Abstract
Alveolar macrophages (AMs) are lung tissue-resident macrophages that can be expanded in culture, but it is unknown to what extent culture affects their in vivo identity. Here we show that mouse long-term ex vivo expanded AMs (exAMs) maintained a core AM gene expression program, but showed culture adaptations related to adhesion, metabolism and proliferation. Upon transplantation into the lung, exAMs reacquired full transcriptional and epigenetic AM identity, even after several months in culture and could self-maintain long-term in the alveolar niche. Changes in open chromatin regions observed in culture were fully reversible in transplanted exAMs and resulted in a gene expression profile indistinguishable from resident AMs. Our results indicate that long-term proliferation of AMs in culture did not compromise cellular identity in vivo. The robustness of exAM identity provides new opportunities for mechanistic analysis and highlights the therapeutic potential of exAMs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All sequencing data generated in this study have been deposited in the Gene Expression Omnibus (GEO) repository under the accession number GSE194144.
References
Davies, L. C., Jenkins, S. J., Allen, J. E. & Taylor, P. R. Tissue-resident macrophages. Nat. Immunol. 14, 986–995 (2013).
Bleriot, C., Chakarov, S. & Ginhoux, F. Determinants of resident tissue macrophage identity and function. Immunity 52, 957–970 (2020).
Gautier, E. L. et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat. Immunol. 13, 1118–1128 (2012).
Lavin, Y. et al. Tissue-resident macrophage enhancer landscapes are shaped by the local microenvironment. Cell 159, 1312–1326 (2014).
Guilliams, M. & Svedberg, F. R. Does tissue imprinting restrict macrophage plasticity? Nat. Immunol. 22, 118–127 (2021).
Sieweke, M. H. Waddington’s valleys and Captain Cook’s islands. Cell Stem Cell 16, 7–8 (2015).
van de Laar, L. et al. Yolk sac macrophages, fetal liver, and adult monocytes can colonize an empty niche and develop into functional tissue-resident macrophages. Immunity https://doi.org/10.1016/j.immuni.2016.02.017 (2016).
McQuattie-Pimentel, A. C. et al. The lung microenvironment shapes a dysfunctional response of alveolar macrophages in aging. J. Clin. Invest. https://doi.org/10.1172/JCI140299 (2021).
Gosselin, D. et al. Environment drives selection and function of enhancers controlling tissue-specific macrophage identities. Cell 159, 1327–1340 (2014).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science https://doi.org/10.1126/science.aal3222 (2017).
Svedberg, F. R. et al. The lung environment controls alveolar macrophage metabolism and responsiveness in type 2 inflammation. Nat. Immunol. 20, 571–580 (2019).
Bohlen, C. J. et al. Diverse requirements for microglial survival, specification, and function revealed by defined-medium cultures. Neuron 94, 759–773 (2017).
Fuchs, E. The impact of cell culture on stem cell research. Cell Stem Cell 10, 640–641 (2012).
Helft, J. et al. GM-CSF mouse bone marrow cultures comprise a heterogeneous population of CD11c(+)MHCII(+) macrophages and dendritic cells. Immunity 42, 1197–1211 (2015).
Misharin, A. V., Saber, R. & Perlman, H. Eosinophil contamination of thioglycolate-elicited peritoneal macrophage cultures skews the functional readouts of in vitro assays. J. Leukoc. Biol. 92, 325–331 (2012).
Lim, W. A. & June, C. H. The principles of engineering immune. Cells Treat. Cancer Cell 168, 724–740 (2017).
Dolgin, E. Cancer-eating immune cells kitted out with CARs. Nat. Biotechnol. 38, 509–511 (2020).
Aziz, A., Soucie, E., Sarrazin, S. & Sieweke, M. H. MafB/c-Maf deficiency enables self-renewal of differentiated functional macrophages. Science 326, 867–871 (2009).
Soucie, E. L. et al. Lineage-specific enhancers activate self-renewal genes in macrophages and embryonic stem cells. Science 351, aad5510 (2016).
Busch, C. J., Favret, J., Geirsdottir, L., Molawi, K. & Sieweke, M. H. Isolation and long-term cultivation of mouse alveolar macrophages. Bio. Protoc. https://doi.org/10.21769/BioProtoc.3302 (2019).
Aegerter, H. et al. Influenza-induced monocyte-derived alveolar macrophages confer prolonged antibacterial protection. Nat. Immunol. 21, 145–157 (2020).
Schneider, C. et al. Induction of the nuclear receptor PPAR-γ by the cytokine GM-CSF is critical for the differentiation of fetal monocytes into alveolar macrophages. Nat. Immunol. 15, 1026–1037 (2014).
Rauschmeier, R. et al. Bhlhe40 and Bhlhe41 transcription factors regulate alveolar macrophage self-renewal and identity. EMBO J. 38, e101233 (2019).
Angelidis, I. et al. An atlas of the aging lung mapped by single cell transcriptomics and deep tissue proteomics. Nat. Commun. 10, 963 (2019).
Yu, X. et al. The cytokine TGF-β promotes the development and homeostasis of alveolar macrophages. Immunity 47, 903–912 (2017).
Misharin, A. V. et al. Monocyte-derived alveolar macrophages drive lung fibrosis and persist in the lung over the life span. J. Exp. Med. 214, 2387–2404 (2017).
Machiels, B. et al. A gammaherpesvirus provides protection against allergic asthma by inducing the replacement of resident alveolar macrophages with regulatory monocytes. Nat. Immunol. 18, 1310–1320 (2017).
de Laval, B. et al. C/EBPβ-dependent epigenetic memory induces trained immunity in hematopoietic stem cells. Cell Stem Cell 26, 793 (2020).
Happle, C. et al. Pulmonary transplantation of macrophage progenitors as effective and long-lasting therapy for hereditary pulmonary alveolar proteinosis. Sci. Transl. Med. 6, 250ra113 (2014).
Suzuki, T. et al. Pulmonary macrophage transplantation therapy. Nature https://doi.org/10.1038/nature13807 (2014).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Heinz, S., Romanoski, C. E., Benner, C. & Glass, C. K. The selection and function of cell type-specific enhancers. Nat. Rev. Mol. Cell Biol. 16, 144–154 (2015).
Glass, C. K. & Natoli, G. Molecular control of activation and priming in macrophages. Nat. Immunol. 17, 26–33 (2016).
Xue, J. et al. Transcriptome-based network analysis reveals a spectrum model of human macrophage activation. Immunity 40, 274–288 (2014).
Imperatore, F. et al. SIRT1 regulates macrophage self-renewal. EMBO J. 36, 2353–2372 (2017).
Green, H. Cultured cells for the treatment of disease. Sci. Am. 265, 96–102 (1991).
Yao, Y. et al. Induction of autonomous memory alveolar macrophages requires T cell help and is critical to trained immunity. Cell 175, 1634–1650 (2018).
Bekkering, S. et al. Metabolic induction of trained immunity through the mevalonate pathway. Cell 172, 135–146 (2018).
Saeed, S. et al. Epigenetic programming of monocyte-to-macrophage differentiation and trained innate immunity. Science 345, 1251086 (2014).
Cheng, S. C. et al. mTOR- and HIF-1α-mediated aerobic glycolysis as metabolic basis for trained immunity. Science 345, 1250684 (2014).
Larsen, S. B. et al. Establishment, maintenance, and recall of inflammatory memory. Cell Stem Cell 28, 1758–1774 (2021).
Klichinsky, M. et al. Human chimeric antigen receptor macrophages for cancer immunotherapy. Nat. Biotechnol. 38, 947–953 (2020).
Morrissey, M. A. et al. Chimeric antigen receptors that trigger phagocytosis. eLife https://doi.org/10.7554/eLife.36688 (2018).
Chan, M. W. Y. & Viswanathan, S. Recent progress on developing exogenous monocyte/macrophage-based therapies for inflammatory and degenerative diseases. Cytotherapy 21, 393–415 (2019).
Starkey Lewis, P. J., Moroni, F. & Forbes, S. J. Macrophages as a cell-based therapy for liver disease. Semin. Liver Dis. 39, 442–451 (2019).
Happle, C. et al. Pulmonary transplantation of human induced pluripotent stem cell-derived macrophages ameliorates pulmonary alveolar proteinosis. Am. J. respiratory Crit. Care Med. 198, 350–360 (2018).
Merad, M. & Martin, J. C. Pathological inflammation in patients with COVID-19: a key role for monocytes and macrophages. Nat. Rev. Immunol. 20, 355–362 (2020).
Gal-Oz, S. T. et al. ImmGen report: sexual dimorphism in the immune system transcriptome. Nat. Commun. 10, 4295 (2019).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Yoshida, H. et al. The cis-regulatory atlas of the mouse immune system. Cell 176, 897–912 (2019).
Robb, L. et al. Hematopoietic and lung abnormalities in mice with a null mutation of the common β subunit of the receptors for granulocyte-macrophage colony-stimulating factor and interleukins 3 and 5. Proc. Natl Acad. Sci. USA 92, 9565–9569 (1995).
Nassif, X. & Sansonetti, P. J. Correlation of the virulence of Klebsiella pneumoniae K1 and K2 with the presence of a plasmid encoding aerobactin. Infect. Immun. 54, 603–608 (1986).
Riottot, M. M., Fournier, J. M. & Jouin, H. Direct evidence for the involvement of capsular polysaccharide in the immunoprotective activity of Klebsiella pneumoniae ribosomal preparations. Infect. Immun. 31, 71–77 (1981).
Corces, M. R. et al. Lineage-specific and single-cell chromatin accessibility charts human hematopoiesis and leukemia evolution. Nat. Genet. 48, 1193–1203 (2016).
Acknowledgements
We thank L. Chasson for histology (CIML), M. Barad, S. Bigot, A. Zouine (CIML), A. Gompf and K. Bernhardt (CMCB Technology Platform, TU Dresden) for flow cytometry, M. Sohn for sequencing and the K. Rajewsky laboratory for hosting and general support (MDC). We also thank L. Razafindramana, CIML and CRTD Animal facilities for mouse husbandry and the light microscopy facility of the CMCB technology platform, TU Dresden. This study was supported by institutional grants from TU Dresden, the Institut National de la Santé et de la Recherche Médicale, Centre National de la Recherche Scientifique and Aix-Marseille University and grants to M.H.S. from the Agence Nationale pour la Recherche (ANR-17-CE15-0007-01 and ANR-18-CE12-0019-03), Fondation ARC pour la Recherche sur le Cancer (PGA1 RF20170205515), an INSERM-Helmholtz cooperation and the European Research Council under the European Union’s Horizon 2020 Research and Innovation program (grant agreement no. 695093 MacAge). S. Subramanian and S.V.A. received doctoral fellowship from the MDC and S.V.A and L.G received doctoral fellowship from the Fondation ARC pour la Recherche sur le Cancer, respectively. K.M. received support from a Human Frontier Science Program long-term fellowship and Stiftung Charité. M.H.S. is an Alexander von Humboldt Professor at TU Dresden (FRA/1188926).
Author information
Authors and Affiliations
Contributions
M.H.S., K.M., S. Subramanian, C.J.B., L.G. and S.V.A. designed experiments and analyzed data. S. Subramanian, C.J.B., K.M. and L.G. performed most experiments. S.V.A., J.F., R.Y.Y.A., M.B. and S.G. contributed to experiments and data analysis. J.M., H.B. and G.G. performed bioinformatic analysis. S. Sarrazin, L.A., B.dL. and P.K.K. supervised experiments and provided advice. S. Subramanian and C.J.B. contributed to writing the manuscript, M.B. and S. Subramanian generated figures. C.J.B. and M.H.S. supervised the project, M.H.S. conceived the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Andrew Macdonald and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Ioana Visan was the primary editor on this article and managed its editorial process and peer review in collaboration with the rest of the editorial team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Freezing exAM does not affect cell death rate or AM phenotype.
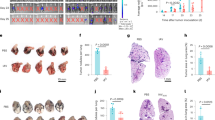
(a) Flow cytometric analysis of early (annexin V+/7-AAD–) and late (annexin V+/7-AAD+) apoptotic cell death in 4 months exAM with 1 hour staurosporine (STS) treated exAM as positive control. Data shown as mean ± s.d. (n = 4). (b) Flow cytometric comparison of AM cell-surface markers SiglecF and CD11c and viability dye (Zombie NIR) on freshly isolated AM and freeze-thawed exAM. Frozen exAM were 2 months expanded, 2 years 5 months frozen and expanded 1 month after thawing. Data shown are representative of 2 independent experiments.
Extended Data Fig. 2 exAM show lysosome activity and phagocytosis similar to AM.
Representative fluorescence images for the lysosomal marker acridine orange (a), lysosomal cathepsin B activity using Magic Red (b) and phagocytosis of green fluorescent latex beads (c) on AM and exAM. Scale bars 10 μm. Data are representative of 2 independent experiments with at least 3 biological replicates each.
Extended Data Fig. 3 Pearson correlation of RNA-seq samples.
Pearson correlation scatter-plots of replicates from pools of 3 mice each for Host AM, exAM, tAM and texAM RNA-seq samples.
Extended Data Fig. 4 Transcriptomic adaptation of exAM in response to regulatory cues in the changed microenvironment.
(a) Gene set enrichment analysis (GSEA) for Reactome: Surfactant metabolism (R-HAS-9006936) in AM vs exAM. Leading-edge surfactant related genes highlighted below. (b) Gene expression heat maps of key genes from GO terms, “carbon metabolism” and “inositol phosphate signaling” of analysis shown in Fig. 3f, showing rlog normalized values from minimum (blue) to maximum (red).
Extended Data Fig. 5 Similar number of differences detected in tAM vs Host AM compared to exAM vs Host AM.
(a) Experimental setup for intra-tracheal transplantation of BAL AM or 2 months expanded exAM (wild type, WT CD45.2) into lungs of WT CD45.1 host mice. Created with Biorender.com. (b) Overlaid volcano plot showing DEGs in exAM (red) or tAM (blue) compared to Host AM. Number of genes in blue and red with threshold FC > 2 and FDR < 0.05.
Extended Data Fig. 6 Pearson correlation of ATAC-seq samples.
Pearson correlation scatter-plots of replicates from pools of 3 mice for BAL AM, Host AM, exAM and texAM ATAC-seq samples.
Extended Data Fig. 7 Differential OCRs in BAL AM vs Host AM and in exAM vs texAM.
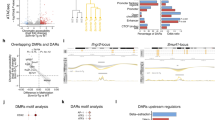
Scatter-plot analysis of ATAC-seq peaks differentially gained (red) or lost (blue) in BAL AM vs Host AM and exAM vs texAM.
Extended Data Fig. 8 Similar chromatin accessibility within in vivo AMs in contrast to exAM.
Heat map of unsupervised clustering of differential ATAC-seq peaks across all conditions, showing rlog normalized values from minimum (blue) to maximum (red).
Extended Data Fig. 9 Association of core myeloid transcription factors with gained and lost exAM OCRs.
Motif enrichment analysis (MEA) for transcription factor (TF) binding sites on open chromatin region (OCR) gained (top panel) or lost (bottom panel) in exAM compared to host AM.
Extended Data Fig. 10 exAM reconstitution of Csf2rb-/- lungs at 3 months post transplantation.
Flow cytometric analysis of exAM contribution to CD45.2 Csf2rb-/- lungs at 3 months post exAM transplantation. FACS plots show engraftment of CD45.1 texAM (top panel) and staining for surface markers SiglecF and CD11c (bottom panel). Quantifications shown in Fig. 6b.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1, ATAC-seq sort gating strategy for AMs.
Rights and permissions
About this article
Cite this article
Subramanian, S., Busch, C.JL., Molawi, K. et al. Long-term culture-expanded alveolar macrophages restore their full epigenetic identity after transfer in vivo. Nat Immunol 23, 458–468 (2022). https://doi.org/10.1038/s41590-022-01146-w
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-022-01146-w
This article is cited by
-
Sexual dimorphism of lung immune-regulatory units imprint biased pulmonary fibrosis
Cellular & Molecular Immunology (2025)
-
Inducible antibacterial responses in macrophages
Nature Reviews Immunology (2025)
-
Human liver immunology: from in vitro models to new insights
Cellular & Molecular Immunology (2025)
-
Clinical landscape of macrophage-reprogramming cancer immunotherapies
British Journal of Cancer (2024)
-
Camouflaging attenuated Salmonella by cryo-shocked macrophages for tumor-targeted therapy
Signal Transduction and Targeted Therapy (2024)