHE by LCSs or GCS is estimated based on the MCMC samples of SNP effects. For a focal set (\(\Omega\)) of SNPs in iteration t, we computed PHE as
where \({\beta }_{\bullet t}\) is the sampled effect for a SNP in iteration t in scaled genotype units. Finally, we computed the posterior mean across MCMC iterations as the estimate for \({{\rm{PHE}}}_{\Omega }\)
where L is the total number of MCMC iterations.
Prediction of power and variance explained for prospective studies
We aim to predict the power of identifying a certain proportion of the causal variants in a prospective fine-mapping analysis, given a sample size (n) and the genetic architecture of the trait, when PIP from a GBMM is used as the test statistic. As shown in Supplementary Note 14, assuming that variance explained by the causal variant is \(v\), the sampling distribution of PIP from the multicomponent mixture model, for example, SBayesRC, is
where \({A}_{k}=\frac{{\pi }_{k}}{{\pi }_{1}}{\lambda }_{k}^{\frac{1}{2}}{C}_{k}^{-\frac{1}{2}}{\rm{and}}\,{B}_{k}=\frac{n{\sigma }_{e}^{2}}{2{C}_{k}}\) are two constants given the genetic architecture parameters \((\pi \,\mathrm{and}\,{h}_{\mathrm{SNP}}^{2},\mathrm{with}\,{\lambda }_{k}=\frac{{\sigma }_{e}^{2}}{{\gamma }_{k}{\sigma }_{g}^{2}}\mathrm{and}\,{C}_{k}=n+{\lambda }_{k}\), and Z is a data-dependent random variable following a noncentral Chi-square distribution with the noncentrality parameter \(\mathrm{NCP}=\frac{{nv}}{{\sigma }_{e}^{2}}\):
Given the threshold of PIP being \(\alpha\) for the hypothesis test, the power to detect this causal variant can be calculated as
where \(f(\mathrm{PIP}{|v})\) is equation (19) above. To compute the power for identifying any causal variant, we integrated out \(v\) by
where
and \({f}_{\beta }(\bullet )\) is the distribution of \(\beta\) estimated from the SBayesRC model.
Therefore, given a sample size, the expected number of causal variants identified from fine-mapping is
The expected proportion of genetic variance explained by the fine-mapped variants is
Since it is challenging to obtain an analytical solution computationally, we opted to estimate these quantities through Monte Carlo simulation (Supplementary Note 14). Although our theoretical prediction does not model LD between SNPs, the extent to which the observed values were consistent with the predicted suggests that LD had been effectively accounted for, albeit not perfectly, by our LCSs. Although we focused on European ancestry in this study, our power prediction method is, in principle, applicable to other ancestries; however, the genetic architecture would need to be estimated using ancestry-specific GWAS data, as it may vary across populations.
Disease sample size at the liability scale
For diseases or binary traits, we converted the GWAS summary statistics from the linear mixed model to the liability scale before running GBMM. Based on the method in ref. 43, we estimated the sample size at the liability scale that gives an equivalent power to detect a locus affecting a quantitative trait with the same properties,
where \(K\) is the disease prevalence, ν is the sample prevalence, \(i=h/K\) with h being the height of standard normal curve at the truncation point \(t=1-K\,\mathrm{and}\,{N}_{01}\) is the total number of cases and controls. Given the z score (zj) from the original GWAS summary data for SNP j, the marginal effect and its standard error at the liability scale can be estimated as following ref. 63
where \({p}_{j}\) is the MAF of the SNP.
The results from GBMM using the converted summary statistics will be directly comparable across traits. In our prediction analysis of power, we compared results between diseases and quantitative traits based on the equivalent sample size estimated from equation (26). Similarly, to estimate the number of cases required, in a case–control study with equivalent number of controls, to achieve a certain power, we rearranged the same equation so that
Simulations based on imputed genotype data from the UKB
To evaluate the performance of GBMM, we ran simulations using imputed genotype data from the UKB after quality control (QC). We selected 300,000 unrelated people and included ~1.2million HapMap3 SNPs with MAF > 0.01, Hardy–Weinberg equilibrium test P > 1 × 10−6, genotyping rate >0.95, and imputation information score >0.8 for simulations.
We randomly sampled \({m}_{c}=\mathrm{10,000}\) casual variants from the genome for 100,000 people and simulated complex trait phenotypes based on the following model:
where X is the genotype matrix for the causal variants, β is the vector of causal variant effects and \(\mathbf{e}{\boldsymbol{ \sim }}N(0,\mathbf{I} {\mathrm{var}}(\mathbf{X}\boldsymbol\beta )/(1/{h}^{2}-1))\,\mathrm{with}\,{h}^{2}=0.5\) being the heritability. We simulated three genetic architectures: sparse architecture with causal effects \({\beta }_{i}{\boldsymbol{ \sim }}N(0,{h}^{2}/{m}_{c})\), large-effects architecture with ten random causal variants with effects from \(N(0,0.2{h}^{2}/10)\) and the remaining causal variants with effects from \(N(0,0.8{h}^{2}/9,990)\), and LDMS architecture where causal variants were sampled only from high LD and high MAF group.
We ran a standard GWAS on simulated phenotypes and applied GBMM (SBayesRC24, SBayesR22 and SBayesC) implemented in GCTB, along with SuSiE6, FINEMAP7, SuSiE-inf8, FINEMAP-inf8 and PolyFun + SuSiE10, to compute PIPs and estimate effect sizes from GWAS summary statistics. For all methods, we followed the recommended default parameter settings. Specifically, for SuSiE and SuSiE-inf, we set the purity parameter to 0.5, the maximum number of SNPs per credible set (n_purity) to 100 and maximum number of causal variants to ten. We used imputed genotypes from 10,000 random samples of European ancestry from the UKB as the LD reference. All fine-mapping methods used the same LD reference sample and identical independent LD blocks to define fine-mapping regions. We repeated the whole process 100 times and quantified the TDR, power, mapping precision and replication rate for each method. TDR was quantified as the proportion of identified α-LCS containing at least one causal variant, whereas power was defined as the proportion of simulated causal variants included in the identified α-LCS. Mapping precision was computed as the physical distance between the identified SNPs and nearest causal variants.
For replication analysis, we split the full simulated dataset randomly into a discovery cohort (n = 100,000) and two independent replication cohorts with n = 100,000 and n = 200,000, respectively. GWAS and GWFM were run separately on each cohort. Replication was assessed by evaluating whether variants identified in the discovery set (for example, with PIP > 0.9) also showed strong support in the replication set at different PIP thresholds.
Real data analysis
We analyzed 597 complex traits from UKB using GWAS summary data from Neale’s lab (‘Data availability’) and SCZ39 and CD41 using previously published GWAS summary data. The 597 UKB traits were selected based on z score > 4 and high-confidence heritability estimates using LD score regression58. We used annotations from the baseline model BaseLineLD (v.2.2)58 and extract imputed SNPs with MAF > 0.001 overlapping with these annotations, resulting in 13,065,104 SNPs after QC. We used 10,000 random UKB samples as the LD reference to run the SBayesRC and other region-specific fine-mapping analysis. Furthermore, we selected 48 well-powered traits with relatively large sample size (n > 100, 000), high heritability estimate (h2 > 0.1) and at least one fine-mapped SNP with PIP > 0.9.
For polygenic score prediction using fine-mapped variants, we performed quality control on the UKB imputed genotype data30. Following ref. 24, we kept SNPs with MAF > 0.01, Hardy–Weinberg equilibrium test P > 10−10, imputation info score >0.6 within each ancestry samples. We removed sex-mismatched, withdrawn and cryptic related samples. The final dataset consisted of four ancestries: European (n = 347,800), East Asian (n = 2,252), South Asian (n = 9,436) and African (n = 7,006). For continuous traits, the phenotypes were filtered within the range of mean ± 7 s.d., followed by rank-based inverse-normal transformed within each ancestry and sex group. GWAS were performed using PLINK264 with sex, age and the first ten principal component as covariates. Linear regression was used for continuous traits, whereas logistic regression was applied for binary traits.
In real data analysis, because causal variants are often unknown, direct evaluation of mapping precision is not feasible. Alternatively, we evaluate the replication rate of the identified variants using an independent sample8. Here we define replication rate as the proportion of high PIP (for example, PIP > 0.9) variants from the discovery GWAS sample to be repeatedly identified in an independent (replication) sample with the same or a smaller PIP threshold. We ran the GWFM analysis using 100,000 samples of European ancestry from the UKB as the discovery cohort for height, red blood cell counts and HDL. We then assessed the replication rate using an independent cohort of 100,000 or 200,000 samples from the same population.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Our SBayesRC-enabled genome-wide fine-mapping results for 599 complex traits are available at https://sbayes.pctgplots.cloud.edu.au/data/SBayesRC/share/Finemap/v1.1/ and via Zenodo at https://doi.org/10.5281/zenodo.15293207 (ref. 65). The UKB data are available through formal application to the UKB (http://www.ukbiobank.ac.uk). The GWAS summary data for 597 complex traits in UKB are from http://www.nealelab.is/uk-biobank/. The LD data used in this study are available at https://cnsgenomics.com/software/gctb/#Download. All the other datasets used in this study are available in the public domain.
Code availability
Summary-data-based genome-wide BMMs are implemented in a public available software GCTB at https://cnsgenomics.com/software/gctb/#Download and are available via Zenodo at https://doi.org/10.5281/zenodo.18478994 (ref. 66). Methods to compute LCS and GCS have also been implemented in GCTB (https://cnsgenomics.com/software/gctb/#Genome-wideFine-mappinganalysis). Online tool for predicting fine-mapping power: https://gctbhub.cloud.edu.au/shiny/power/.
References
Abdellaoui, A., Yengo, L., Verweij, K. J. H. & Visscher, P. M. 15 years of GWAS discovery: realizing the promise. Am. J. Hum. Genet. 110, 179–194 (2023).
Uffelmann, E. et al. Genome-wide association studies. Nat. Rev. Methods Primers 1, 59 (2021).
Zeng, J. et al. Widespread signatures of natural selection across human complex traits and functional genomic categories. Nat. Commun. 12, 1164 (2021).
O’Connor, L. J. et al. Extreme polygenicity of complex traits is explained by negative selection. Am. J. Hum. Genet. 105, 456–476 (2019).
Schaid, D. J., Chen, W. & Larson, N. B. From genome-wide associations to candidate causal variants by statistical fine-mapping. Nat. Rev. Genet. 19, 491–504 (2018).
Wang, G., Sarkar, A., Carbonetto, P. & Stephens, M. A simple new approach to variable selection in regression, with application to genetic fine mapping. J. R. Stat. Soc. Series B 82, 1273–1300 (2020).
Benner, C. et al. FINEMAP: efficient variable selection using summary data from genome-wide association studies. Bioinformatics 32, 1493–1501 (2016).
Cui, R. et al. Improving fine-mapping by modeling infinitesimal effects. Nat. Genet. 56, 162–169 (2024).
Zou, Y., Carbonetto, P., Wang, G. & Stephens, M. Fine-mapping from summary data with the ‘Sum of Single Effects’ model. PLoS Genet. 18, e1010299 (2022).
Weissbrod, O. et al. Functionally informed fine-mapping and polygenic localization of complex trait heritability. Nat. Genet. 52, 1355–1363 (2020).
Kichaev, G. et al. Integrating functional data to prioritize causal variants in statistical fine-mapping studies. PLoS Genet. 10, e1004722 (2014).
Yang, Z. et al. CARMA is a new Bayesian model for fine-mapping in genome-wide association meta-analyses. Nat. Genet. 55, 1057–1065 (2023).
Li, X., Sham, P. C. & Zhang, Y. D. A Bayesian fine-mapping model using a continuous global-local shrinkage prior with applications in prostate cancer analysis. Am. J. Hum. Genet. 111, 213–226 (2024).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
So, H.-C., Gui, A. H. S., Cherny, S. S. & Sham, P. C. Evaluating the heritability explained by known susceptibility variants: a survey of ten complex diseases. Genet. Epidemiol. 35, 310–317 (2011).
Chen, W. et al. Fine mapping causal variants with an approximate Bayesian method using marginal test statistics. Genetics 200, 719–736 (2015).
Kichaev, G. & Pasaniuc, B. Leveraging functional-annotation data in trans-ethnic fine-mapping studies. Am. J. Hum. Genet. 97, 260–271 (2015).
Zhang, Y., Qi, G., Park, J.-H. & Chatterjee, N. Estimation of complex effect-size distributions using summary-level statistics from genome-wide association studies across 32 complex traits. Nat. Genet. 50, 1318–1326 (2018).
Erbe, M. et al. Improving accuracy of genomic predictions within and between dairy cattle breeds with imputed high-density single nucleotide polymorphism panels. J. Dairy Sci. 95, 4114–4129 (2012).
MacLeod, I. M. et al. Exploiting biological priors and sequence variants enhances QTL discovery and genomic prediction of complex traits. BMC Genomics 17, 144 (2016).
Habier, D., Fernando, R. L., Kizilkaya, K. & Garrick, D. J. Extension of the Bayesian alphabet for genomic selection. BMC Bioinformatics 12, 186 (2011).
Lloyd-Jones, L. R. et al. Improved polygenic prediction by Bayesian multiple regression on summary statistics. Nat. Commun. 10, 5086 (2019).
Moser, G. et al. Simultaneous discovery, estimation and prediction analysis of complex traits using a Bayesian mixture model. PLoS Genet. 11, e1004969 (2015).
Zheng, Z. et al. Leveraging functional genomic annotations and genome coverage to improve polygenic prediction of complex traits within and between ancestries. Nat. Genet. 56, 767–777 (2024).
Xiang, R. et al. Genome-wide fine-mapping identifies pleiotropic and functional variants that predict many traits across global cattle populations. Nat. Commun. 12, 860 (2021).
Shrestha, M. et al. Enhanced genetic fine mapping accuracy with Bayesian Linear Regression models in diverse genetic architectures. PLoS Genet. 21, e1011783 (2025).
Carbonetto, P. & Stephens, M. Scalable variational inference for Bayesian variable selection in regression, and its accuracy in genetic association studies. Bayesian Analysis 7, 73–108 (2012).
Blei, D. M., Alp, K. & McAuliffe, J. D. Variational inference: a review for statisticians. J. Am. Stat. Assoc. 112, 859–877 (2017).
Zeng, J. et al. Signatures of negative selection in the genetic architecture of human complex traits. Nat. Genet. 50, 746–753 (2018).
Bycroft, C. et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 562, 203–209 (2018).
Berisa, T. & Pickrell, J. K. Approximately independent linkage disequilibrium blocks in human populations. Bioinformatics 32, 283–285 (2016).
Zanella, G. & Roberts, G. Scalable importance tempering and Bayesian variable selection. J. R. Stat. Soc. Series B Stat. Methodol. 81, 489–517 (2019).
Gelman, A. & Rubin, D. B. Inference from iterative simulation using multiple sequences. Stat. Sci. 7, 457–472, 416 (1992).
Wu, Y., Zheng, Z., Visscher, P. M. & Yang, J. Quantifying the mapping precision of genome-wide association studies using whole-genome sequencing data. Genome Biol. 18, 86 (2017).
Hou, K. et al. Causal effects on complex traits are similar for common variants across segments of different continental ancestries within admixed individuals. Nat. Genet. 55, 549–558 (2023).
Yengo, L. et al. A saturated map of common genetic variants associated with human height. Nature 610, 704–712 (2022).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Graham, S. E. et al. The power of genetic diversity in genome-wide association studies of lipids. Nature 600, 675–679 (2021).
Trubetskoy, V. et al. Mapping genomic loci implicates genes and synaptic biology in schizophrenia. Nature 604, 502–508 (2022).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and in regions under strong background selection. Nat. Genet. 50, 381–389 (2018).
de Lange, K. M. et al. Genome-wide association study implicates immune activation of multiple integrin genes in inflammatory bowel disease. Nat. Genet. 49, 256–261 (2017).
Lee, S. H., Wray, N. R., Goddard, M. E. & Visscher, P. M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Yang, J., Wray, N. R. & Visscher, P. M. Comparing apples and oranges: equating the power of case-control and quantitative trait association studies. Genet. Epidemiol. 34, 254–257 (2010).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Novo, I., López-Cortegano, E. & Caballero, A. Highly pleiotropic variants of human traits are enriched in genomic regions with strong background selection. Hum. Genet. 140, 1343–1351 (2021).
Claussnitzer, M. et al. FTO obesity variant circuitry and adipocyte browning in humans. N. Engl. J. Med. 373, 895–907 (2015).
Yang, J. et al. Conditional and joint multiple-SNP analysis of GWAS summary statistics identifies additional variants influencing complex traits. Nat. Genet. 44, 369–375 (2012).
Li, A. et al. mBAT-combo: a more powerful test to detect gene-trait associations from GWAS data. Am. J. Hum. Genet. 110, 30–43 (2023).
Li, S. et al. The schizophrenia-associated missense variant rs13107325 regulates dendritic spine density. Transl. Psychiatry 12, 361 (2022).
Singh, T. et al. Rare coding variants in ten genes confer substantial risk for schizophrenia. Nature 604, 509–516 (2022).
Luo, J. et al. Genetic regulation of human brain proteome reveals proteins implicated in psychiatric disorders. Mol. Psychiatry 29, 3330–3343 (2024).
Liu, J., Li, X. & Luo, X.-J. Proteome-wide association study provides insights into the genetic component of protein abundance in psychiatric disorders. Biol. Psychiatry 90, 781–789 (2021).
Bryois, J. et al. Cell-type-specific cis-eQTLs in eight human brain cell types identify novel risk genes for psychiatric and neurological disorders. Nat. Neurosci. 25, 1104–1112 (2022).
Sullivan, P. F., Yao, S. & Hjerling-Leffler, J. Schizophrenia genomics: genetic complexity and functional insights. Nat. Rev. Neurosci. 25, 611–624 (2024).
Zhang, M. J. et al. Pervasive correlations between causal disease effects of proximal SNPs vary with functional annotations and implicate stabilizing selection. Preprint at medRxiv https://doi.org/10.1101/2023.12.04.23299391 (2023).
Stephens, M. False discovery rates: a new deal. Biostatistics 18, 275–294 (2016).
Gazal, S. et al. Functional architecture of low-frequency variants highlights strength of negative selection across coding and non-coding annotations. Nat. Genet. 50, 1600–1607 (2018).
Berisa, T. & Pickrell, J. K. Approximately independent linkage disequilibrium blocks in human populations. Bioinformatics 32, 283–285 (2015).
Casella, G. & Robert, C. P. Rao-Blackwellisation of sampling schemes. Biometrika 83, 81–94 (1996).
Zhu, X. & Stephens, M. Bayesian large-scale multiple regression with summary statistics from genome-wide association studies. Ann. Appl. Stat. 11, 1561–1592 (2017).
Andrew, G. & Donald, B. R. Inference from iterative simulation using multiple sequences. Stat. Sci. 7, 457–472 (1992).
Zhu, Z. et al. Integration of summary data from GWAS and eQTL studies predicts complex trait gene targets. Nat. Genet. 48, 481–487 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Wu, Y. Genome-wide fine-mapping improves identification of causal variants. Zenodo https://doi.org/10.5281/zenodo.15293206 (2026).
Wu, Y. GCTB software to perform genome-wide fine-mapping analysis. Zenodo https://doi.org/10.5281/zenodo.15238830 (2026).
Acknowledgements
This research was supported by the Fundamental Research Funds for the Central Universities (to Y.W.), the 1·3·5 project for disciplines of excellence, West China Hospital, Sichuan University (ZYYC24006 to Y.W.), National Supercomputing Center in Chengdu, the Australian National Health and Medical Research Council (1177268 and 2043111 to J.Z., 1113400 to P.M.V., and 1173790 to N.R.W.) and the Australian Research Council (FL180100072 to P.M.V., DP220101947 to J.Z. and DP230101352 to M.E.G.). P.M.V. acknowledges funding from the European Research Council (grant no. 101198904). L.Y. is supported by the Australian Research Council (grant no. FT220100069) and the Snow Medical Research Foundation. Numerical computations were performed on Hefei advanced computing center and the clusters from the Research Computing Centre (RCC) at The University of Queensland. This study makes use of data from the UKB (project ID: 12505).
Author information
Authors and Affiliations
Contributions
J.Z. conceived and supervised the study. J.Z. and Y.W. developed the methods. J.Z., P.M.V. and Y.W. designed the experiment. Y.W. conducted all analyses with the assistance or guidance from J.Z., Z.Z., L.T., T.L., Q.F., H.C., L.Y., P.M.V., N.R.W. and M.E.G. Y.W., and J.Z. wrote the manuscript with the participation of all authors. All the authors approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Out-of-sample prediction accuracy using identified variants from different fine-mapping methods.
The comparison of results for different methods in the simulation was based on the sparse genetic architecture. The reported power was calculated based on 100 simulation replicates. Data are presented as mean values +/- standard deviation. The number above each bar is the average number of fine-mapped SNPs across 100 simulation replicates from each method at different PIP cut-offs.
Extended Data Fig. 2 Comparison between GWAS P-values and the proportion of SNP-based heritability explained (PHE) for local credible sets associated with Crohn’s disease and a simulated trait.
a, GWAS Manhattan plot for Crohn’s disease, with red triangles indicating SNPs with smallest P-value within each 0.9-LCSs identified by GWFM. b, GWAS Manhattan plot for a simulated trait, where red triangles indicate local credible set includes at least a true causal variant and the grey triangles represent sets without any simulated causal variants. c,d, Proportion of SNP-based heritability explained (PHE) for the identified 0.9-LCSs (orange triangles) and for SNPs not included in any credible set for Crohn’s disease (c) and a simulated trait (d). The genomic position of each local credible set is mapped to the SNP with highest PIP within the set. The GWAS P values are based on two-sided t-test and adjustments are made for multiple comparisons.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–36, Tables 1 and 2 and Notes 1–14.
Supplementary Tables (download XLSX )
Supplementary Tables 3–5.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wu, Y., Zheng, Z., Thibaut, L. et al. Genome-wide fine-mapping improves identification of causal variants. Nat Genet (2026). https://doi.org/10.1038/s41588-026-02549-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41588-026-02549-3