Abstract
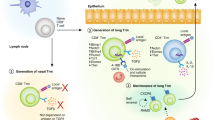
Tissue-resident memory T cells (TRM cells) are critical for cellular immunity to respiratory pathogens and reside in both the airways and the interstitium. In the present study, we found that the airway environment drove transcriptional and epigenetic changes that specifically regulated the cytolytic functions of airway TRM cells and promoted apoptosis due to amino acid starvation and activation of the integrated stress response. Comparison of airway TRM cells and splenic effector-memory T cells transferred into the airways indicated that the environment was necessary to activate these pathways, but did not induce TRM cell lineage reprogramming. Importantly, activation of the integrated stress response was reversed in airway TRM cells placed in a nutrient-rich environment. Our data defined the genetic programs of distinct lung TRM cell populations and show that local environmental cues altered airway TRM cells to limit cytolytic function and promote cell death, which ultimately leads to fewer TRM cells in the lung.
This is a preview of subscription content, access via your institution
Access options







Similar content being viewed by others
Data availability
All sequencing data are available from the National Center for Biotechnology Information Gene Expression Omnibus under accession no. GSE118112. All code, data processing scripts and additional data that support the findings of this study are available from the corresponding author upon request.
References
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Schenkel, J. M., Fraser, K. A., Vezys, V. & Masopust, D. Sensing and alarm function of resident memory CD8+ T cells. Nat. Immunol. 14, 509–513 (2013).
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ TRM cells providing global skin immunity. Nature 483, 227–231 (2012).
Kohlmeier, J. E., Cookenham, T., Roberts, A. D., Miller, S. C. & Woodland, D. L. Type I interferons regulate cytolytic activity of memory CD8+ T cells in the lung airways during respiratory virus challenge. Immunity 33, 96–105 (2010).
Mackay, L. K. et al. T-box transcription factors combine with the cytokines TGF-β and IL-15 to control tissue-resident memory T cell fate. Immunity 43, 1101–1111 (2015).
Mackay, L. K. et al. Hobit and Blimp1 instruct a universal transcriptional program of tissue residency in lymphocytes. Science 352, 459–463 (2016).
Liang, S. H., Mozdzanowska, K., Palladino, G. & Gerhard, W. Heterosubtypic immunity to influenza type-A virus in mice—effector mechanisms and their longevity. J. Immunol. 152, 1653–1661 (1994).
Wu, T. et al. Lung-resident memory CD8 T cells (TRM) are indispensable for optimal cross-protection against pulmonary virus infection. J. Leukoc. Biol. 95, 215–224 (2014).
Purwar, R. et al. Resident memory T cells (TRM) are abundant in human lung: diversity, function, and antigen specificity. PLoS One 6, e16245 (2011).
Lee, L. Y. et al. Memory T cells established by seasonal human influenza A infection cross-react with avian influenza A (H5N1) in healthy individuals. J. Clin. Invest. 118, 3478–3490 (2008).
Hogan, R. J. et al. Activated antigen-specific CD8+ T cells persist in the lungs following recovery from respiratory virus infections. J. Immunol. 166, 1813–1822 (2001).
Hou, S., Doherty, P. C., Zijlstra, M., Jaenisch, R. & Katz, J. M. Delayed clearance of Sendai virus in mice lacking class I MHC-restricted CD8+ T cells. J. Immunol. 149, 1319–1325 (1992).
Eichelberger, M., Allan, W., Zijlstra, M., Jaenisch, R. & Doherty, P. C. Clearance of influenza virus respiratory infection in mice lacking class I major histocompatibility complex-restricted CD8+ T cells. J. Exp. Med. 174, 875–880 (1991).
McMaster, S. R. et al. Pulmonary antigen encounter regulates the establishment of tissue-resident CD8 memory T cells in the lung airways and parenchyma. Mucosal Immunol. 11, 1071–1078 (2018).
McMaster, S. R., Wilson, J. J., Wang, H. & Kohlmeier, J. E. Airway-resident memory CD8 T cells provide antigen-specific protection against respiratory virus challenge through rapid IFN-γ production. J. Immunol. 195, 203–209 (2015).
Slutter, B. et al. Dynamics of influenza-induced lung-resident memory T cells underlie waning heterosubtypic immunity. Sci. Immunol. 2, eaag2031 (2017).
Farber, D. L., Yudanin, N. A. & Restifo, N. P. Human memory T cells: generation, compartmentalization and homeostasis. Nat. Rev. Immunol. 14, 24–35 (2014).
Kumar, B. V. et al. Human tissue-resident memory T cells are defined by core transcriptional and functional signatures in lymphoid and mucosal sites. Cell Rep. 20, 2921–2934 (2017).
Hombrink, P. et al. Programs for the persistence, vigilance and control of human CD8+ lung-resident memory T cells. Nat. Immunol. 17, 1467–1478 (2016).
Hikono, H. et al. Activation phenotype, rather than central- or effector-memory phenotype, predicts the recall efficacy of memory CD8+ T cells. J. Exp. Med. 204, 1625–1636 (2007).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Buenrostro, J. D., Giresi, P. G., Zaba, L. C., Chang, H. Y. & Greenleaf, W. J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Scharer, C. D. et al. ATAC-seq on biobanked specimens defines a unique chromatin accessibility structure in naive SLE B cells. Sci. Rep. 6, 27030 (2016).
Barwick, B. G., Scharer, C. D., Bally, A. P. & Boss, J. M. Plasma cell differentiation is coupled to division-dependent DNA hypomethylation and gene regulation. Nat. Immunol. 17, 1216–1225 (2016).
Scharer, C. D., Bally, A. P., Gandham, B. & Boss, J. M. Cutting edge: chromatin accessibility programs CD8 T cell memory. J. Immunol. 198, 2238–2243 (2017).
Pakos-Zebrucka, K. et al. The integrated stress response. EMBO Rep. 17, 1374–1395 (2016).
de la Ballina, L. R. et al. Amino acid transport associated to cluster of differentiation 98 heavy chain (CD98hc) is at the cross-road of oxidative stress and amino acid availability. J. Biol. Chem. 291, 9700–9711 (2016).
Ely, K. H., Cookenham, T., Roberts, A. D. & Woodland, D. L. Memory T cell populations in the lung airways are maintained by continual recruitment. J. Immunol. 176, 537–543 (2006).
Li, T. et al. DDIT3 and KAT2A proteins regulate TNFRSF10A and TNFRSF10B expression in endoplasmic reticulum stress-mediated apoptosis in human lung cancer cells. J. Biol. Chem. 290, 11108–11118 (2015).
Pino, S. C. et al. CHOP mediates endoplasmic reticulum stress-induced apoptosis in Gimap5-deficient T cells. PLoS One 4, e5468 (2009).
Teske, B. F. et al. CHOP induces activating transcription factor 5 (ATF5) to trigger apoptosis in response to perturbations in protein homeostasis. Mol. Biol. Cell 24, 2477–2490 (2013).
Iyer, V. R. et al. The transcriptional program in the response of human fibroblasts to serum. Science 283, 83–87 (1999).
Kohlmeier, J. E., Miller, S. C. & Woodland, D. L. Cutting edge: antigen is not required for the activation and maintenance of virus-specific memory CD8+ T cells in the lung airways. J. Immunol. 178, 4721–4725 (2007).
Takamura, S. et al. Specific niches for lung-resident memory CD8+ T cells at the site of tissue regeneration enable CD69-independent maintenance. J. Exp. Med. 213, 3057–3073 (2016).
Pizzolla, A. et al. Resident memory CD8+ T cells in the upper respiratory tract prevent pulmonary influenza virus infection. Sci. Immunol. 2, eaam6970 (2017).
Caminschi, I., Lahoud, M. H., Pizzolla, A. & Wakim, L. M. Zymosan by-passes the requirement for pulmonary antigen encounter in lung tissue-resident memory CD8+ T cell development. Mucosal Immunol. 12, 403–412 (2019).
Van Braeckel-Budimir, N., Varga, S. M., Badovinac, V. P. & Harty, J. T. Repeated antigen exposure extends the durability of influenza-specific lung-resident memory CD8+ T cells and heterosubtypic immunity. Cell Rep. 24, 3374–3382.e3 (2018).
Zhou, A. C., Wagar, L. E., Wortzman, M. E. & Watts, T. H. Intrinsic 4-1BB signals are indispensable for the establishment of an influenza-specific tissue-resident memory CD8 T-cell population in the lung. Mucosal Immunol. 10, 1294–1309 (2017).
Mintern, J. D., Guillonneau, C., Carbone, F. R., Doherty, P. C. & Turner, S. J. Cutting edge: tissue-resident memory CTL down-regulate cytolytic molecule expression following virus clearance. J. Immunol. 179, 7220–7224 (2007).
Kohlmeier, J. E. et al. Inflammatory chemokine receptors regulate CD8+ T cell contraction and memory generation following infection. J. Exp. Med. 208, 1621–1634 (2011).
Kohlmeier, J. E. et al. The chemokine receptor CCR5 plays a key role in the early memory CD8+ T cell response to respiratory virus infections. Immunity 29, 101–113 (2008).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Hsu, F. et al. The UCSC Known Genes. Bioinformatics 22, 1036–1046 (2006).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Huang da, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Acknowledgements
We thank the New York University Genome Technology Center and University of Albama at Birmingham Helfin Genomics Core for Illumina sequencing, the Emory Integrated Genomics Core for sequencing library Bioanalyzer expertise, Children’s Healthcare of Atlanta and Emory University Pediatric Flow Cytometry Core for cell sorting and the NIH Tetramer Core Facility (contract no. HHSN272201300006C). This project was supported by NIH grants (nos. R01HL122559 and R01HL138508) and Centers of Excellence in Influenza Research and Surveillance contracts (no. HHSN272201400004C (to J.E.K.), and nos. 1R01AI113021 and P01AI125180-01 (to J.M.B.)). S.L.H. was supported by an NIH grant (no. F31 HL136101).
Author information
Authors and Affiliations
Contributions
S.L.H., C.D.S., J.M.B. and J.E.K. designed the study. S.L.H., C.D.S., E.K.C., Z.-R.T.L. and S.T. performed the experiments. S.L.H., C.D.S., E.K.C., Z.-R.T.L., S.T. and J.E.K. analyzed the data. S.L.H., C.D.S., J.M.B. and J.E.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information I. Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Fig. 1 Sendai-specific CD8 TRM cells in the airways and interstitium gradually decline over time.
(a) Example staining reflecting Sendai NP (SenNP) tetramer staining of BAL, interstitium, and spleen on days 30 and 90 post-Sendai virus infection. (b) Number of SenNP+ CD8+ T cells in the airway (A-TRM), lung interstitium (I-TRM), and spleen (S-TEM) following infection with Sendai virus. n=10 for all timepoints. Data represented as mean ± SEM.
Supplementary Fig. 2 Lung TRM cells decline more rapidly than circulating TEM cells.
(a) Staining of CXCR3 and CD62L on FluNP+ S-TEM over time. (b) Number of FluNP+ S-TEM in wild-type mice on days 35, 60, 90, and 180 post-x31 infection. (c) Fold change in S-TEM and subsets of I-TRM FluNP+ cells defined by CD69 and CD103 expression between day 35 and day 60 (left graph) and between day 35 and day 180 (right graph). n=10 for all time points and data are derived from mice in Fig. 1. Data represented as mean ± SEM.
Supplementary Fig. 3 Validation of A-TRM-specific epigenomic and transcriptome profiles.
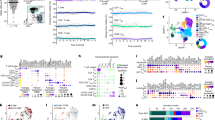
(a) Principal component analysis of 9,970 detected genes in S-TEM (n=3), lung vascular TEM (N=3), I-TRM (N=3) and A-TRM (n=3) FluNP+ CD8+ T cells following RNA-Seq. Points denote samples and circles show 99% confidence intervals for each cell type. (b) Bar plots of FPKM normalized gene expression for the indicated genes. Data represent mean ± SD. (c) Principal component analysis of 31,049 accessible peaks in S-TEM (n=3), lung vascular TEM (N=3), I-TRM (N=3) and A-TRM (n=3) FluNP+ CD8+ T cells following ATAC-Seq. Points denote samples and circles show 99% confidence intervals for each cell type. (d) Genome plot showing the Slc7a5, Asns, and Gzma loci. Accessibility for the indicated sample is shown along with gene structure and transcription direction. Locations of DAR are boxed. Data represent the mean of three replicates for each group. (e) Percent AnnexinV+ among I-TRM and lung vascular TEM FluNP+ CD8+ T cells. N=13 and the I-TRM data are from Fig. 2h. P value: *p<0.05. Data represented as mean ± SEM.
Supplementary Fig. 4 BCL2 is up-regulated in A-TRM compared to I-TRM.
(a) Frequency of BCL2 on FluNP+ CD8+ I-TRM cells and A-TRM cells (n=5). (b) gMFI of BCL2 on FluNP+ CD8+ I-TRM cells and A-TRM cells (n=5). Data represented as mean ± SEM. P value: ** = p<0.01. (c) Example histogram of BCL2 gated on FluNP+ CD8+ I-TRM cells and A-TRM cells.
Supplementary Fig. 5 A-TRM cells from WT and Ddit3-/- mice provide similar protection following influenza challenge.
(a) Experimental design for intratracheal (IT) transfer of WT or Ddit3-/- A-TRM cells into naïve recipient mice. (b) Viral titers measured on day 4 post-challenge in mice receiving WT (n=8) or Ddit3-/- (n=10) A-TRM cells. Data represented as mean ± SD.
Supplementary Fig. 6 Alveolar macrophages do not up-regulate stress response pathways.
Gene Set Enrichment Analysis comparing transcriptome profiles of alveolar versus interstitial macrophages for the indicated gene sets. The FDR q-value for each comparison is indicated.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6.
Rights and permissions
About this article
Cite this article
Hayward, S.L., Scharer, C.D., Cartwright, E.K. et al. Environmental cues regulate epigenetic reprogramming of airway-resident memory CD8+ T cells. Nat Immunol 21, 309–320 (2020). https://doi.org/10.1038/s41590-019-0584-x
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-019-0584-x
This article is cited by
-
Heterogeneity of T cells regulates tumor immunity mediated by Helicobacter pylori infection in gastric cancer
BMC Cancer (2025)
-
Physiological microbial exposure normalizes memory T cell surveillance of the brain and modifies host seizure outcomes
Nature Immunology (2025)
-
Dynamics of pulmonary mucosal cytotoxic CD8 T-cells in people living with HIV under suppressive antiretroviral therapy
Respiratory Research (2024)
-
Prevention of respiratory virus transmission by resident memory CD8+ T cells
Nature (2024)
-
PU.1 and BCL11B sequentially cooperate with RUNX1 to anchor mSWI/SNF to poise the T cell effector landscape
Nature Immunology (2024)