Abstract
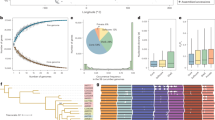
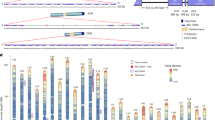
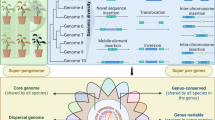
Pangenomes are increasingly important for harnessing crop genetic diversity, yet their resolution and utility are often limited by insufficient sampling of high-quality genome assemblies. Here we present a population-level watermelon super-pangenome constructed from 138 reference-grade assemblies, including 135 newly generated genomes representing all seven species. This super-pangenome captures approximately 1 million structural variants (SVs), enabling accurate variant genotyping across 914 accessions. Broader sampling within the pangenome provides insights into watermelon genome evolution and the origin of cultivated watermelon. Incorporating SVs into genome-wide association studies improves mapping resolution and reveals a copy number variant upstream of ClFCI1 that regulates flesh color intensity in a dosage-dependent manner. Leveraging this comprehensive variation map, we developed high-accuracy genomic prediction models for 18 agronomic traits. Together, these findings and genomic resources establish a foundation for dissecting complex traits and accelerating precision breeding in watermelon, while offering a valuable model for SV-resolved pangenomics in crops.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
Raw HiFi, ONT, Hi–C and genome resequencing reads, as well as genome assemblies, have been deposited in the NCBI BioProject database under accession PRJNA1272048. Individual accession numbers for genome assemblies and raw sequencing datasets are provided in Supplementary Tables 1 and 10. Genome assemblies and annotations, and variant files in VCF format are available at CuGenDBv2 (http://cucurbitgenomics.org/v2/ftp/pan-genome/watermelon/graph_pangenome/).
Code availability
No custom code was generated in this study. All software used in this study is publicly or commercially available as described in the Methods and Reporting Summary.
References
Renner, S. S. How Russian breeders discovered Citrullus mucosospermus and Citrullus lanatus var. cordophanus, the likely closest relatives of domesticated watermelon. Cucurbit Genet. Coop. Rep. 42, 10–12 (2019).
Wu, S. et al. A Citrullus genus super-pangenome reveals extensive variations in wild and cultivated watermelons and sheds light on watermelon evolution and domestication. Plant Biotechnol. J. 21, 1926–1928 (2023).
Paris, H. S. Origin and emergence of the sweet dessert watermelon, Citrullus lanatus. Ann. Bot. 116, 133–148 (2015).
Zhang, Y. et al. Telomere-to-telomere Citrullus super-pangenome provides direction for watermelon breeding. Nat. Genet. 56, 1750–1761 (2024).
Grumet, R., Katzir, N. & Garcia-Mas, J. (eds) Genetics and Genomics of Cucurbitaceae 87–110 (Springer, 2017).
Guo, S. et al. Resequencing of 414 cultivated and wild watermelon accessions identifies selection for fruit quality traits. Nat. Genet. 51, 1616–1623 (2019).
Renner, S. S., Sousa, A. & Chomicki, G. Chromosome numbers, Sudanese wild forms, and classification of the watermelon genus Citrullus, with 50 names allocated to seven biological species. Taxon 66, 1393–1405 (2017).
Zhang, L. et al. An update on genomics and molecular breeding in watermelon. Acta Hortic. 1411, 305–318 (2024).
Zhou, Y. et al. Graph pangenome captures missing heritability and empowers tomato breeding. Nature 606, 527–534 (2022).
Groza, C. et al. Pangenome graphs improve the analysis of structural variants in rare genetic diseases. Nat. Commun. 15, 657 (2024).
Khan, A. W. et al. Super-pangenome by integrating the wild side of a species for accelerated crop improvement. Trends Plant Sci. 25, 148–158 (2020).
Igolkina, A. A. et al. A comparison of 27 Arabidopsis thaliana genomes and the path toward an unbiased characterization of genetic polymorphism. Nat. Genet. 57, 2289–2301 (2025).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biol. 21, 245 (2020).
Levin, D. A. The Role of Chromosomal Change in Plant Evolution (Oxford University Press, 2002).
Wellenreuther, M. & Bernatchez, L. Eco-evolutionary genomics of chromosomal inversions. Trends Ecol. Evol. 33, 427–440 (2018).
Deng, Y. et al. A telomere-to-telomere gap-free reference genome of watermelon and its mutation library provide important resources for gene discovery and breeding. Mol. Plant 15, 1268–1284 (2022).
Renner, S. S. et al. A chromosome-level genome of a Kordofan melon illuminates the origin of domesticated watermelons. Proc. Natl Acad. Sci. USA 118, e2101486118 (2021).
Zhang, J. et al. A unique chromosome translocation disrupting ClWIP1 leads to gynoecy in watermelon. Plant J. 101, 265–277 (2020).
Jiao, D. et al. Identification of allelic relationship and translocation region among chromosomal translocation lines that leads to less-seed watermelon. Hortic. Res. 11, uhae087 (2024).
Branham, S. E. et al. QTL mapping of resistance to Fusarium oxysporum f. sp. niveum race 2 and Papaya ringspot virus in Citrullus amarus. Theor. Appl. Genet. 133, 677–687 (2020).
Gao, L. Transcriptome Analysis and Fine Mapping of Major Genes Controlling Flesh Firmness and Sour Flesh in Watermelon (Chinese Academy of Agricultural Sciences, 2018).
Ren, Y. et al. An integrated genetic map based on four mapping populations and quantitative trait loci associated with economically important traits in watermelon (Citrullus lanatus). BMC Plant Biol. 14, 33 (2014).
Maples, B. K., Gravel, S., Kenny, E. E. & Bustamante, C. D. RFMix: a discriminative modeling approach for rapid and robust local-ancestry inference. Am. J. Hum. Genet. 93, 278–288 (2013).
Connallon, T. & Olito, C. Natural selection and the distribution of chromosomal inversion lengths. Mol. Ecol. 31, 3627–3641 (2022).
Schaefer, H., Heibl, C. & Renner, S. S. Gourds afloat: a dated phylogeny reveals an Asian origin of the gourd family (Cucurbitaceae) and numerous oversea dispersal events. Proc. Biol. Sci. 276, 843–851 (2009).
Meyer, R. S. & Purugganan, M. D. Evolution of crop species: genetics of domestication and diversification. Nat. Rev. Genet. 14, 840–852 (2013).
Pérez-Escobar, O. A. et al. Genome sequencing of up to 6,000-year-old Citrullus seeds reveals use of a bitter-fleshed species prior to watermelon domestication. Mol. Biol. Evol. 39, msac168 (2022).
Martin, S. H., Davey, J. W. & Jiggins, C. D. Evaluating the use of ABBA–BABA statistics to locate introgressed loci. Mol. Biol. Evol. 32, 244–257 (2015).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
Andolfo, G., Sánchez, C. S., Cañizares, J., Pico, M. B. & Ercolano, M. R. Large-scale gene gains and losses molded the NLR defense arsenal during the Cucurbita evolution. Planta 254, 82 (2021).
Schemberger, M. O. et al. Transcriptome profiling of non-climacteric ‘yellow’ melon during ripening: insights on sugar metabolism. BMC Genomics 21, 262 (2020).
Dou, J. L. et al. Genetic mapping reveals a candidate gene (ClFS1) for fruit shape in watermelon (Citrullus lanatus L.). Theor. Appl. Genet. 131, 947–958 (2018).
Katuuramu, D. N., Levi, A. & Wechter, W. P. Genome-wide association study of soluble solids content, flesh color, and fruit shape in citron watermelon. Plant Genome 16, e20391 (2023).
Wang, J. et al. A natural variant of NON-RIPENING promotes fruit ripening in watermelon. Plant Cell 37, koae313 (2024).
Yin, J. et al. A single amino acid substitution in the AAA-type ATPase LRD6-6 activates immune responses but decreases grain quality in rice. Front. Plant Sci. 15, 1451897 (2024).
Zhang, B. et al. The mitochondrial outer membrane AAA ATPase AtOM66 affects cell death and pathogen resistance in Arabidopsis thaliana. Plant J. 80, 709–727 (2014).
Waldo, B. D., Branham, S. E., Levi, A., Patrick Wechter, W. & Rutter, W. B. Distinct genomic loci underlie quantitative resistance to Meloidogyne enterolobii galling and reproduction in Citrullus amarus. Plant Dis. 107, 2126–2132 (2023).
Sun, T. et al. ChIP–seq reveals broad roles of SARD1 and CBP60g in regulating plant immunity. Nat. Commun. 6, 10159 (2015).
Larkin, R. M. et al. Reduced chloroplast coverage genes from Arabidopsis thaliana help to establish the size of the chloroplast compartment. Proc. Natl Acad. Sci. USA 113, E1116–E1125 (2016).
Crossa, J. et al. Genomic selection in plant breeding: methods, models, and perspectives. Trends Plant Sci. 22, 961–975 (2017).
Yan, J. et al. LightGBM: accelerated genomically designed crop breeding through ensemble learning. Genome Biol. 22, 271 (2021).
Liu, S. et al. Nucleotide variation in the phytoene synthase (ClPsy1) gene contributes to golden flesh in watermelon (Citrullus lanatus L.). Theor. Appl. Genet. 135, 185–200 (2022).
Branham, S. et al. Genetic mapping of a major codominant QTL associated with β-carotene accumulation in watermelon. Mol. Breed. 37, 146 (2017).
Nie, H. et al. High-quality genome assembly and genetic mapping reveal a gene regulating flesh color in watermelon (Citrullus lanatus). Front. Plant Sci. 14, 1142856 (2023).
Zhang, J. et al. Decreased protein abundance of lycopene β-cyclase contributes to red flesh in domesticated watermelon. Plant Physiol. 183, 1171–1183 (2020).
Fang, X. F. et al. Clorf encodes carotenoid isomerase and regulates orange flesh color in watermelon (Citrullus lanatus L.). J. Agric. Food Chem. 71, 15445–15455 (2023).
Liu, S. et al. Identification of chromosome region and candidate genes for canary-yellow flesh (Cyf) locus in watermelon (Citrullus lanatus). Plant Sci. 329, 111594 (2023).
Zhang, W. et al. Clpf encodes pentatricopeptide repeat protein (PPR5) and regulates pink flesh color in watermelon (Citrullus lanatus L.). Theor. Appl. Genet. 137, 126 (2024).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Sim, S. B., Corpuz, R. L., Simmonds, T. J. & Geib, S. M. HiFiAdapterFilt, a memory efficient read processing pipeline, prevents occurrence of adapter sequence in PacBio HiFi reads and their negative impacts on genome assembly. BMC Genomics 23, 157 (2022).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Guan, D. et al. Identifying and removing haplotypic duplication in primary genome assemblies. Bioinformatics 36, 2896–2898 (2020).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi–C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol. 23, 258 (2022).
Lin, Y. et al. quarTeT: a telomere-to-telomere toolkit for gap-free genome assembly and centromeric repeat identification. Hortic. Res. 10, uhad127 (2023).
Guo, S. G. et al. The draft genome of watermelon (Citrullus lanatus) and resequencing of 20 diverse accessions. Nat. Genet. 45, 51–58 (2013).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome Biol. 20, 275 (2019).
Cantarel, B. L. et al. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-seq data without a reference genome. Nat. Biotechnol. 29, 644–652 (2011).
Li, Q. et al. A chromosome-scale genome assembly of cucumber (Cucumis sativus L.). Gigascience 8, giz072 (2019).
Castanera, R., Ruggieri, V., Pujol, M., Garcia-Mas, J. & Casacuberta, J. M. An improved melon reference genome with single-molecule sequencing uncovers a recent burst of transposable elements with potential impact on genes. Front. Plant Sci. 10, 1815 (2020).
Sun, H. et al. Karyotype stability and unbiased fractionation in the paleo-allotetraploid Cucurbita genomes. Mol. Plant 10, 1293–1306 (2017).
Fu, A. et al. Combined genomic, transcriptomic, and metabolomic analyses provide insights into chayote (Sechium edule) evolution and fruit development. Hortic. Res. 8, 35 (2021).
Ma, L. et al. The genome and transcriptome analysis of snake gourd provide insights into its evolution and fruit development and ripening. Hortic. Res. 7, 199 (2020).
Xie, D. et al. The wax gourd genomes offer insights into the genetic diversity and ancestral cucurbit karyotype. Nat. Commun. 10, 5158 (2019).
Cheng, C. Y. et al. Araport11: a complete reannotation of the Arabidopsis thaliana reference genome. Plant J. 89, 789–804 (2017).
The UniProt Consortium. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res. 49, D480–D489 (2021).
Stanke, M., Tzvetkova, A. & Morgenstern, B. AUGUSTUS at EGASP: using EST, protein and genomic alignments for improved gene prediction in the human genome. Genome Biol. 7, S11 (2006).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biol. 9, R7 (2008).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Gotz, S. et al. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 36, 3420–3435 (2008).
Li, P. et al. RGAugury: a pipeline for genome-wide prediction of resistance gene analogs (RGAs) in plants. BMC Genomics 17, 852 (2016).
Song, B. et al. AnchorWave: sensitive alignment of genomes with high sequence diversity, extensive structural polymorphism, and whole-genome duplication. Proc. Natl Acad. Sci. USA 119, e2113075119 (2022).
Lovell, J. T. et al. GENESPACE tracks regions of interest and gene copy number variation across multiple genomes. Elife 11, e78526 (2022).
Pickrell, J. K. & Pritchard, J. K. Inference of population splits and mixtures from genome-wide allele frequency data. PLoS Genet. 8, e1002967 (2012).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49 (2012).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Wu, S. et al. The bottle gourd genome provides insights into Cucurbitaceae evolution and facilitates mapping of a Papaya ring-spot virus resistance locus. Plant J. 92, 963–975 (2017).
Matsumura, H. et al. Long-read bitter gourd (Momordica charantia) genome and the genomic architecture of nonclassic domestication. Proc. Natl Acad. Sci. USA 117, 14543–14551 (2020).
Marrano, A. et al. High-quality chromosome-scale assembly of the walnut (Juglans regia L.) reference genome. Gigascience 9, giaa050 (2020).
Abascal, F., Zardoya, R. & Telford, M. J. TranslatorX: multiple alignment of nucleotide sequences guided by amino acid translations. Nucleic Acids Res. 38, W7–W13 (2010).
Bouckaert, R. et al. BEAST 2: a software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537 (2014).
Terhorst, J., Kamm, J. A. & Song, Y. S. Robust and scalable inference of population history from hundreds of unphased whole genomes. Nat. Genet. 49, 303–309 (2017).
Jeong, S. et al. GenoCore: a simple and fast algorithm for core subset selection from large genotype datasets. PLoS ONE 12, e0181420 (2017).
Wu, S. et al. Genome of ‘Charleston Gray’, the principal American watermelon cultivar, and genetic characterization of 1,365 accessions in the U.S. National Plant Germplasm System watermelon collection. Plant Biotechnol. J. 17, 2246–2258 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Khelik, K., Lagesen, K., Sandve, G. K., Rognes, T. & Nederbragt, A. J. NucDiff: in-depth characterization and annotation of differences between two sets of DNA sequences. BMC Bioinformatics 18, 338 (2017).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, giab008 (2021).
Ebler, J. et al. Pangenome-based genome inference allows efficient and accurate genotyping across a wide spectrum of variant classes. Nat. Genet. 54, 518–525 (2022).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Lippert, C. et al. FaST linear mixed models for genome-wide association studies. Nat. Methods 8, 833–835 (2011).
Li, M.-X., Yeung, J. M. Y., Cherny, S. S. & Sham, P. C. Evaluating the effective numbers of independent tests and significant p-value thresholds in commercial genotyping arrays and public imputation reference datasets. Hum. Genet. 131, 747–756 (2012).
Mansfeld, B. N. & Grumet, R. QTLseqr: an R package for bulk segregant analysis with next-generation sequencing. Plant Genome 11, 180006 (2018).
Meng, L., Li, H., Zhang, L. & Wang, J. QTL IciMapping: integrated software for genetic linkage map construction and quantitative trait locus mapping in biparental populations. Crop J. 3, 269–283 (2015).
Curtis, M. D. & Grossniklaus, U. A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133, 462–469 (2003).
Zhang, J. et al. High-level expression of a novel chromoplast phosphate transporter ClPHT4;2 is required for flesh color development in watermelon. New Phytol. 213, 1208–1221 (2017).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Acknowledgements
The authors thank S.S. Renner from University of Munich for providing seeds of C. lanatus subsp. cordophanus. This research was supported by the Beijing Rural Revitalization Agricultural Science and Technology Project (grant NY2401130025 to Y.X.), the National Natural Science Foundation of China (grants 32172592 to J. Zhang and 32330093 to Y.X.), the Scientific and Technological Innovation Capacity Building Project of BAAFS (KJCX20251008 to J. Zhang), the Scientist Training Program of BAAFS (JKZX202401 to J. Zhang), Ministry of Agriculture of China (CARS-25 to Y.X.) and USDA National Institute of Food and Agriculture Specialty Crop Research Initiative (2015-51181-24285 and 2020-51181-32139 to Z.F., R.G., A.L., C.K., S.E.B., P.W. and C.M.).
Author information
Authors and Affiliations
Contributions
Z.F. and Y.X. designed and supervised the project. S.G., S.A.H., C.M., R.J., S.E.B., P.W., C.K., A.L. and R.G. contributed to sample collection and DNA extraction. S.G., H.S., S. Liao, J. Zhang, R.J. and Z.F. coordinated genome sequencing. S.G., S. Liao, J. Zhang, G.G., J.W., M.L., Y.Y., Y.R., S.T., S. Li and H.Z. performed phenotyping for fruit-quality traits. H.S., Z.Z., X.Z. and S.W. contributed to genome assembly and annotation, as well as pangenome and population genetic analyses. H.S. and J. Zhao conducted the genomic prediction analysis. J. Zhang, H.S. and S. Liao contributed to genetic mapping and gene functional characterization. H.S., Z.Z., J. Zhang, X.Z. and S.W. wrote the manuscript. Z.F. and Y.X. revised the manuscript.
Corresponding authors
Ethics declarations
Competing interests
All authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks Michael Bevan, Hongbo Li, and the other anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Closely located large inversions on chromosome 6 encompassing QTLs associated with Fusarium wilt resistance, flesh sweetness and flesh firmness.
a, A genomic region on chromosome 6 (green box) in cultivated watermelon introgressed from C. amarus, overlapping with three QTLs related to Fusarium wilt resistance (qFon2-6), flesh sweetness (QBrix6), and flesh firmness. b, Alignments of chromosome 6 between reference ‘97103’ and the accessions ‘SugarleeXZ’, ‘PI 264341-FR’, and ‘USVL252’, all of which harbor resistance to Fusarium wilt.
Extended Data Fig. 2 Haplotypes of the fruit shape-related gene ClFS1.
a, Structure diagram of the missense SNP (ClFS1Hap2) and 159-bp deletion (ClFS1Hap3) in ClFS1. Representative fruits carrying the corresponding haplotypes are shown. ClFS1Hap1 denotes the reference allele from ‘97103’. FSI, fruit shape index. Scale bars, 10 cm. b, Allele frequencies of the three haplotypes across Citrullus groups. c, Effect of different haplotypes of ClFS1 on fruit shape. In c, P values were calculated using two-sided Student’s t-test.
Extended Data Fig. 3 Genetic mapping of flesh color intensity in watermelon.
a, Genome-wide ΔSNP-index profile from bulked-segregant analysis (BSA) of an F2 population derived from the cross ‘Ming 58’ (scarlet flesh) x ‘JX2’ (pink flesh). The black curve represents ΔSNP-index values, while red and blue envelopes mark the 95% and 99% confidence thresholds, respectively. b,c, Recombinant-based fine-mapping of the ClFCI1 locus in 141 F2 plants from the ‘Ming 58’ x ‘JX2’ cross (b), and 636 F2 plants from the ‘JLM’ x ‘Cream of Saskatchewan’ (CS) cross (c). Top, fruits of parental lines and F1 hybrids. Scale bars, 10 cm. Middle, QTL curves showing logarithm of odds (LOD) scores using KASP markers between 23 Mb and 26 Mb on chromosome 6. Bottom, graphical genotypes of key recombinant individuals. Pink, red, and scarlet segments denote homozygous ‘JX2’, heterozygous, and homozygous ‘Ming 58’ regions, respectively; pale yellow, orange, and yellow segments denote homozygous ‘CS’, heterozygous, and homozygous ‘JLM’ regions, respectively. d, Relative expression levels of candidate genes in the four parental lines (‘Ming 58’, ‘JX2’, ‘JLM’, and ‘CS’). Error bars indicate the standard deviation of three biological replicates. Different lowercase letters indicate significant differences according to Turkey’s multiple range test (P < 0.05).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–5.
Supplementary Tables (download XLSX )
Supplementary Tables 1–20.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Sun, H., Zhang, J., Liao, S. et al. Population-level super-pangenome reveals genome evolution and empowers precision breeding in watermelon. Nat Genet (2026). https://doi.org/10.1038/s41588-026-02598-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41588-026-02598-8