Abstract
Regulatory T (Treg) cells are vital for immune suppression. The role of the coreceptor programmed cell death 1 receptor (PD-1) in Treg cell function is controversial. Here, we demonstrate that PD-1 deficiency enhances the function of Treg cells through expression of a compensatory network of coinhibitory receptors. CD30 has a central role within this network, driving the Treg cell suppressive function within the tumor microenvironment. Mechanistically, PD-1 deficiency enhances STAT5 signaling in Treg cells, which induces CD30 expression. These data indicate a role for PD-1 as a checkpoint that negatively controls CD30 expression in Treg cells to limit their suppressive function. Understanding the functional changes that PD-1 has on Treg cells might enable combination therapies with better treatment outcomes in cancer.
Similar content being viewed by others
Main
Regulatory T (Treg) cells are a subset of CD4+ T helper cells that restrain the immune system. Treg cells express the forkhead transcription factor FoxP3 (ref. 1), along with several coinhibitory receptors, namely cytotoxic T lymphocyte antigen 4 (CTLA4) and programmed cell death 1 receptor (PD-1), to which specific functions have been attributed2,3,4. PD-1 binds to its cognate ligand programmed death ligand 1 (PD-L1) or PD-L2 (ref. 5). When bound to PD-L1, PD-1 enhances FoxP3 expression in stimulated murine CD4+ T helper cells3 and maintains FoxP3 expression in human and murine Tbet+ T helper 1 cells4,6.
PD-1 and PD-L1 antibodies have been used successfully in the treatment of several cancers, yet many cancer patients do not respond to this treatment, and in some cases PD-1 inhibition may enhance cancer growth. The underlying mechanisms that drive this resistance within the immune system are unclear. Correlative studies in human cancers have shown that immune resistance after PD-1 therapy can be attributed to Treg cells,7 and PD-1 deficiency drives Treg cell dysfunction8. Current observations9,10,11 are in line with findings by Chen et al.12 that PD-1 deficiency has an impact on peripheral Treg cell generation. Hence, PD-1 can negatively affect Treg cell function in autoimmunity and infections, potentially contributing to immunotherapy resistance in cancer and driving hyperprogressive disease7,10. Therefore, the current literature reports both negative and positive regulatory functions for PD-1 in Treg cells, with a lack of consensus on the role of PD-1 in driving Treg cell function.
Understanding of PD-1 signaling in Treg cells might provide insight into its function. The downstream signaling of PD-1 is dependent on its ITSM motif13, which recruits and activates Src homology phosphatase 1/2 (SHP1/2)14,15, leading to dephosphorylation and inactivation of downstream signaling pathways. PD-1 signaling in CD8+ T cells and (to some extent) in Treg cells has been shown to inhibit downstream TCR and CD28 signaling16,17. This includes inhibition of the PI3K–Akt signaling pathway in naive murine T cells, which drives functional plasticity of these cells and their differentiation into Treg cells3. However, the opposite has also been shown to enhance Treg cell function, with unchecked PI3K–Akt signaling enhancing the proliferation of Treg cells, leading to superior function in the absence of PD-1 (ref. 9).
We have previously found that PD-1 can dampen STAT1/4/5 phosphorylation4,18, and PD-1–STAT5 signaling in Treg cells has been reported to be important in human viral infections19,20. The outcome of enhanced STAT5 signaling in Treg cells is unclear; suggesting a crucial role for enhanced STAT5 in driving a regulatory phenotype in Treg cells, independent of FoxP3 expression.
In this study, we use BD Rhapsody mouse and human single-cell RNA sequencing (scRNA-seq) analysis and 1000-plex CosMx spatial analysis to identify an alternative regulatory pathway in PD-1-deficient Treg cells that can enhance their function. In the absence of PD-1, an increase in specific coinhibitory receptor expression is noted that enhances Treg cell function within the tumor microenvironment (TME). Mechanistically, PD-1 deficiency enhances STAT5 signaling, which in turn results in upregulation of CD30 receptor on Treg cells, driving increased Treg cell function. We confirm that CD30 expression in human Treg cells is driven by blockade of the PD-1–PD-L1 pathway. These findings highlight the importance of PD-1 in Treg cell function and suggest combination strategies that could benefit patients who are refractory to PD-1 therapy owing to increased Treg cell numbers.
Results
PD-1 intrinsically controls CD30 expression by Treg cells
A role of PD-1 in modifying Treg cell frequency in steady state has been reported9, but the expression and function of other coinhibitory receptors within these Treg cells has not been explored. We found a significant increase in FoxP3 frequency in splenocytes from Pd1−/−Foxp3RFP mice compared with wild-type (WT) animals (note that Pd1 has been used here in place of the official gene symbol Pdcd1) (Fig. 1a,b and Supplementary Fig. 1a). No difference was noted in the proportion of Treg cells that expressed Nrp1, Helios, CD25, Tbet, Gata3 or RORγt21,22,23 (Fig. 1c,d and Supplementary Fig. 1b–f). We investigated the expression of CD30, CTLA4, GARP, GITR and TIGIT coinhibitory receptors on WT and Pd1−/− Treg cell subsets and found significant increases in absolute numbers of all measured coreceptors except TIGIT in the cells from Pd1−/− mice compared with WT mice (Fig. 1e–j). Following Odorizzi et al.24, we performed Boolean analysis of coinhibitory receptor networks and found a significant increase in the coinhibitory receptor network CD30+GITR+CTLA4+TIGIT+ subset in Pd1−/− Treg cells (Fig. 1k,l and Supplementary Table 1). No change in frequency or coreceptor landscape was noted in CD4+FoxP3− cells, suggesting specific regulation driven by PD-1 in Treg cells (Supplementary Fig. 1g–i). Similar to Perry et al.11, we observed an increase in CD62L−CD44+ Treg cells and a reduction in CD62L+CD44− Treg cells; no difference was found in IL-10, IFNγ or IL-17 expression (Supplementary Fig. 1j–o). We next tested whether the coinhibitory receptors found in effector Treg cells could be attributed to an intrinsic or extrinsic requirement of PD-1. For this analysis, we generated Pd1fl/fl-DTR-Tdtomato (Pd1fl/fl) mice, in which loxP sites were inserted either side of exons 2 and 3 of the Pd1 gene25. In addition, an inserted IRES drove expression of the tdTomato reporter and the diphtheria toxin receptor (DTR). We crossed this strain to Foxp3ERT2-Cre-eGFP (ref. 26) to generate Pd1fl/fl-DTR-TdtomatoFoxp3ERT2-Cre-eGFP (Pd1fl/flFoxp3ERT2Cre) mice (Supplementary Fig. 2a–d). Using these animals, we repeated experiments after tamoxifen treatment. We observed an increase in FoxP3 expression (Fig. 1m,n) but no change in CD25, Helios, Tbet, Gata3 or RORγt expression or in the frequency of CD62L−CD44+ Treg cells or CD62L+CD44− Treg cells (Supplementary Fig. 2e–k). Consistent with our findings in Pd1−/−Foxp3RFP mice, only CD30 showed significantly increased expression in Pd1fl/flFoxp3ERT2Cre Treg cells; no changes were noted in CTLA4, GARP, GITR or TIGIT (Fig. 1o–p and Supplementary Fig. 2l–o). Again, a compensatory increase in the coinhibitory receptor network was noted (Fig. 1q,r and Supplementary Table 2). By contrast, no change in cytokine expression was observed (Supplementary Fig. 2p–r). We also found significant upregulation of Tnfrsf8 (CD30) mRNA (Fig. 1s,t). Finally, we tested whether PD-1 intrinsically controlled CD30. Using Pd1fl/fl-DTR-Tdtomato Foxp3ERT2-Crehet-eGFP (Pd1fl/flFoxp3ERT2Cre+/−) mice, we found that CD30 was intrinsically inhibited by PD-1 in Treg cells (Fig. 1u–w). No change in GARP or TIGIT expression was noted, but there was intrinsic upregulation of GITR and CTLA4 (Supplementary Fig. 2s–v, right panel).
a–j, Foxp3RFP (WT) and Pd1−/−Foxp3RFP (Pd1−/−) spleens were immunophenotyped using flow cytometry (n = 7 per group): flow cytometry profiles (a) and summaries of FoxP3 expression (b) in WT and Pd1−/− mice; frequencies of neuropilin 1 (Nrp1) (c) and Helios (d) expression in Treg cells; flow cytometry profiles of CD30, CTLA4, GARP, GITR and TIGIT expression in Treg cells (e); and absolute numbers of CD30 (f), CTLA4 (g), GARP (h), GITR (i) and TIGIT (j) in Treg cells are shown. k, Boolean analysis of expression of five different coinhibitory receptors in Treg cells. l, Linear analysis of CD30 network of coreceptors in Treg cells. m–r, Spleens of Pd1fl/flFoxp3ERT2Cre mice that had undergone preadministration of 1 mg tamoxifen or PBS for 5 consecutive days via i.p. injection (n = 7 per group) were analyzed: representative flow cytometry analysis of FoxP3 expression (m); summary data of FoxP3 expression (n); a representative flow plot of CD30, CTLA4, GARP, GITR and TIGIT expression in Treg cells (o); absolute numbers of CD30 in Treg cells (p); Boolean analysis of expression of five different coinhibitory receptors in Treg cells (q); and absolute counts of the CD30 network of coreceptors (r) are shown. s,t, RT–qPCR analysis of Tnfrsf8 mRNA expression in Treg cells from Pd1−/−Foxp3RFP (n = 7) (s) and Pd1fl/flFoxp3ERT2Cre+/− (n = 6) (t) female mice treated with tamoxifen. u, Representative flow cytometry analysis of CD30 expression in CD4+FoxP3+CD44+PD-1+ and CD4+FoxP3+CD44+PD-1− T cells. v,w, Absolute numbers of CD30 in PD-1+ and PD-1− Treg cells from Foxp3ERT2Cre (v) and Pd1fl/flFoxp3ERT2Cre+/− (w) mice. Data are shown as mean ± s.e.m., with each data point representing an animal from an independent experiment, except in v and w, in which each pair of data points are from the same animal’s Treg cells with or without PD-1 expression and from independent experiments. Two-tailed unpaired Student’s t-tests were performed for the results shown in b–d, f–j, l, n, p and r–t and two-tailed paired Student’s t-tests for those shown in v and w. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001. FMO, fluorescence minus one; NS, not significant.
PD-1 deficiency does not alter proliferation or apoptosis of Treg cells
In ex vivo splenocyte experiments, we found higher proportions of Ki67+ expression in both Pd1−/− Treg cells and CD4+FoxP3− T cells compared with WT cells; however, contrary to previous reports9,11, these differences were not significant (Supplementary Fig. 3a–c). WT and Pd1−/− Treg cells stimulated with αCD3, αCD28 and IL-2 also showed no difference in proliferation (Supplementary Fig. 3d,e). Treg cells expressing CD30, CTLA4, GITR and TIGIT were highly proliferative in both WT and Pd1−/− animals (Supplementary Fig. 3f,g). No change in Annexin V staining or total live cells was observed between WT and Pd1−/− Treg cell populations (Supplementary Fig. 3h,i). A similar profile was observed for CD4+FoxP3− T cells (Supplementary Fig. 3j,k). No change in Bcl2 expression was noted in either Treg cells or CD4+FoxP3− T cells (Supplementary Fig. 3l,m) between the WT and Pd1−/− cohort. On further analysis of the FoxP3 compartment, WT Treg cells expressing the CD30 network of coinhibitory receptors showed a significantly higher apoptotic index, whereas no such change was observed in Pd1−/−CD30+ Treg cells (Supplementary Fig. 3n,o).
CD30 can enhance Pd1 −/− Treg function within the TME
We established B16 tumors in WT and Pd1−/− mice. Tumor growth was monitored (Supplementary Fig. 4a), and tumor-infiltrating lymphocytes (TILs) were harvested and subjected to scRNA-seq analysis. We used germline Pd1−/− mice to generate a resource that would be relevant to the clinical setting. Immune cell clusters in WT and Pd1−/− TILs were defined (Fig. 2a–d); then, Treg cell clusters were determined based on Cd4 and Foxp3 (Fig. 2c and Supplementary Table 3a). Clusters were visualized with uniform manifold approximation and projection (UMAP) following Louvain clustering. Differential gene expression analysis, which was illustrated using a volcano plot, showed that Tnfrsf8 was significantly upregulated in Pd1−/− Treg cell TILs compared with other canonical coinhibitory receptors (Fig. 2e and Supplementary Table 3b). Among effector T (Teff) cells, we found an increase in Gzm transcripts in Pd1−/− TILs compared with WT TILs (Supplementary Fig. 4c,d and Supplementary Table 3c,d). Gene ontology (GO) enrichment analysis and gene set enrichment analysis (GSEA) demonstrated that the Pd1−/− Treg cell cohorts had enhanced cytokine signaling (Fig. 2f,g and Supplementary Table 3e,f). We interrogated whether the changes in the coinhibitory network in steady state in Pd1−/− Treg cells compared with WT Treg cells was preserved in the TME. There was a significant increase in Tnfrsf8 expression in Pd1−/− Treg cells compared with WT Treg cells (Fig. 2h and Supplementary Fig. 4b). Using Boolean analysis, we found that the altered coinhibitory receptor network observed in steady-state Pd1−/− Treg cells was preserved at the transcript level in Pd1−/− Treg cell TILs (Fig. 2i).
WT and Pd1−/− mice TILs were harvested on day 14 and subjected to scRNA-seq. a, UMAP view of 1,810 enriched TILs color-coded by assigned cell type. b, WT and Pd1−/− TILs, decoupled to show similar clustering between the samples. c, UMAP view of CD4 and Foxp3 expression. d, Dot plot of candidate gene expression in immune clusters. e, Differential gene expression analysis of Pd1−/− Treg cells versus WT Treg cells. f, GO enrichment analysis identifying the functions of upregulated genes in Pd1−/− Treg cells. g, GSEA showing enhanced regulation of cytokine production of Pd1−/− Treg cells. NES, normalized enrichment score. h, mRNA expression of Tnfrsf8, Ctla4, Tnfrsf18 (GITR), Lrrc32 (GARP) and Tigit transcripts in WT and Pd1−/− Treg cells in individual animals. i, Average percentages of Treg cells expressing 0–5 Tnfrsf8 (CD30), Ctla4, Tnfrsf18 (GITR), Lrrc32 (GARP) and Tigit transcripts in WT and Pd1−/− Treg cells in the TME. j, Pseudotime trajectory analysis of WT (left) and Pd1−/− Treg (right) cells. k,l, Heatmap showing cell patterns and received signaling patterns of WT (k) and Pd1−/− (l) TILs. m, Chord diagram visualizing numbers of interactions between Treg cells and TILs in WT (top) and Pd1−/− (bottom) cohorts. The inner bar size is proportional to the signal strength received by the TILs from Treg cells. Statistical analyses were performed using two-tailed Wilcoxon rank-sum test with Bonferroni correction in e, Fisher’s exact test with FDR correction in f, two-tailed Kolmogorov–Smirnov test with FDR correction in g, and two-tailed unpaired Student’s t-test in h. Data are mean ± s.e.m. from n = 5 mice per cohort in h. ***P < 0.001. For the results shown in k–m, CellChat was used to model communication probabilities based on the law of mass action and identify significant communications using permutation tests. Padj., adjusted P value.
Pseudotime and trajectory analysis using the Monocle plugin in SeqGeq demonstrated that WT and Pd1−/− Treg cells combined had seven states of differentiation (Supplementary Fig. 4e,g). When WT and Pd1−/− Treg cells were separated, pseudotime trajectory analysis revealed five differentiation states in Pd1−/− Treg cells and three in WT Treg cell TILs (Fig. 2j and Supplementary Table 3g,h). WT Treg cells showed a tissue-resident state marked by St2 (state 1), an immunosuppressive phenotype marked by Tigit (state 2) and a cytotoxic state (state 3). By contrast, Pd1−/− Treg cells comprised a type 1 phenotype characterized by either IFNγR (state 1) or Stat1 expression (state 3), a cytotoxic phenotype (state 2), an immune-regulatory phenotype (state 4) and a TNFR-enriched differentiation program (state 5).
Using CellChat analysis, we determined the communication program of Treg cells within the TME. Whereas WT Treg cells showed a single incoming pattern dominated by CD86 (activating CTLA4) and PD-L1 (through PD-1), Pd1−/− Treg cells showed dual incoming signaling patterns dominated by CD86 and TNF (activating TNF receptors through CD30; red boxes in Fig. 2k,l). We next investigated the signals contributed by all immune cells among the TILs; we found no significant difference in incoming signaling patterns between the WT and Pd1−/− cohorts, but Pd1−/− natural killer (NK) cells contributed less to outgoing signaling patterns (Supplementary Fig. 5a,b). We next determined the crosstalk of Treg cells with other immune cells within the TME. Analysis of receptor–ligand interactions, the results of which were visualized using a chord diagram, showed increases in expression of several binding partners of Pd1−/− Treg cells with NK cells and myeloid dendritic cells (DCs) (Fig. 2m) compared with WT Treg cells. In turn, communication of NK cells with Teff cells was predicted to be diminished, whereas no changes in myeloid DC communication were noted (Supplementary Fig. 5c,d). This predictive analysis suggests that the interaction between Pd1−/− Treg cells and NK cells may influence the interactions of NK cells with Teff cells within the TME; however this would need to be validated in future studies and is beyond the scope of the current work.
Functional validation of scRNA-seq and CellChat analysis of the TME
Similar to previous work, we found that Pd1−/− Treg cells were significantly more suppressive in vitro27 (Fig. 3a,b). For analysis of in vivo suppressive function, Rag1−/− mice bearing tumors were reconstituted with either WT or Pd1−/− CD45.2+ Treg cells along with WT CD45.1+ naive CD4+ T cells. The Rag1−/− cohort reconstituted with Pd1−/− Treg cells showed significantly enhanced tumor growth compared with those reconstituted with WT Treg cells (Fig. 3c). No significant differences in Treg cell frequency, cytokine production or cytokine production in CD4+ Teff cells were observed between the two groups (Supplementary Fig. 6a–d). We next sought to confirm the results of our scRNA-seq and CellChat analysis of CD30 expression in Pd1−/− Treg cells. Using the same Rag1−/− tumor model, we found that CD30 frequency was significantly increased in Treg cells but not in CD45.1+ Teff cells among TILs (Fig. 3d). We next tested whether Treg cells in Pd1fl/flFoxp3ERT2Cre mice had a tumor-protective role similar to that of global Pd1−/− Treg cells. For this purpose, we established tumors in control mice treated with phosphate-buffered saline (PBS) and Pd1fl/flFoxp3ERT2Cre mice after tamoxifen treatment. Significantly increased tumor growth was noted in the cohort with PD-1-deficient Treg cells (Fig. 3e), along with significant increases in absolute numbers of CD30+ TIL Treg cells in Pd1fl/flFoxp3ERT2Cre mice compared with Pd1fl/fl control mice (Fig. 3f). By contrast, we found no change in absolute numbers of FoxP3+, IFNγ+, IL-10+, GITR+ or CTLA4+ TIL Treg cells (Supplementary Fig. 6e–i).
a,b, Suppressive function of Treg cells from Foxp3RFP and Pd1−/−Foxp3RFP mice (n = 8 per group) as demonstrated by a representative CellTrace Violet flow plot (a) and percentages of Teff cell proliferation (b). c,d, For Rag1−/− mice with B16F10 melanoma cells and reconstituted with CD45.1+ T cells and CD45.2+ Treg cells from either WT or Pd1−/− mice, tumor volumes (n = 8 for WT and n = 6 for Pd1−/− groups) (c) and frequencies of CD30 expression on CD45.1+ Teff and CD45.2+ Pd1−/− Treg cells in TILs (n = 5) (d) are shown. e,f, For tamoxifen-treated Pd1fl/fl and Pd1fl/flFoxp3ERT2Cre mice with tumors, tumor volumes (n = 10 for Pd1fl/fl and n = 9 for Pd1fl/flFoxp3ERT2Cre) (e) and absolute counts of CD30+ Treg cells (n = 12 for Pd1fl/fl and n = 11 for Pd1fl/flFoxp3ERT2Cre) (f) are shown. g, Tumor-bearing Rag1−/− mice were reconstituted with different T cell populations (n = 3 for Teff plus WT Treg cell group and n = 7 for Teff plus Pd1−/− Treg cell groups); in the Teff plus Pd1−/− Treg cohorts, mice were treated with either 0.1 mg anti-IgG2a (n = 5) or anti-CD153 (CD30L, n = 4), and tumors were measured. h, Tumor-bearing Rag1−/− mice were reconstituted with Teff plus WT Treg (treated with either 0.1 mg anti-IgG2a (n = 5) or anti-CD153 (n = 5)), and tumors were measured. Ab, antibody. i–m, Tamoxifen-treated and tumor-bearing Pd1fl/fl (n = 10) and Pd1fl/flFoxp3ERT2Cre (n = 8 or 9) mice were immunophenotyped on day 19: frequencies of FoxP3 (i); ST2, TIGIT and GzmB in Pd1fl/fl mice (j); IFNγR (CD119), GzmB, Tbet, TIGIT and IL-10 in Pd1fl/flFoxp3ERT2Cre mice (k); NK cells (l); and DCs (m) are shown. Data are presented as the mean ± s.e.m. Each data point represents an in vitro biological replicate in b or an individual animal in f, i, l and m. Data points in each group were matched to the same individual mouse in d, j and k. One-way analysis of variance (ANOVA) with Sidak’s multiple comparison was used in b; two-way ANOVA with Sidak’s multiple comparison in c, e, g and h, two-tailed paired Student’s t-test in d; two-tailed unpaired Student’s t-test in f, i, l and m; and matched one-way ANOVA with Sidak’s multiple comparison in j and k. Cumulative data from n = 3 independent experiments are shown. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Next, Rag1−/− mice were reconstituted with either WT or Pd1−/− CD45.2+ Treg cells along with WT CD45.1+ naive CD4+ T cells, but animals receiving Pd1−/− CD45.2+ Treg cells were injected with either an isotype control or anti-CD30L (also known as anti-CD153). We found that blocking the CD30 pathway with anti-CD30L significantly inhibited tumor growth after 14 days in the Pd1−/− CD45.2+ Treg cell cohort compared with animals that received the isotype control. This decrease in tumor growth caused by anti-CD153 treatment reversed most of the increase observed in animals reconstituted with Pd1−/− CD45.2+ Treg cells compared with those reconstituted with WT Treg cells (Fig. 3g). Next, we tested whether anti-CD153 treatment also had an impact on the protumor function of WT Treg cells. In Rag1−/− animals reconstituted with B16 melanoma, WT CD45.2+ Treg cells and CD45.1+ naive CD4+ T cells, we found no difference in tumor growth between those in which reconstitution was followed by anti-CD153 treatment and those who received the isotype control (Fig. 3h). These data indicate that the CD30/CD30L pathway is a specific target for PD-1-deficient Treg cell function and plays a minimal part in WT Treg cell function. Immunophenotyping data showed that anti-CD153 treatment did not deplete the frequency of either Treg cells or Teff cells in any condition, in contrast to previous reports28 (Supplementary Fig. 6j–m). We next sought to confirm our Monocle-predicted differentiation states and the results of our CellChat analysis. Pd1fl/flFoxp3ERT2Cre mice were reconstituted with tumors after tamoxifen treatment, and Treg cells were characterized. We found a significant increase in FoxP3 expression among the TILs (Fig. 3i). We then subcharacterized Treg cells based on our pseudotime trajectory analysis and detected the presence of all three substates in the WT and five different substates in the PD-1-deficient Treg cell TILs (Fig. 3j,k). Next, we tested whether PD-1 deficiency in Treg cells affected NK and DCs among TILs per the predictive CellChat analysis. We observed significant decreases in NK and DC frequencies among the TILs (Fig. 3l,m) but no changes for other functional markers (Supplementary Fig. 6n–t).
CD30 expression is driven by STAT5 signaling in Pd1 −/− Treg cells
We measured the ability of PD-1 to inhibit anti-CD3, anti-CD28 stimulation and IL-2-induced expression of CD30 in WT FoxP3+ T cells. In the absence of IL-2, neither PD-1 inhibition nor activation using a PD-L1 Fc chimera altered CD30 expression. In the presence of IL-2, PD-1 inhibition led to significant increases in both protein and mRNA CD30 expression compared with the PD-L1 Fc condition (Fig. 4a–c). Similarly, we observed a significant increase in CD30 expression in both WT and Pd1−/− Treg cells stimulated with anti-CD3/28 in the presence of IL-2 (Fig. 4d).
a–c, WT Treg cells from C57BL/6 mice were stimulated with αCD3 and αCD28 and cultured with either anti-mouse IL-2 (αmIL-2) or αmIL-2 with recombinant human IL-2 (rhIL-2). Cells were either cultured in recombinant mouse PD-L1 Fc (rmPD-L1-Fc)-coated well plates or with αPD-1 to block the PD-1–PD-L1 interaction: a representative plot of CD30 expression (a), a summary of CD30 expression in five independent experiments (b) and CD30 mRNA expression in three independent experiments (c) are shown. d, Treg cells from C57BL/6 (WT) and Pd1−/− mice were stimulated with αCD3 and αCD28 and cultured with either αmIL-2 or rhIL-2. At the indicated time points, CD30 expression on Treg cells was plotted as the fold change in expression compared with the unstimulated condition (0 h). Data from seven independent experiments are shown. e, WT Treg cells were stimulated with αCD3, αCD28 and αIL-2 in a PD-L1 Fc-coated plate. αPD-1 was added to block the PD-1–PD-L1 interaction. After 48 h, cells were stimulated with IL-2 for 15 min, and phospho-STAT5 (pSTAT5) was measured; the results are shown on the left. A summary of pSTAT5 in CD30+ Treg cells is shown on the right. Uns., unstimulated. f, Representative ChIP–seq tracks and peaks detected by STAT5 (upper) and p300 (lower) at the Tnfrsf8 gene locus of naive WT CD4+ cells stimulated with αCD3, αCD28 and rhIL-2 for 3 days. g,h, Naive WT and Stat5−/− CD4+ cells were stimulated with αCD3, αCD28 in the presence of αIL-2 for 72 h, washed and restimulated with αCD3 along with either αIL-2 or rhIL-2 for 4 days: GSEA plots showing Tnfrsf family gene rankings in IL-2-treated WT CD4+ cells compared with αIL-2-treated WT CD4+ cells (left) and Stat5−/− CD4+ cells (right) (g); and heatmaps showing expression of coreceptors and ligands in WT and Stat5−/− CD4+ cells (h) are shown. Data are presented as the mean ± s.e.m.; each data point represents an independent experiment. Statistical analyses were performed using two-tailed paired Student’s t-tests (b, c and e) or two-way ANOVA with Sidak’s multiple comparison (d). *P < 0.05, **P < 0.01, ****P < 0.0001.
We next found that PD-1 activation through PD-L1 downregulated STAT5, whereas anti-PD-1 enhanced STAT5 phosphorylation in Treg cells (Fig. 4e). Using previously published RNA-seq and CHIP–seq (chromatin immunoprecipitation followed by sequencing) datasets, we explored the effects of STAT5 on CD30 expression29. We found that STAT5 bound to the Tnfrsf8 gene locus in WT CD4+ T lymphoblasts stimulated with IL-2 for 48 h and shared two sites with p300 on either side of the gene (Fig. 4f). To confirm that STAT5 positively regulated CD30 expression, we performed GSEA, comparing WT CD4+ T cells subjected to IL-2 stimulation with the anti-IL-2 condition, and IL-2-stimulated WT CD4+ T cells with Stat5−/− CD4+ T cells. Genes were then ranked with respect to their differential expression between the comparison groups; we found that the gene encoding CD30, a member of the TNFRSF family, was highly ranked among the genes induced by IL-2 in the presence of STAT5 (Fig. 4g). By contrast, the TNFRSF family as a group was not significantly enriched in either condition. Next, we evaluated whether a similar pattern in fold change could be observed for other common coinhibitory receptors and found that STAT5 did not significantly regulate clinically relevant coinhibitory receptors (Fig. 4h).
Spatial transcriptomic analysis of Pd1 −/− Treg cell interactions in the TME
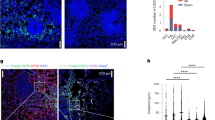
To visualize the predicted CellChat interactions and pseudotime trajectory analysis, we performed CosMx spatial transcriptomics. Rag1−/− mice engrafted with tumors were reconstituted with either WT or Pd1−/−CD45.2+ Treg cells, along with WT CD45.1+ naive CD4+ T cells. Tumor sections stained with hematoxylin and eosin (H&E) from both groups of animals were evaluated, and identical tissues were used for the CosMx analysis. Fields of view (FOVs) that were enriched for both immune infiltration and surrounding tumor tissue were chosen for analysis. A combination of cell segmentation analysis and immune cell profiling enabled us to identify single cells and their corresponding cell types within the tissue (Supplementary Fig. 7a,b). Immune cell profiling was performed using Louvain clustering analysis across all FOVs from the tested samples, integrating RNA transcripts for enhanced resolution (Fig. 5a, Supplementary Fig. 7c and Supplementary Table 4a). This approach allowed subdivision and classification of the ‘immune cells’ cluster based on specific immune cell markers, which were then mapped to single cells within the tissue (Supplementary Fig. 7b, right). Frequencies of the various cell populations in the two cohorts were analyzed (Fig. 5b), with the ‘immune cells’ cluster further subdivided by specific markers (Fig. 5c). Pathway analysis indicated upregulation of leukocyte migration pathways in Pd1−/− Treg cells compared with WT Treg cells (Supplementary Fig. 7d). The CosMx dataset was then used for several descriptive visual analyses of Treg cells in the TME. First, all Treg cell IDs and cell coordinates were obtained from the FOV object (Supplementary Table 4b). Each Treg cell interaction with neighboring cells was assigned as an event, and the number of events was determined through manual annotation of the FOV images. These interaction events were classified as single, dual or multiple (Fig. 5d). Treg cells were grouped based on their interactions with respective tumor or immune cells as follows. All the Treg cells interacting with a specific cell type were grouped using their unique cell ID (Supplementary Table 4c). Enriched transcripts in each group were determined using differential gene expression analysis to identify the short-range (SR) and long-range (LR) communication signals expressed by Treg cells (Supplementary Table 4d,e). We found that Treg cells used combinations of SR signals (ligand or receptor expression) to interact with CD4+ Teff cells, NK cells, DCs and macrophages (Fig. 5e), together with tissue-resident cells within the TME. However, it should be noted these were descriptive analyses using manual annotation that suggested possible interactions with other immune cells. We found potential interactions of Treg cells with NK and DCs using either SR or LR communication signals. Hence, we analyzed the mRNA of DCs and NK cells for functional changes. Although we observed slight increases in Il10 and Gzmb expression in DCs and NK cells (Fig. 5f,g), when we attempted to validate these results in vivo using tumor-bearing Pd1fl/fl and Pd1fl/flFoxp3ERT2Cre mice, no difference in IL-10 or GzmB protein expression in DCs or NK cells was found (Fig. 5h,i).
a–g, Tumor-bearing Rag1−/− mice were reconstituted with different T cell subsets and then subjected to CosMx spatial transcriptomics: UMAPs of 9,231 cells derived from WT Teff/WT Treg cell and WT Teff/Pd1−/− Treg cell cohorts, color-coded by assigned cell type (left), and WT and Pd1−/− Treg cell cohorts decoupled (middle and right) (a); frequencies of all cells in the TME (b); immune cells subclusters and frequencies of immune cell subsets among WT and Pd1−/− Treg cells (c); a graphical representation of cell–cell interaction analysis (left) and numbers of cell–cell interactions of WT Treg cells (middle) and Pd1−/− Treg cells (right) (d); a graphical representation of SR and LR gene definition by manual annotation (left), and frequencies of LR and SR interactions of WT (middle) and Pd1−/− Treg cells (right) (e); and DC Il10 expression (f) and Gzmb expression in NK cells (g) according to clustered Wilcoxon rank-sum test are shown. h,i, Tamoxifen-treated, tumor-bearing Pd1fl/fl and Pd1fl/flFoxp3ERT2Cre mice were immunophenotyped on day 19: the frequencies of IL-10 expression in TIL DCs (n = 5 per group) (h) and GzmB expression in TIL NK cells (n = 10 for Pd1fl/fl and n = 9 for Pd1fl/flFoxp3ERT2Cre) (i) are shown. j, For mice as in a–g, FOVs showing colocalization of different WT and Pd1−/− Treg cell states and their interactions within the TME. k,l, Tamoxifen-treated, tumor-bearing Pd1fl/flFoxp3ERT2Cre mice were immunophenotyped (n = 8): representative flow cytometry results (k) and a summary of CD30 expression in Treg cell subsets (l) are shown. Data represent individual cells in f and g. In h, i and l, data are from three independent experiments, and the mean ± s.e.m. is shown. In h and i, each data point represents an individual mouse. In l, data points are matched within animals. Two-tailed clustered Wilcoxon rank-sum test was used for f and g, two-tailed unpaired Student’s t-test for h and i, and one-way ANOVA with Sidak’s multiple comparison test for l. *P < 0.05, **P < 0.01. Illustrations in d and e created using BioRender.com. ILC, innate lymphoid cell.
As our CosMx panel did not include Tnfrsf8, we investigated whether we could identify which cell type within the TME predominantly communicated through Tnfsf8 (CD30L) with Treg cells, as CD30 is the only binding partner for CD30L. We found CD30L expression in tissue-resident cells, tumor cells and immune cells in the cohorts treated with WT and Pd1−/− Treg cells (Supplementary Fig. 7e and Supplementary Table 4f). We next visualized cells expressing CD30L that were in contact with Treg cells across all the FOVs. The frequency of each cell type was visualized, and the possible interactions of all cells with Treg cells via CD30L within the TME were identified (Supplementary Fig. 7f). The mRNA data suggested that CD30L was ubiquitously expressed in all cell types in the TME.
We next sought to visually validate our pseudotime trajectory analysis in the TME and found three different subsets of Treg cells in the WT Treg cell-treated cohorts and five different subsets in the Pd1−/− Treg cell-treated cohort (Fig.5j). Taken together, the results of our descriptive analysis of the CosMx data indicated the presence of CD30L in all cells in the TME and identified different Treg cell subsets in the TME. Finally, we validated the expression of CD30 in these Treg cell subsets in the TME in Pd1fl/flFoxp3ERT2Cre mice. We observed CD30 expression in the regulatory and cytotoxic Treg cell subsets (Fig. 5k,l). Hence, our validation studies suggest that it is the regulatory subsets of the Pd1−/− Treg cell population in the TME that express CD30, and that blocking CD30/CD30L inhibits this regulatory axis.
PD-1 disruption not the TME enhances CD30 expression in human Treg cells
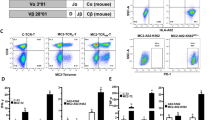
We tested whether CD30 expression in human Treg cells was driven by the TME or by anti-PD-1 effects by means of in-house BD Rhapsody scRNA-seq analysis of peripheral blood mononuclear cells (PBMCs) from stage IV human melanoma patients and healthy controls (HC). We also used a publicly available scRNA-seq dataset from melanoma TILs to validate our findings in naive melanoma TILs (Supplementary Table 5). Immune cell clusters in HC and melanoma samples (Fig. 6a–c) were defined using AbSeq protein expression and the top five gene transcripts expressed in each cluster (Supplementary Fig. 8a–c and Supplementary Table 6a,b), whereas the public dataset was analyzed using gene transcripts (Fig. 6d,e, Supplementary Fig. 9a and Supplementary Table 6c). We found no alteration of mRNA expression of CD30 or any other coreceptor in tumor-derived Treg cells among either TILs or melanoma PBMCs compared with HC PBMCs (Fig. 6f–i and Supplementary Fig. 8d–g). Notably, expression of TIGIT was significantly enhanced when measured in individual patients, suggesting regulation of TIGIT by the TME. The results of this analysis suggested that the TME alone does not drive enhanced CD30 expression in Treg cells among PBMCs or TILs in melanoma.
a–c, PBMCs from HC and stage IV melanoma patients (Mel) were subjected to scRNA-seq: a UMAP view of 11,634 cells, with all cells colored by cell type (a); HC and Mel PBMC UMAPs showing homogeneity of clusters between the two cohorts (b); and frequencies of each cluster within individual samples (c) are shown. H, healthy; M, melanoma. d,e, Publicly available TIL and tumor cell data were mined and analyzed: a UMAP view of 1603 cells, with all cells colored by cell type (d), and frequencies of each cluster in individual patients (e) are shown. Endo, endothelial. f–i, mRNA expression of TNFRSF8 (f), CTLA4 (g), TIGIT (h) and LRRC32 (GARP) (i) in Treg cells from PBMCs of HC individuals (n = 3) and Mel patients (n = 3), and Treg cells from TILs (n = 4), where each data point represents an individual. j, HC and Mel PBMCs were cultured with isotype control (IC) or anti-PD-L1 antibody (αPD-L1): a representative flow plot of CD30 expression in Treg cells (left) and a summary of CD30 expression in Treg cells (right) are shown. Pt, patient. k–o, mRNA expression of TNFRSF8 (k), CTLA4 (l), GARP (m), TNFRSF18 (n) and TIGIT (o) in Treg cells from individual patients at baseline and after anti-PD-1 treatment. p, TNFRSF8 mRNA expression in Treg cells in each patient identified as a responder (Res) or nonresponder (Nonres) to anti-PD-1 immunotherapy. q, Kaplan–Meier analysis of overall survival of patients diagnosed with different cancers corresponding to CD30 expression, with P values indicated. Data are presented as the mean ± s.e.m., with each data point representing an individual from at least three independent experiments in f–i and k–p. Statistical analysis was performed using one-way ANOVA with multiple Sidak’s comparison (f–i) or two-tailed unpaired Student’s t-test (k–p). *P < 0.05. The survival curve statistical analysis for q was performed using a two-tailed log-rank test with the R2 Genomics Analysis and Visualization Platform.
We next tested the effect of blocking the PD-1 pathway on CD30 expression in Treg cells using in vitro cultures. Treg cells cultured in the presence of an anti-PD-L1 antibody showed upregulation of CD30 for all three human melanoma patients (Fig. 6j), suggesting that it is not the TME but anti-PD-1 activity that drives CD30 expression in tumor Treg cells.
We next analyzed the expression of CD30 in Treg cells from a cohort of anti-PD-1-treated melanoma patients using a publicly available dataset. We first performed cluster analysis to classify immune cells (Supplementary Fig. 9b–e, and Supplementary Table 6d), followed by two types of analysis. In the first analysis, we tested CD30 expression and that of other coreceptors in patient samples both at baseline and after anti-PD-1 therapy. We found that when individual patient data were analyzed separately, TNFRSF8 expression was significantly enhanced after anti-PD-1 therapy, whereas no changes were found for CTLA4, LRRC32 (GARP), TNFRSF18 (GITR) or TIGIT (Fig. 6k–o). The results of the single-cell analysis were consistent with these findings, with a significant difference in TNFRSF8 expression and no differences for other coreceptors (Supplementary Fig. 9f–j). In the second analysis, we compared the expression of TNFRSF8 in Treg cells from responders and nonresponders to anti-PD-1 therapy. Again, we found that TNFRSF8 was predominantly expressed within the nonresponder cohort, but no significant difference was noted for other coinhibitory receptors (Fig. 6p and Supplementary Fig. 9k–n). Finally, we tested whether CD30 could be used as a biomarker for prediction of cancer-free survival. We found that elevated CD30 expression in PBMCs was associated with poor survival in patients with uveal melanoma and those with breast, lung or colon cancer, indicating that CD30 could be used as a biomarker and therapeutic target that may determine disease outcomes after checkpoint therapy (Fig. 6q).
Discussion
The use of antibodies against PD-1 and CTLA4 has been transformative in the management of many cancers, particularly melanoma. Despite the success of these treatments in certain cases, most cancer patients either fail to respond, relapse after an initial response or develop hyperprogressive disease in which PD-1 inhibition paradoxically stimulates tumor growth. The emerging literature suggests that anti-PD-1 immunotherapy can counterintuitively enhance Treg cell numbers in a proportion of patients, driving resistance to immunotherapy. In this study, using clinically relevant global Pd1−/− and biologically specific Pd1fl/fl animals, we elucidated three key aspects of PD-1 biology in Treg cells in steady state and in the TME. Specifically, we showed that PD-1 could inhibit CD30 coinhibitory receptor expression in (1) Treg cells in steady state, (2) Treg cells in the TME and (3) human Treg cells in melanoma. These data provide insight into the complex relationship between PD-1 and Treg cell function.
Our finding that Pd1−/− Treg cells were superior in function to WT Treg cells was in line with recent reports on autoimmunity and infectious diseases9,11. This increase in function was not due to increased proliferation as measured in vitro or increased accumulation as measured in vivo. Hence, we explored an alternative hypothesis that an increase in Treg cell function could result from compensatory upregulation of other well-known coinhibitory receptors such as GITR, CTLA4, TIGIT and GARP. However, the dominant coinhibitory receptor that was intrinsically regulated by PD-1 deficiency was CD30. This finding was unexpected but enabled us to identify a unique regulatory pathway in Treg cells that could be manipulated to enhance checkpoint therapies. Indeed, the roles of CD30 in immunotherapy and Treg cell biology have shown that CD30 plays a key part in Treg cell function in graft versus host disease30. Hence, our data are complimentary to previous observations and are relevant to anti-PD-1 resistance, as patients who have enhanced Treg cell numbers after PD-1 therapy could be treated with anti-CD30 antibodies. Murine studies have found that combining anti-CD30 therapy with other coreceptors can delete Treg cells28; this finding was not corroborated in our system, in which we found that blocking CD30/CD30L preserved Treg cell numbers within the TME.
The functional plasticity of Treg cells has been well established, with important roles for cytokines31 and coreceptors32. PD-1 is vital for maintaining FoxP3 expression in both human4 and murine Tbet+ Treg cells6. A recent study showed that Pd1−/− Treg cells were functionally plastic in the TME8 by acquiring an altered metabolic phenotype. This observation confirmed previous work by us and others, in which PD-1 deficiency was found to induce a glycolytic phenotype in T cells and innate immune cells (ILC2/ILC1)33,34. We found a similar phenotype in Pd1−/− Treg cells, with pseudotime analysis identifying type 1 differentiation states in Treg cells. Therefore, we concluded that PD-1 does control the emergence of Treg cells with type 1 phenotype; however, within the TME, it also controls Treg cell subsets that can drive immune regulation through TNFRs, TIGIT and IL-10. We proposed that within the TME, several Treg cell states would coexist, and their function would be determined by the predominant state during a particular space and time. We sought to confirm this using spatial transcriptomics to identify which Treg cell states were associated with either immune cells or tumor cells. We found that Treg cells expressing Il10, Tigit, and Tnfrsf were in proximity to tumor cells. Hence, Pd1−/− Treg cells could exhibit both functional plasticity and exert an immune suppressive function within the TME. Validation in mouse models further confirmed an increase in CD30 expression in the regulatory Treg cell subsets within the TME.
The communication patterns of WT and Pd1−/− Treg cells in TME were of particular note. We showed that deleting PD-1 enhanced a second pattern of communication among Treg cells that was primarily driven by TNF signaling, which may occur through TNFRs. Although GITR seemed to play a part in this communication pattern, it was CD30 that was predominantly activated. These data could potentially explain the lack of success of combination trials in which anti-PD-1 and GITR agonists have been used35. Similarly, anti-LAG3 combination trials have been proposed to overcome resistance; however, in our single-cell analysis, LAG3 was not altered in all T cell subsets. These data highlight the need for unsupervised analysis of T cells in clinical models of established large tumors as a proof of concept before building combination therapy trials using checkpoint antibodies.
Our data suggests that of all Treg differentiation states, the predominant phenotype should be targeted to overcome tumor resistance. In the TME, the predominant Treg cell phenotypes associated with tumor cells were immunosuppressive. These included TNFR and IL-10 phenotypes, which also expressed CD30; hence, targeting CD30+ Treg cells could substantially overcome Treg cell-mediated resistance to tumor growth.
Our data also highlight the interplay between PD-1 and STAT5 signaling. We have previously shown that PD-1 can inhibit STAT5 signaling in group 2 innate lymphoid cells, and this molecular mechanism is conserved in Treg cells. We found that PD-1 could inhibit STAT5 phosphorylation in Treg cells, albeit with a subtle effect, which then prevented expression of CD30 in these cells. There have been contradictory findings regarding the outcomes of PD-1 signaling in Treg cells in general, with some studies reporting inhibition of CD28 signaling, whereas others have shown TCR blockade. However, none of the studies considered the IL-2 that was present within the culture system or produced by contaminating T cells in the culture or Treg cells undergoing functional plasticity. By using an anti-IL-2 blocking antibody, we showed that in the absence of IL-2, CD30 is not induced in Treg cells. Hence, IL-2 through STAT5 drives CD30 expression, which is inhibited by PD-1 but only in the presence of anti-IL-2. Hence, within the TME resistance landscape, combination therapies targeting checkpoint receptors that are driven by IL-2 signaling could substantially dampen Treg cell function in the TME.
Our unbiased scRNA-seq analysis showed that CD30 expression post PD-1/PDL1 blockade was preserved in human Treg cells. However, our work using human samples was limited by the sample number; further confirmatory phenotyping could enhance the validity of our findings regarding the use of CD30 as a combination drug in melanoma patients who develop resistance. Of note, in silico analyses of published datasets show that CD30 is a predictive biomarker of survival in human cancers.
In summary, using unbiased analysis and murine validation studies, we report on the mechanistic role of PD-1 in Treg cell biology and function. This report consolidates previous contradictory findings and provides mechanistic insight into the importance of Treg cells within the TME in anti-PD-1 resistance.
Methods
Mice
C57BL/6, CD45.1 C57BL/6 and Rag1−/− mice were purchased from Charles River Laboratories, and Foxp3RFP and Foxp3ERT2-Cre-eGFP mice were purchased from Jackson Laboratory. C57BL/6 Pdcd1−/− (Pd1−/−) mice were provided by L. Chen (Yale University School of Medicine) and were further bred to Foxp3RFP mice to generate Pd1−/−Foxp3RFP mice. Pd1fl/fl-DTR-Tdtomato (Pd1fl/fl) mice were engineered by Ozgene. To ablate PD-1-expressing cells, Pd1fl/fl mice were given up to ten doses of 10 μg kg−1 of diphtheria toxin from Corynebacterium diphtheriae (Sigma-Aldrich) in PBS via intraperitoneal (i.p.) injection. Pd1fl/fl mice were further bred to Foxp3ERT2-Cre-GFP (Foxp3ERT2cre) mice to generate Pd1fl/flFoxp3ERT2cre and Pd1fl/flFoxp3ERT2cre+/− mice. To induce Cre-mediated PD-1 excision, Pd1fl/flFoxp3ERT2cre mice were given up to five doses of 1 mg tamoxifen (Sigma-Aldrich) in corn oil (Sigma-Aldrich) via i.p. injection. All mice were bred and maintained in a pathogen-free facility at Newcastle University under a project license approved by the Home Office. Male and female mice aged 8 to 12 weeks were used for all mouse strains, except for Foxp3ERT2cre and Pd1fl/flFoxp3ERT2cre+/− mice, where only females were used. All mouse experimental procedures were performed at Newcastle University incorporating the NC3R guidelines for animal research, and the data are presented according to the ARRIVE guidelines. Mice were maintained on a 12-h light–dark cycle, at a temperature of 21 ± 2 °C and humidity of 55 ± 5%. Mice were maintained in individually ventilated cages under positive pressure.
Patient characteristics
Human PBMCs from healthy donors were derived from NHS Blood and Transplant in Newcastle upon Tyne. Patients with stage IV metastatic melanoma were recruited from Royal Victoria Infirmary in Newcastle upon Tyne under Research Ethics Committee reference 24/NE/0014. Patients were treated with a combination therapy of ipilimumab and nivolumab. Blood samples were collected at the subsequent posttreatment clinic visit, at which point the antibody plasma concentration was three half-lives for both antibodies. All samples were collected after informed consent had been provided. In keeping with the literature,36 the effects of anti-PD-1 treatment were deemed minimal in this cohort, and the samples were labeled as ‘baseline’ samples. We also mined a publicly available dataset that included TILs from treatment-naive patients37 and patients treated with anti-PD-1 therapy, comprising both baseline and posttreatment samples. In this cohort, both responders and nonresponders to anti-PD-1 monotherapy were used for analysis. For responders, only posttreatment biopsy samples were used, to ensure comparability with the nonresponders37.
Reagents
Mouse complete medium was made up of DMEM (Gibco) supplemented with 10% fetal bovine serum (FBS), 2 mM l-glutamine (Corning), 0.1 mM nonessential amino acids, 1 mM sodium pyruvate (Sigma-Aldrich), 50 µM 2-mercaptoethanol, and penicillin and streptomycin (100 units ml−1 penicillin, 100 µg ml−1 streptomycin; Corning). Melanoma culture media was made up of DMEM (Gibco) supplemented with 10% FBS and penicillin and streptomycin (100 units ml−1 penicillin, 100 µg ml−1 streptomycin). Flow cytometry staining (FACS) buffer was made up of PBS supplemented with 0.5% bovine serum albumin (BSA) (Sigma-Aldrich) and 0.01% sodium azide. Miltenyi buffer was made up of PBS supplemented with 0.5% BSA and 2 mM EDTA (Sigma-Aldrich). Human PBMC culture medium was made up of X-VIVO 15 with gentamicin and phenol red (Lonza) formulated with 5% FBS. All murine and human antibodies were obtained from BioLegend unless otherwise stated.
Tumor model
The B16 melanoma model was set up using established protocols6,33. Mice were inoculated with 3 × 105 B16F10 melanoma cells via subcutaneous injection into the flank. Tumors were measured daily from day 5 postinoculation, and tumor volumes were calculated using the following equation: volume = π/6 × 0.5 × length × width2. In certain experiments, Pd1fl/fl and Pd1fl/flFoxp3ERT2cre mice were used and treated with PBS or 1 mg tamoxifen in corn oil, respectively, via i.p. injection for 5 consecutive days before tumor reconstitution. For Treg cell adoptive transfer experiments, Rag1−/− mice were reconstituted with 5 × 104 sorted CD4+CD25− cells from CD45.1 WT mice as Teff cells, along with 5 × 104 sorted CD4+FoxP3RFP cells as Treg cells from either Foxp3RFP or Pd1−/−Foxp3RFP mice via intravenous injection at day 8 postinoculation. In certain experiments, mice were treated with 0.1 mg isotype control IgG2a (2A3, Bio X Cell) or anti-CD153 (RM153, 2BScientific) via i.p. injection five times at 2-day intervals, starting from day 9. Mice were euthanized when tumor volumes reached 800–1,500mm3.
Flow cytometry
Single-cell suspensions from the indicated organs were stained with a Live/Dead fixable dead cell staining kit (Invitrogen) per the manufacturer’s instructions. Surface staining was performed for 30 min at 4 °C in FACS buffer with the following antibodies: CD3ε (clone: 145-2C11), CD30 (clone: mCD30.1), CD4 (clone: GK1.5), CD44 (clone: IM7), CD45.1 (clone: A20), CD45.2 (clone: 104), CD62L (clone: MEL-14), GARP (clone: YGIC86, eBioscience), GITR (clone: DTA-1, BD Biosciences), Nrp1 (clone: 3E12), PD-1 (clone: 29F.1A12), TIGIT (clone: 1G9), CD25 (clone: PC61), CD119 (clone: 2E2, eBioscience), ST2 (clone: DJ8, mdbioproducts), NK1.1 (clone: PK136), CD11c (clone: N418), PD-L1 (clone: 10F9G2), PD-L2 (clone: TY25), CD86 (clone: GL-1), CD80 (clone: 16-10A1), IA/IE (clone: M5/114.15.2), H2Kb (clone: AP6-88.5) and CD16/32 (clone: 93). For intracellular cytokine staining, cells were stimulated with a cell-stimulation cocktail containing protein transport inhibitors (eBioscience) for 4 h at 37 °C. Cells were then fixed and permeabilized using a FoxP3 Transcription Factor Staining Buffer Set (eBioscience) per the manufacturer’s instructions. Intracellular staining was performed overnight at 4 °C in 1× permeabilization buffer (eBioscience) with the following antibodies: Bcl2 (clone: 10C4, eBioscience), CD152 (clone: UC10-4B9), FoxP3 (clone: FJK-16S, eBioscience), Ki67 (clone: SolA15, eBioscience), Gata3 (clone: TWAJ, eBioscience), RORγt (clone: AFKJS-9, eBioscience), Tbet (clone: 4B10), Helios (clone: 22F6), IFNγ (clone: XMG1.2), IL-10 (clone: JES5-16ES), IL-17A (clone: TC11-18H10.1), GzmB (clone: QA16A02) and CD16/32 (clone: 93).
Human PBMCs were stained with a Live/Dead fixable dead cell staining kit (Invitrogen) per the manufacturer’s instructions. Cells were stained for 30 min at 4 °C with surface antibodies follows: CD4 (clone: RPA-T4), CD30 (clone: BY88), GARP (clone: 7B11), GITR (clone: 108-17) TIGIT (clone: A15153G) and CD16/32 (clone: 3G8, FUN-2 and 10.1). Cells were then fixed and permeabilized using a FoxP3 staining kit (eBioscience) per the manufacturer’s instructions. Intracellular staining was performed with antibodies including FoxP3 (clone: 206D) and CD152 (clone: BNI3) overnight at 4 °C.
Flow cytometry was performed using a BD LSR Fortessa X20. Sort purification was performed on a BD FACSAria Fusion. All data were analyzed using FlowJo 10 (Tree Star). Graphs and statistical analyses were done with SPICE 6.1 (https://niaid.github.io/spice/) and Prism v.10 (GraphPad Software).
Annexin V/7-AAD staining
Treg cells (CD4+CD25+) were isolated from Foxp3RFP or Pd1−/−Foxp3RFP mice using a CD4+CD25+ Regulatory T Cell Isolation Kit (Miltenyi Biotec) per the manufacturer’s instructions. Isolated Treg cells were cultured in mouse complete medium at 1 × 106 cells ml−1 and stimulated with 0.5 μg ml−1 anti-CD3 (clone: 145-2C11), 2 μg ml−1 anti-CD28 (clone: 37.51) and 80 ng ml−1 IL-2 (Miltenyi Biotec) for 72 h. After washing with cold PBS, Treg cells were stained with 5 μl Annexin V and 5 μl 7-AAD in 100 μl annexin V binding buffer (BD Biosciences) for 15 min in the dark at room temperature and analyzed by flow cytometry within 1 h.
Mouse and human PBMC cell preparation for BD Rhapsody
BD Rhapsody single-cell technology was utilized. For murine samples, isolated B16F10 melanoma TILs from C57BL/6 WT and Pd1−/− mice were stained with lineage markers for 20 min at 4 °C as follows: CD3ε (clone: 145-2C11), CD4 (clone: GK1.5), CD5 (clone: 53-7.3), CD8α (clone: 53-6.7), CD11b (clone: M1/70), CD11c (clone: N418), CD19 (clone: 6D5), CD49b (clone: DX5), Ter119 (clone: TER-119), F4/80 (clone: BM8), B220 (clone: RA3-6B2) and Gr1 (clone: RB6-8C5). After washing with Miltenyi buffer, cells were incubated with anti-Thy1 (clone: 30-H12) and streptavidin-Percp cy5.5 for 20 min at 4 °C. Cells were washed again and incubated with AbSeq antibodies and unique sample tags using a BD Mouse Single-Cell Multiplexing Kit (BD Biosciences) for 60 min on ice. The following AbSeq antibody-oligos were used: CD25, CD103, CD119, CD37, CD223, CD272, CD273, CD274, CD278, CD279, IL-17Rb, IL-23R, IL-33R, CD335, CD3 and NK1.1. After washing three times, cells were stained with DAPI and flow-enriched for the Lineage+ Thy1+ population consisting of T cells, innate immune cells and NK cells. For human samples, PBMC processing and Rhapsody experiments were performed as previously described38,39.
Single-cell library preparation and sequencing
Single-cell libraries were prepared according to the protocol provided by BD Biosciences. Briefly, a pooled sample tag, AbSeq and mRNA library was sequenced with minimum read lengths of 51 bp (R1) and 71 bp (R2) and a 200-cycle sequencing protocol. Libraries were sequenced on a NovaSeq 6000 (Illumina) using an S2 Reagent Kit v.1.5 (200 cycles).
Preprocessing of scRNA-seq data and dimensionality reduction
For murine TIL and human PBMC scRNA-seq datasets, FastQ files were processed using the BD Rhapsody analysis pipeline (https://www.sevenbridges.com/bdgenomics/), which generated unique molecular identifier (UMI) counts. In RStudio (R v.3.3), the Seurat package (v.5.0.3) was used to further analyze transcriptomic and protein expression data. For murine samples, cells were filtered to include those with ≥10 UMIs and 20–300 detected genes per cell. For human samples, cells were filtered to include those with 1,200–60,000 UMIs and 50–200 detected genes per cell. Multiplets and indeterminates were removed from downstream analysis. Expression matrices were log-normalized, and the top 2,000 variable features were identified. Samples were integrated using an anchor set and scaled by z-score conversion. Dimensionality reduction (npcs = 50) was performed using principal component analysis (PCA). The top ten and top 20 principal components (PCs) were used to compute UMAP for murine and human scRNA-seq. A k-nearest neighbor graph (KNN) was constructed using the FindNeighbors function (k.param = 20, dims = 1:10 for murine and 1:20 for human, annoy.metric = ‘euclidean’), and Louvain clustering (resolution = 0.6) was performed.
For mining of publicly available datasets, the data files were preprocessed37,40, and the processed gene expression data were retrieved from the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) database (http://www.ncbi.nlm.nih.gov/geo). In RStudio, the Seurat package was used to identify the top 2,000 variable features. Dimensionality reduction (npcs = 50) was performed using PCA. The top ten and top 20 PCs were used to compute UMAPs for treatment-naive melanoma patients and for anti-PD-1-treated melanoma patients at baseline and after treatment, respectively. A KNN graph was constructed using the FindNeighbors function (k.param = 20, dims = 1:10 for treatment-naive melanoma patients and 1:20 for anti-PD-1-treated melanoma patients, annoy.metric = ‘euclidean’). Louvain clustering was performed with a resolution of 0.2 for treatment-naive melanoma patients and 0.1 for anti-PD-1-treated melanoma patients.
scRNA-seq analysis
For scRNA-seq analysis of murine TILs, cluster identities were assigned using the ScType function based on cell-type-specific gene sets (Supplementary Table 3a), resulting in nine clusters (basophils, Teff cells, macrophages, megakaryocytes, myeloid DCs, B cells, NK cells, plasmacytoid DCs and Treg cells) in murine TILs. The Teff cell cluster was further subclustered into CD4 central memory T, CD4 Teff, CD8 central memory T and CD8 Teff cell clusters. For scRNA-seq analysis, of human PBMCs, cluster identities were assigned using the ScType function based on cell-type-specific gene sets (Supplementary Table 6a). In addition, cells were annotated using a manual review process based on algorithmically defined marker gene expression for each cluster with the FindAllMarkers function (method = ‘Wilcoxon’ with Bonferroni correction), together with protein expression data, resulting in 16 clusters of human PBMCs (B cells, CD8+ Tc1 cells, CD11c+CD14+, CD11c+CD16+, CD4+ Teff, CD8+ Teff, γδ, CD4+ memory T, CD8+ memory T, CD4+ T naive, CD8+ T naive, CD161+ NK, NK, CD8+ Tc17, Treg and other). For public datasets, cluster identities were annotated based on algorithmically defined marker gene expression for each cluster with the FindAllMarkers function (method = ‘Wilcoxon’ with Bonferroni correction). This process revealed nine clusters (CD4 T cells, CD8 T cells, CD4−CD8− T cells, B cells, macrophages, endothelial cells, cancer-associated fibroblasts, melanoma cells and NK cells) for treatment-naive patients and six clusters (CD4 T cells, CD8 T cells, B cells, monocytes and macrophages, DCs, and NK and NK T cells) for anti-PD-1-treated patients in which the top preferentially expressed genes (adjusted P < 0.001) included multiple known markers of particular cell types. The CD4 T cell cluster was further divided into CD4 T cells without FoxP3 expression and Treg cells with FoxP3 expression in both datasets. Gene expression of clusters was visualized as a dot plot using the JJDotPlot function in the scRNAtoolVis package (v.0.1.0).
Differential gene expression and pseudotime trajectory analysis
Differential gene expression was determined using the FindMarkers function (method = ‘Wilcoxon’ with Bonferroni correction). For this analysis, AbSeq antibodies were excluded. Genes were considered to be differentially expressed if they achieved log2 (fold change) > 0.585 and adjusted P < 0.05. The results were illustrated as a volcano plot using the ggplot2 (v.3.5.0) package. To overcome the limitations of sample pooling, mRNA expression levels of coinhibitory receptors were obtained and plotted to show expression in each individual sample. The average mRNA expression of selected genes in Treg cells across individuals was calculated using the mean function with summarise from the dplyr package (v.1.1.4). Unpaired Student’s t-test was performed to measure differences between groups. In addition, single-cell mRNA expression was visualized using violin plots generated with the VlnPlot function in the Seurat package, and clustered Wilcoxon rank-sum test41 was performed using the clusWilcox.test function (method = ‘ds’) in the clusrank (v.1.0-4) package to overcome sample pooling effects while measuring differences in coinhibitory receptor expression between two groups. GO overrepresentation analysis and GSEA were performed using the clusterProfiler (v.4.10.1) package. Gene symbols were converted into Entrez ID using the biomaRt package (v.3.18). The enriched GO terms were acquired from the list of differentially expressed genes using the enrichGO function and visualized as a dot plot using the dotplot function. Statistical analysis was performed using Fisher’s exact test with false discovery rate (FDR) correction, and adjusted P values are shown. GSEA was conducted using the gseGO function; genes were ranked using the default Kolmogorov–Smirnov test with FDR correction, and a P value cutoff of 0.05 was used to identify significant GO terms. Pseudotime single-cell trajectory analysis was performed using the Monocle2 plugin in SeqGeq.
Cell–cell communications
To determine cell–cell interactions and signaling patterns, we applied the cell communications CellChat (v.2) R package to our single-cell analysis (https://github.com/sqjin/CellChat)42. CellChat cross-references a ligand–receptor interaction database (CellChatDB) including extracellular matrix–receptor interactions, cell–cell contact interactions and secreted signals. To identify significant cell–cell interactions, differentially expressed ligands and receptors were first determined using the identifyOverExpressedGenes and identifyOverExpressedInteractions functions with default parameters. Communications with fewer than three cells were excluded with the filterCommunication function. The communication probability was calculated with the computeCommunProbPathway function, and significant communications were identified using a permutation test. An aggregated cell–cell communication network was then compiled using the aggregateNet function; this network was visualized as a circle plot using the netVisual_circle function and as a chord plot using the netVisual_chord_gene function. Network centrality scores were then calculated using the netAnalysis_computeCentrality function, and the role of each cell type as a signal sender or receiver within the inferred communication network were quantified using the netAnalysis_signalingRole_heatmap function. Cophenetic and Silhouette values were evaluated by using the selectK function to identify the optimal number of communication patterns of cell types. Outgoing and incoming patterns were then determined using the identifyCommunicationPatterns function to reveal how the sender cells coordinated with certain outgoing signaling pathways to drive communication or how the target cells responded to incoming signaling pathways.
PD-L1 molecular mechanistic studies
Cell-culture 96-well plates were prepared by coating with either 5 μg ml−1 anti-CD3 alone or 5 μg ml−1 anti-CD3 and recombinant PD-L1 Fc chimera protein for 3 h at 37 °C. The wells were washed with PBS three times. Treg cells were then seeded in the wells and cultured in mouse complete medium at 1 × 106 cells ml−1, along with 2 μg ml−1 anti-CD28 (clone: 37.51). In certain wells, 10 μg ml−1 anti-PD-1 (clone: RMP1.14) was added to block PD-1–PD-L1 interaction. After 72 h, cells were washed with complete medium and further stimulated with either 10 μg ml−1 anti-IL-2 (clone: S4B6, Invitrogen) alone or 10 μg ml−1 anti-IL-2 and 80 ng ml−1 IL-2 (Miltenyi Biotec) for 24 h in a humidified incubator at 37 °C with 5% CO2. Cells were then harvested for flow cytometry and quantitative reverse transcription polymerase chain reaction (RT–qPCR) assays.
Treg cell suppressor assays
The Treg cell suppressor assay was performed as previously published27. CD90.2T cells were purified from splenocytes isolated from Foxp3RFP or Pd1−/−Foxp3RFP mice using CD90.2 MicroBeads (Miltenyi Biotec) per the manufacturer’s instructions. Cells were stained with anti-CD4, and cell viability was assessed by DAPI before sorting. Sorted Teff cells were stained in PBS with cell trace violet. Then, 5 × 104 Teff cells and the indicated ratios of sorted Treg cells from either Foxp3RFP or Pd1−/−Foxp3RFP mice were stimulated using 0.5 μg ml−1 anti-CD3 along with irradiated splenocytes from C57BL/6 WT mice as antigen-presenting cells using a 4:1 ratio of antigen-presenting cells to Teff cells. Cells were cultured in complete medium for 72 h at 37 °C with 5% CO2, and proliferation was assessed as the dilution of cell trace violet using flow cytometry.
Phospho-STAT5 assays
Phospho-STAT5 assays using coated PD-L1 Fc chimera proteins were performed using our established protocols4,18. Cell-culture plates were prepared by coating with 5 μg ml−1 anti-CD3 and 5 μg ml−1 recombinant PD-L1 Fc chimera protein for 3 h at 37 °C. Control wells were coated with anti-CD3 alone. The wells were washed with PBS three times. Isolated Treg cells were then cultured in mouse complete medium at 1 × 106 cells ml−1 and incubated with 2 μg ml−1 anti-CD28 and 10 μg ml−1 anti-IL-2 for 48 h in a humidified incubator at 37 °C with 5% CO2. In certain wells, 10 μg ml−1 anti-PD-1 was added to block PD-1–PD-L1 interaction. After 48 h, cells were washed with complete medium and further stimulated with 80 ng ml−1 IL-2 for 15 min at 37 °C. Cell fixation and permeabilization were performed using BD Phosflow Lyse/Fix buffer and BD Phosflow Perm Buffer III per the manufacturer’s instructions. Cells were then stained with anti-pSTAT5 (clone: 47; BD Biosciences) for 20 min at room temperature and analyzed by flow cytometry
RNA extraction from Treg cells
mRNA was extracted from Treg cells using mRNeasy Kits (Qiagen) per the manufacturer’s protocol. cDNA synthesis from mRNA was carried out using TaqMan Reverse Transcription Reagents (Applied Biosystems).
RT–qPCR
cDNAs were used as the template for RT–qPCR using a LightCycler 480 II (Roche). TaqMan Gene Expression assays (FAM) were purchased from Life Technologies Thermo Fisher Scientific for quantification of mouse CD30 mRNA (Mm00437140_m1 (Tnfrsf8)), as well as actin mRNA (Mm00607939_s1 (Actb)), which was used as a housekeeping control. All TaqMan assay RT–qPCRs were performed using TaqMan Fast Advanced Master Mix (Applied Biosystems). The RT–qPCR reaction included an initial step of 10 min at 95 °C followed by 50 cycles of 15 s denaturation at 95 °C and then 60 s annealing and extension at 60 °C.
RT–qPCR data analysis
Results were processed using LightCycler 480 software. Samples were assigned to control (WT Treg) and experimental (Pd1−/− Treg) groups. Cycle threshold (Ct) values were then normalized, with Actb as a reference gene, using the following formula: ΔCt = average Ct (Tnfrsf8) − average Ct (Actb). The difference between the ΔCt values for the WT Treg cells and Pd1−/− Treg cells was calculated using the formula ΔΔCt = ΔCt (Pd1−/− Treg) − ΔCt (WT Treg). The expression fold change was then calculated as follows: fold change = \(2^{-\Delta\Delta C_{\rm{t}}}\).
CosMx tissue processing
Tumors were harvested from B16F10 melanoma-bearing Rag1−/− mice reconstituted with CD45.1+ WT Teff cells and either WT Treg cells or Pd1−/− Treg cells and fixed in 10% neutral buffered formalin (Sigma-Aldrich) overnight. Tumor tissues were embedded in paraffin, punched as 3-mm biopsies and mounted onto the centers of Leica Bond Plus slides. Formalin-fixed paraffin-embedded slides were air-dried at room temperature after sectioning. Hematoxylin and eosin staining of slides was performed by NovoPath at Newcastle Hospitals, and the stained slides were observed under a Zeiss Axio Imager 2 and analyzed using ZEISS ZEN software to assess tissue block quality and select FOVs.
CosMx image acquisition and cell segmentation
CosMx experiments were performed by NanoString as part of the Technology Access Program. Briefly, standard methods for fluorescence in situ hybridization were applied by NanoString to formalin-fixed paraffin-embedded tissue slides to expose RNA targets as previously described43. Fluorescent bead-based fiducials fixed to the tissue were introduced to provide an optical reference for cyclic image acquisition, followed by hybridization of RNA-specific probes and antibodies. After washing, each sample was assembled into a flow cell and loaded onto a spatial molecular imaging (SMI) instrument for RNA readout and morphological imaging. Raw images were transformed to decoded RNA transcripts at subcellular resolution through a workflow including (1) three-dimensional primary image processing to identify and register reporter spots; (2) decoding of reporter spots to RNA transcripts with registered x, y and z spatial locations; (3) outlining of nucleus and cell boundaries with DAPI and antibodies after cyclic reporter readout for morphology-based cell segmentation; and (4) assigning RNA transcripts to single cells.
CosMx data analysis
CosMx SMI data and flat files were exported from AtoMx SIP to a QWS S3 Bucket. The CosMx SMI data output consisted of a Seurat object, TileDB array, and a ‘Flow Cell ID’ folder containing raw decoded files and images for each FOV, whereas the flat file output consisted of a count matrix, cell metadata, global transcripts, global cell boundaries and global FOV positions. Both output files were downloaded using the S3 console and used for data analysis. Samples were loaded into RStudio (R v.4.3.3) using the LoadNanostring function, and expression profiles of cells were clustered using the Seurat package (v.5.0.0). Briefly, data were normalized and scaled using the SCTransform function. Dimensionality reduction (number of principle components, npcs = 5) was performed using PCA, and the top five PCs were used to compute UMAPs. A KNN graph was constructed using the FindNeighbors function (k.param = 20, dims = 1:5, annoy.metric = ‘euclidean’), followed by Louvain clustering with a resolution of 0.2. Cluster identities were assigned using the ScType function based on cell-type-specific gene sets and published papers on cell identification in skin and tumor tissue44,45, resulting in five clusters in the murine TME (basal cells, immune cells, Schwann cells, tumor cells and melanocytes). The ‘immune cells’ cluster was further subclustered into monocytes, DCs, innate lymphoid cells, mast cells, Treg cells, NK cells, macrophages and CD4+ T cells. Each FOV region was zoomed in and visualized using the Crop function. Using ImageDimPlot function, we overlaid the cell identities onto the FOV images. Spatial expression pattern of individual molecules was visualized using the ImageFeaturePlot function. Visualization of gene expression, calculation of average gene expression, differential gene expression analysis and enrichment analysis were performed as described for the scRNA-seq analysis. Clustered Wilcoxon rank-sum tests were performed using the clusWilcox.test or pairwise.clusWilcox.test function (method = ‘ds’) in clusrank (v.1.0-4) to measure differences in coinhibitory receptor expression between two groups or among multiple groups, with FDR correction applied for comparisons involving more than two groups.
CosMx spatially resolved cell–cell interaction analysis
The Treg cell population was further divided into several groups based on spatial location and cell interactions in FOVs using specific Cell_IDs. The numbers of physical interactions of Treg cells with other cell types in the FOV images were manually counted. Using the FindAllMarkers function (method = ‘Wilcoxon’ with Bonferroni correction), specific genes in each Treg cell subset interacting with neighboring cells or alone within the FOV were determined by visually annotating cells. Treg cell-specific genes were further subdivided into either SR or LR communication categories based on their biological function by manual annotation. Genes involved in cell–cell contact were annotated as SR, and secretion or signaling genes were annotated as LR. The SR category consisted of genes for cell surface receptors, proliferation markers and apoptotic markers, whereas LR genes encoded cytokines, chemokines, transcription factors that could drive cytokine and chemokine production, and so on.
CHIP–seq analysis
The STAT5 CHIP–seq dataset was derived from NCBI GEO under accession number GSE207265 (ref. 29). In brief, naive WT CD4+ cells were stimulated with 10 μg ml−1 plate-bound anti-CD3 (17A2, Bio X Cell) and anti-CD28 (37.51, Bio X Cell) in the presence of anti-mouse IL-2 (S4B6) for 48 h. Cells were then washed and cultured with 100 U ml−1 human IL-2 (NIH/National Cancer Institute Biological Resources Branch Preclinical Repository) for 48 h. Genome browser files were rendered with Integrated Genome Viewer.
RNA-seq analysis
The RNA-seq dataset was derived from NCBI GEO under accession number GSE207265 (ref. 29). In brief, naive WT and Stat5−/− CD4+ cells were cultured with 10 μg ml−1 plate-bound anti-CD3 (clone: 17A2) and anti-CD28 (clone: 37.51) in the presence of anti-mouse IL-2 (clone: S4B6) for 72 h. Cells were then washed and restimulated with an anti-TCR agonist with either anti-IL-2 or 100 U ml−1 human IL-2 (NIH/National Cancer Institute Biological Resources Branch Preclinical Repository) for 4 days. For GSEA-based ranking of TNFRSF family members, transcripts of the TNF family including Tnfrsf8, Tnfrsf4, Tnfrsf13b, Tnfrsf11b, Tnfrsf18, Tnfrsf21, Tnfrsf1b, Tnfrsf9, Tnfrsf17, Tnfrsf19, Ltbr, Eda2r, Tnfrsf11a, Ngfr, Tnfrsf1a, Tnfrsf12a, Tnfrsf10a, Cd40, Tnfrsf13c, Fas, Tnfrsf25, Cd27 and Tnfrsf14 were ranked by rank metric score.
In vitro culture of melanoma patient PBMCs
Human PBMCs derived from patients with stage IV metastatic melanoma were plated at 1 × 106 cells ml−1 in a 96-well U-bottomed plate. Cells were stimulated with 0.5 μg ml−1 anti-CD3 (clone: OKT3) and 20 μg ml−1 of the isotype control IgG2b (clone: MPC-11) or anti-PD-L1 (clone: 29E.2A3) for 72 h in a humidified incubator at 37 °C with 5% CO2.
CD30 biomarker analysis
Analysis of CD30 as a biomarker in different cancers was performed on the R2 Genomics Analysis and Visualization Platform. The dataset analyzed was derived from The Cancer Genome Atlas and NCBI GEO under accession numbers GSE42568 (ref. 46) and GSE33114 (ref. 47). Kaplan–Meier survival curves with the default log-rank test were used to determine the optimum expression cutoff for survival analyses.
Statistical analysis
All data shown are expressed as individual data points with the line at the mean unless otherwise stated. Error bars on graphs show the standard error of the mean. Paired or unpaired two-tailed Student’s t-test or two-tailed clustered Wilcoxon rank-sum test was used for comparisons between two groups; and one- or two-way analysis of variance corrected by Sidak’s multiple comparison test, Wilcoxon rank-sum test corrected by Bonferroni correction or clustered Wilcoxon rank-sum test corrected by FDR was used for comparisons among multiple groups.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
scRNA-seq data from treatment-naive and anti-PD-1-treated melanoma patients used in this paper were accessed via GEO under accession codes GSE72056 and GSE120575 for the Tirosh and Sade-Feldman datasets, respectively. All raw data from the mouse single-cell BD Rhapsody RNA-seq experiments and human single-cell BD Rhapsody experiments, along with the NanoString CosMx data, have been uploaded to GEO: scRNA-seq data have been deposited under accession code GSE273532, human scRNA-seq data under accession code GSE280319, and the NanoString CosMx SMI dataset (both the Seurat object and flat files) under accession code GSE273530. The flat files were used for the analysis presented in this manuscript. The CellChat database for mouse datasets used in the cell–cell communication analysis is available via GitHub at https://github.com/sqjin/CellChat.
Code availability
The code used in analyzing the dataset is available via GitHub at https://github.com/JXLIM99/scRNAseq-and-Nanostring-Cosmx-SMI.
References
Abbas, A. K. et al. Regulatory T cells: recommendations to simplify the nomenclature. Nat. Immunol. 14, 307–308 (2013).
Amarnath, S. et al. Regulatory T cells and human myeloid dendritic cells promote tolerance via programmed death ligand-1. PLoS Biol. 8, e1000302 (2010).
Francisco, L. M. et al. PD-L1 regulates the development, maintenance, and function of induced regulatory T cells. J. Exp. Med. 206, 3015–3029 (2009).
Amarnath, S. et al. The PDL1-PD1 axis converts human TH1 cells into regulatory T cells. Sci. Transl. Med. 3, 111ra120 (2011).
Keir, M. E., Butte, M. J., Freeman, G. J. & Sharpe, A. H. PD-1 and its ligands in tolerance and immunity. Annu Rev. Immunol. 26, 677–704 (2008).
Stathopoulou, C. et al. PD-1 inhibitory receptor downregulates asparaginyl endopeptidase and maintains Foxp3 transcription factor stability in induced regulatory T cells. Immunity 49, 247–263.e247 (2018).
Kamada, T. et al. PD-1+ regulatory T cells amplified by PD-1 blockade promote hyperprogression of cancer. Proc. Natl Acad. Sci. USA 116, 9999–10008 (2019).
Kim, M. J. et al. Deletion of PD-1 destabilizes the lineage identity and metabolic fitness of tumor-infiltrating regulatory T cells. Nat. Immunol. 24, 148–161 (2023).
Tan, C. L. et al. PD-1 restraint of regulatory T cell suppressive activity is critical for immune tolerance. J. Exp. Med. 218, e20182232 (2021).
van Gulijk, M. et al. PD-L1 checkpoint blockade promotes regulatory T cell activity that underlies therapy resistance. Sci. Immunol. 8, eabn6173 (2023).
Perry, J. A. et al. PD-L1-PD-1 interactions limit effector regulatory T cell populations at homeostasis and during infection. Nat. Immunol. 23, 743–756 (2022).
Chen, X. et al. PD-1 regulates extrathymic regulatory T-cell differentiation. Eur. J. Immunol. 44, 2603–2616 (2014).
Okazaki, T., Maeda, A., Nishimura, H., Kurosaki, T. & Honjo, T. PD-1 immunoreceptor inhibits B cell receptor-mediated signaling by recruiting src homology 2-domain-containing tyrosine phosphatase 2 to phosphotyrosine. Proc. Natl Acad. Sci. USA 98, 13866–13871 (2001).
Chemnitz, J. M., Parry, R. V., Nichols, K. E., June, C. H. & Riley, J. L. SHP-1 and SHP-2 associate with immunoreceptor tyrosine-based switch motif of programmed death 1 upon primary human T cell stimulation, but only receptor ligation prevents T cell activation. J. Immunol. 173, 945–954 (2004).
Yokosuka, T. et al. Programmed cell death 1 forms negative costimulatory microclusters that directly inhibit T cell receptor signaling by recruiting phosphatase SHP2. J. Exp. Med. 209, 1201–1217 (2012).
Kamphorst, A. O. et al. Rescue of exhausted CD8 T cells by PD-1-targeted therapies is CD28-dependent. Science 355, 1423–1427 (2017).
Hui, E. et al. T cell costimulatory receptor CD28 is a primary target for PD-1-mediated inhibition. Science 355, 1428–1433 (2017).
Taylor, S. et al. PD-1 regulates KLRG1+ group 2 innate lymphoid cells. J. Exp. Med. 214, 1663–1678 (2017).
Franceschini, D. et al. PD-L1 negatively regulates CD4+CD25+Foxp3+ Tregs by limiting STAT-5 phosphorylation in patients chronically infected with HCV. J. Clin. Invest. 119, 551–564 (2009).
Laba, S., Mallett, G. & Amarnath, S. The depths of PD-1 function within the tumor microenvironment beyond CD8+ T cells. Semin. Cancer Biol. 86, 1045–1055 (2022).
Weiss, J. M. et al. Neuropilin 1 is expressed on thymus-derived natural regulatory T cells, but not mucosa-generated induced Foxp3+ T reg cells. J. Exp. Med. 209, 1723–1742 (2012).
Yadav, M. et al. Neuropilin-1 distinguishes natural and inducible regulatory T cells among regulatory T cell subsets in vivo. J. Exp. Med. 209, 1713–1722 (2012).
Thornton, A. M. et al. Expression of Helios, an Ikaros transcription factor family member, differentiates thymic-derived from peripherally induced Foxp3+ T regulatory cells. J. Immunol. 184, 3433–3441 (2010).
Odorizzi, P. M., Pauken, K. E., Paley, M. A., Sharpe, A. & Wherry, E. J. Genetic absence of PD-1 promotes accumulation of terminally differentiated exhausted CD8+ T cells. J. Exp. Med. 212, 1125–1137 (2015).
Strauss, L. et al. Targeted deletion of PD-1 in myeloid cells induces antitumor immunity. Sci. Immunol. 5, eaay1863 (2020).
Rubtsov, Y. P. et al. Stability of the regulatory T cell lineage in vivo. Science 329, 1667–1671 (2010).
Thornton, A. M. & Shevach, E. M. CD4+CD25+ immunoregulatory T cells suppress polyclonal T cell activation in vitro by inhibiting interleukin 2 production. J. Exp. Med. 188, 287–296 (1998).
Gaspal, F. et al. Abrogation of CD30 and OX40 signals prevents autoimmune disease in FoxP3-deficient mice. J. Exp. Med. 208, 1579–1584 (2011).
Villarino, A. V. et al. A central role for STAT5 in the transcriptional programing of T helper cell metabolism. Sci. Immunol. 7, eabl9467 (2022).
Zeiser, R. et al. Early CD30 signaling is critical for adoptively transferred CD4+CD25+ regulatory T cells in prevention of acute graft-versus-host disease. Blood 109, 2225–2233 (2007).
Laurence, A. et al. STAT3 transcription factor promotes instability of nTreg cells and limits generation of iTreg cells during acute murine graft-versus-host disease. Immunity 37, 209–222 (2012).
Amarnath, S. & Brown, M. L. Harnessing proteases for T regulatory cell immunotherapy. Eur. J. Immunol. 50, 770–778 (2020).
Lim, J. X. et al. Programmed cell death-1 receptor-mediated regulation of Tbet+NK1.1− innate lymphoid cells within the tumor microenvironment. Proc. Natl Acad. Sci. USA 120, e2216587120 (2023).
Helou, D. G. et al. PD-1 pathway regulates ILC2 metabolism and PD-1 agonist treatment ameliorates airway hyperreactivity. Nat. Commun. 11, 3998 (2020).
Heinhuis, K. M. et al. Safety, tolerability, and potential clinical activity of a glucocorticoid-induced TNF receptor-related protein agonist alone or in combination with nivolumab for patients with advanced solid tumors: a phase 1/2a dose-escalation and cohort-expansion clinical trial. JAMA Oncol. 6, 100–107 (2020).
Brahmer, J. R. et al. Phase I study of single-agent anti-programmed death-1 (MDX-1106) in refractory solid tumors: safety, clinical activity, pharmacodynamics, and immunologic correlates. J. Clin. Oncol. 28, 3167–3175 (2010).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998–1013.e1020 (2018).
McTaggart, T. et al. Deep phenotyping of T regulatory cells in psoriatic arthritis highlights targetable mechanisms of disease. J. Biol. Chem. 301, 108059 (2025).
McTaggart, T. et al. Deciphering novel communication patterns in T regulatory cells from very old adults. Aging Cell https://doi.org/10.1111/acel.70044 (2025).
Tirosh, I. et al. Single-cell RNA-seq supports a developmental hierarchy in human oligodendroglioma. Nature 539, 309–313 (2016).
Jiang, Y., Lee, M.-L. T., He, X., Rosner, B. & Yan, J. Wilcoxon rank-based tests for clustered data with R package clusrank. J. Stat. Softw. 96, 1–26 (2020).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Jacob, T. et al. Molecular and spatial landmarks of early mouse skin development. Dev. Cell 58, 2140–2162.e2145 (2023).
Kim, Y. S. et al. Single-cell RNA sequencing reveals the existence of pro-metastatic subpopulation within a parental B16 murine melanoma cell line. Biochem. Biophys. Res. Commun. 613, 120–126 (2022).
Clarke, C. et al. Correlating transcriptional networks to breast cancer survival: a large-scale coexpression analysis. Carcinogenesis 34, 2300–2308 (2013).
de Sousa E Melo, F. et al. Methylation of cancer-stem-cell-associated Wnt target genes predicts poor prognosis in colorectal cancer patients. Cell Stem Cell 9, 476–485 (2011).
Acknowledgements
We thank E. Kowalczyk and D. Hinson (BD, Biosciences Bioinformatics Support Team) for their help with quality control analysis of our data and for providing us with the support for data analysis. We also thank the organizers of the British Society of Immunology RStudio training program and the RStudio single-cell analysis training provided by BD Rhapsody. This work was funded by an MRC project grant (MR/T015586/1), Leo Foundation Future Leader Award (LF-AW_EMEA-19-4000), Leo Foundation Serendipity Grant (LF-SE-23-800015), Lister Prize, Academy of Medical Sciences Springboard Award (SBF003_1129_6), the NIHR Newcastle Biomedical Research Centre awarded to the Newcastle upon Tyne Hospitals NHS Foundation Trust, Newcastle University and Cumbria, Northumberland, Tyne and Wear Foundation Trust. J.X.L. is supported by a Newcastle University Overseas Research Scholarship award and a NC3R PhD studentship (NC/W002043/1), and T.M. is supported by a Barbour foundation PhD studentship. Finally, we thank R. Heiser, B. Grogan and B. P. O’Connor from Seagen/Pfizer for their insightful advice on CD30 and its related expression in human Treg cells in cancers; Y. Wang, Postdoctoral Research Fellow at the Department of Physics and Astronomy, University of Exeter, for insightful discussions on code generation for single-cell analysis; and J. O’Shea, Scientific Director of NIAMS, NIH, for reading the manuscript and for assistance with obtaining statistical help with analysis of our scRNA-seq and spatial transcriptomics data.
Author information
Authors and Affiliations
Contributions
J.X.L. and T.M. performed experiments, analyzed data, wrote methods and edited the manuscript. K.J.S., G.H., S.L., R.H. and J.C. performed experiments. I.C. recruited melanoma patients for the study. S.J., J.N. and H.S. provided intellectual input and guidance on computational biology analysis. P.L. holds ethics for the human work, and provided patient samples and edited the manuscript. G.A. provided intellectual input and edited the manuscript. A.L. and S.A. conceptualized the idea, wrote the manuscript and supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Tobias Bopp, Gabriela Fragiadakis and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Nick Bernard, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–9.
Supplementary Tables 1–6 (download XLSX )
This file contains multiple tabs of supplementary tables, each clearly labeled in the spreadsheet.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lim, J.X., McTaggart, T., Jung, S.K. et al. PD-1 receptor deficiency enhances CD30+ Treg cell function in melanoma. Nat Immunol 26, 1074–1086 (2025). https://doi.org/10.1038/s41590-025-02172-0
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-025-02172-0
This article is cited by
-
Post-translational modifications of immune checkpoints: molecular mechanisms, tumor microenvironment remodeling, and therapeutic implications
Journal of Biomedical Science (2026)
-
KIRA6 restrains the generation of myeloid-derived suppressor cells and overcomes resistance to anti-PD-1 therapy
Cell Death & Disease (2025)