Abstract
Pathogen-specific CD4⁺ T cells expand and contract during infection, generating memory clones that shape subsequent immune responses. How distinct tissue environments influence differentiation and clonal selection of polyclonal T cells remains unclear. Here we develop Tracking Recently Activated Cell Kinetics (TRACK) mice, a dual-recombinase fate-mapping system enabling the spatial and temporal labeling of recently activated CD4⁺ T cells. Using TRACK mice during influenza infection, we observed organ-specific transcriptional differentiation and clonal selection in lung, mediastinal lymph nodes (medLNs) and spleen. During the effector phase, spleen-derived CD4⁺ T cells adopted a stem-like migratory phenotype, whereas medLN-activated cells differentiated into T follicular helper cells. T cell receptor sequencing showed low clonal overlap between tissues during the effector response, consistent with distinct antigenic landscapes. During memory formation, overlap increased between lung- and medLN-derived cells, while splenic clones retained a distinct repertoire. These findings define tissue-dependent mechanisms that shape CD4⁺ T cell fate and clonal architecture.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
scRNA-seq data generated in this study have been deposited in the Gene Expression Omnibus under accession code GSE317035. Source data are provided with this paper.
Code availability
Custom scripts supporting the data analysis are available via GitHub at https://github.com/tiagobrc/parsa_2024.
References
Littman, D. R. & Rudensky, A. Y. Th17 and regulatory T cells in mediating and restraining inflammation. Cell 140, 845–858 (2010).
Taniuchi, I. CD4 helper and CD8 cytotoxic T cell differentiation. Annu. Rev. Immunol. 36, 579–601 (2018).
O’Shea, J. J. & Paul, W. E. Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science 327, 1098–1102 (2010).
Zhu, J., Yamane, H. & Paul, W. E. Differentiation of effector CD4 T cell populations. Annu. Rev. Immunol. 28, 445–489 (2010).
Sallusto, F. Heterogeneity of human CD4+ T cells against microbes. Annu. Rev. Immunol. 34, 317–334 (2016).
Tanoue, T., Atarashi, K. & Honda, K. Development and maintenance of intestinal regulatory T cells. Nat. Rev. Immunol. 16, 295–309 (2016).
Honda, K. & Littman, D. R. The microbiota in adaptive immune homeostasis and disease. Nature 535, 75–84 (2016).
Jenkins, M. M. et al. Lung dendritic cells migrate to the spleen to prime long-lived TCF1hi memory CD8+ T cell precursors after influenza infection. Sci. Immunol. 6, eabg6895 (2021).
Masopust, D., Vezys, V., Marzo, A. L. & Lefrancois, L. Preferential localization of effector memory cells in nonlymphoid tissue. Science 291, 2413–2417 (2001).
Schenkel, J. M. & Masopust, D. Tissue-resident memory T cells. Immunity 41, 886–897 (2014).
Masopust, D. & Soerens, A. G. Tissue-resident T cells and other resident leukocytes. Annu. Rev. Immunol. 37, 521–546 (2019).
Mueller, S. N. & Mackay, L. K. Tissue-resident memory T cells: local specialists in immune defence. Nat. Rev. Immunol. 16, 79–89 (2016).
Mayassi, T., Barreiro, L. B., Rossjohn, J. & Jabri, B. A multilayered immune system through the lens of unconventional T cells. Nature 595, 501–510 (2021).
Faria, A. M. C., Reis, B. S. & Mucida, D. Tissue adaptation: implications for gut immunity and tolerance. J. Exp. Med. 214, 1211–1226 (2017).
Parsa, R. et al. Newly recruited intraepithelial Ly6A+CCR9+CD4+ T cells protect against enteric viral infection. Immunity 55, 1234–1249 (2022).
Crowl, J. T. et al. Tissue-resident memory CD8+ T cells possess unique transcriptional, epigenetic and functional adaptations to different tissue environments. Nat. Immunol. 23, 1121–1131 (2022).
Sathaliyawala, T. et al. Distribution and compartmentalization of human circulating and tissue-resident memory T cell subsets. Immunity 38, 187–197 (2013).
Gebhardt, T. et al. Different patterns of peripheral migration by memory CD4+ and CD8+ T cells. Nature 477, 216–219 (2011).
Dash, P. et al. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 547, 89–93 (2017).
Moon, J. J. et al. Tracking epitope-specific T cells. Nat. Protoc. 4, 565–581 (2009).
Yang, Y. et al. Focused specificity of intestinal TH17 cells towards commensal bacterial antigens. Nature 510, 152–156 (2014).
Bousbaine, D. et al. A conserved Bacteroidetes antigen induces anti-inflammatory intestinal T lymphocytes. Science 377, 660–666 (2022).
Ansaldo, E. et al. Akkermansia muciniphila induces intestinal adaptive immune responses during homeostasis. Science 364, 1179–1184 (2019).
Qi, Q. et al. Diversity and clonal selection in the human T-cell repertoire. Proc. Natl Acad. Sci. USA 111, 13139–13144 (2014).
Krueger, P. D. et al. Two sequential activation modules control the differentiation of protective T helper-1 (Th1) cells. Immunity 54, 687–701 (2021).
Turner, D. L., Bickham, K. L., Farber, D. L. & Lefrançois, L. Splenic priming of virus-specific CD8 T cells following influenza virus infection. J. Virol. 87, 4496–4506 (2013).
Ciucci, T. et al. The emergence and functional fitness of memory CD4+ T cells require the transcription factor Thpok. Immunity 50, 91–105 (2019).
Cardenas, M. A. et al. Differentiation fate of a stem-like CD4 T cell controls immunity to cancer. Nature https://doi.org/10.1038/s41586-024-08076-7 (2024).
Stolley, J. M. et al. Retrograde migration supplies resident memory T cells to lung-draining LN after influenza infection. J. Exp. Med. 217, e20192197 (2020).
Rodger, B., Stagg, A. J. & Lindsay, J. O. The role of circulating T cells with a tissue resident phenotype (ex-TRM) in health and disease. Front. Immunol. 15, 1415914 (2024).
Heim, T. A. et al. Lymphatic vessel transit seeds cytotoxic resident memory T cells in skin draining lymph nodes. Sci. Immunol. 9, eadk8141 (2024).
Hutchinson, E. C. et al. Conserved and host-specific features of influenza virion architecture. Nat. Commun. 5, 4816 (2014).
Bogdanow, B. et al. The dynamic proteome of influenza A virus infection identifies M segment splicing as a host range determinant. Nat. Commun. 10, 5518 (2019).
Andreatta, M. et al. A CD4+ T cell reference map delineates subtype-specific adaptation during acute and chronic viral infections. Elife 11, e76339 (2022).
Victora, G. D. & Nussenzweig, M. C. Germinal centers. Annu. Rev. Immunol. 40, 1–30 (2022).
Ferreira, D. P. et al. Central memory CD8+ T cells derive from stem-like Tcf7hi effector cells in the absence of cytotoxic differentiation. Immunity 53, 985–1000 (2020).
Lee, J.-Y. et al. The transcription factor KLF2 restrains CD4+ T follicular helper cell differentiation. Immunity 42, 252–264 (2015).
Carlson, C. M. et al. Kruppel-like factor 2 regulates thymocyte and T-cell migration. Nature 442, 299–302 (2006).
Wu, H. et al. Modeling of influenza-specific CD8+ T cells during the primary response indicates that the spleen is a major source of effectors. J. Immunol. 187, 4474–4482 (2011).
Heath, W. R. & Carbone, F. R. Dendritic cell subsets in primary and secondary T cell responses at body surfaces. Nat. Immunol. 10, 1237–1244 (2009).
León, B., Ballesteros-Tato, A., Randall, T. D. & Lund, F. E. Prolonged antigen presentation by immune complex-binding dendritic cells programs the proliferative capacity of memory CD8 T cells. J. Exp. Med. 211, 1637–1655 (2014).
Kedmi, R. & Littman, D. R. Antigen-presenting cells as specialized drivers of intestinal T cell functions. Immunity 57, 2269–2279 (2024).
Friedl, P. & Gunzer, M. Interaction of T cells with APCs: the serial encounter model. Trends Immunol. 22, 187–191 (2001).
Wherry, E. J. & Kurachi, M. Molecular and cellular insights into T cell exhaustion. Nat. Rev. Immunol. 15, 486–499 (2015).
Wijeyesinghe, S. et al. Expansible residence decentralizes immune homeostasis. Nature https://doi.org/10.1038/s41586-021-03351-3 (2021).
Khatun, A. et al. Single-cell lineage mapping of a diverse virus-specific naive CD4 T cell repertoire. J. Exp. Med. 218, e20200650 (2021).
Gerlach, C. et al. One naive T cell, multiple fates in CD8+ T cell differentiation. J. Exp. Med. 207, 1235–1246 (2010).
Tubo, N. J. et al. Single naive CD4+ T cells from a diverse repertoire produce different effector cell types during infection. Cell 153, 785–796 (2013).
Künzli, M. et al. Long-lived T follicular helper cells retain plasticity and help sustain humoral immunity. Sci. Immunol. 5, eaay5552 (2020).
Snook, J. P., Kim, C. & Williams, M. A. TCR signal strength controls the differentiation of CD4+ effector and memory T cells. Sci. Immunol. 3, eaas9103 (2018).
Fazilleau, N., McHeyzer-Williams, L. J., Rosen, H. & McHeyzer-Williams, M. G. The function of follicular helper T cells is regulated by the strength of T cell antigen receptor binding. Nat. Immunol. 10, 375–384 (2009).
Zehn, D., Lee, S. Y. & Bevan, M. J. Complete but curtailed T-cell response to very low-affinity antigen. Nature 458, 211–214 (2009).
Tubo, N. J. & Jenkins, M. K. TCR signal quantity and quality in CD4+ T cell differentiation. Trends Immunol. 35, 591–596 (2014).
Takahashi, M. et al. Intratumoral antigen signaling traps CD8+ T cells to confine exhaustion to the tumor site. Sci. Immunol. 9, eade2094 (2024).
Jones, M. C. et al. CD4 Eeffector TCR avidity for peptide on APC determines the level of memory generated. J. Immunol. 210, 1950–1961 (2023).
Quadros, R. M. et al. Easi-CRISPR: a robust method for one-step generation of mice carrying conditional and insertion alleles using long ssDNA donors and CRISPR ribonucleoproteins. Genome Biol. 18, 92 (2017).
Harms, D. W. et al. Mouse genome editing using the CRISPR–Cas system. Curr. Protoc. Hum. Genet. 83, 15.7.1–15.7.27 (2014).
Dash, P. et al. Paired analysis of TCRα and TCRβ chains at the single-cell level in mice. J. Clin. Invest. 121, 288–295 (2011).
Han, A., Glanville, J., Hansmann, L. & Davis, M. M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G. & Neufeld, J. D. PANDAseq: paired-end assembler for illumina sequences. BMC Bioinformatics 13, 31 (2012).
Brochet, X., Lefranc, M.-P. & Giudicelli, V. IMGT/V-QUEST: the highly customized and integrated system for IG and TR standardized V-J and V-D-J sequence analysis. Nucleic Acids Res. 36, W503–W508 (2008).
Nagashima, K. et al. Mapping the T cell repertoire to a complex gut bacterial community. Nature https://doi.org/10.1038/s41586-023-06431-8 (2023).
Acknowledgements
We thank A. Rogoz and J. Bortolatto for embryo bioengineering, animal care, mouse colony management and genotyping; RU Genomics core for assistance with sequencing; K. Gordon and J. P. Truman for sorting; and Y. Khan and additional Rockefeller University employees for continuous assistance. We thank the NIH Tetramer Facility for providing the LLO and NP tetramers. We thank all the members of the Mucida, Victora (Rockefeller) and Lafaille (NYU) laboratories for fruitful discussions. This work was supported by The Swedish-American Foundation and The Swedish Research Council (to R.P.; grant no. 2023/02507-7), the São Paulo Research Foundation (H.CA.) and a Life Sciences Research Foundation postdoctoral fellowship (to T.B.R.d.C.). G.L.d.R. was supported by a scholarship from Fundação Maria Emília, FME (Brazil); NIH grants R01DK093674 and R01DK113375; NIAID grant P01AI179273; the CZI Science, Food Allergy FARE/FASI Consortium; Stavros Niarchos Foundation Institute for Global Infectious Diseases; and The Howard Hughes Medical Institute (to D.M.).
Author information
Authors and Affiliations
Contributions
Conceptualization by R.P. and D.M. Investigation by R.P., H.C.A., T.B.R.d.C., G.L.d.R., A.S. and A.M.B. Writing—original draft by R.P. and D.M. Editing by all authors. Funding acquisition by R.P. and D.M. Resources by T.B.R.d.C. and H.H. Supervision by D.M.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Immunology thanks Adrian Liston and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: S. Houston in collaboration with the Nature Immunology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Tracking of pathogen-specific CD4 + T cells during intestinal and respiratory infection.
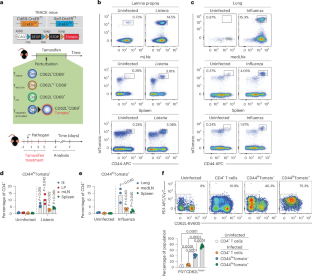
a, Overview of genetic strategy regarding Cd69-CreER and Sell-DreER mice. b, Gating strategy for CD4+CD44hiTomato+ T cells. c, TRACK mice were infected with L. monocytogenes and CD8+CD44hiTomato+ T cells were analyzed at day 9 post infection. Representative plot. Minimum 3 independent experiments, For infected, n = 4 for IE, LP and spleen, n = 3 for medLN. For control, n = 3. Statistical significance was tested between infected organ to uninfected organ by two-tailed Student’s T-test. d, TRACK mice were treated with tamoxifen as described and medLN, spleen and lung were analyzed for tdTomato labeling of CD4+CD44hi T cells 9 days post-influenza PR8 infection. n = 3/group, data from one experiment. e, Stated mice were infected with L. monocytogenes and CD4+CD44hiTomato+ were analyzed in the LP 9-day post infection. Representative plot from 2 independent experiments, n = 3 per group. One-way analysis of variance (ANOVA) with Tukey’s multiple-comparison test. f, T cells from the spleen of TRACK mice were isolated and stimulated with anti-CD3/anti-CD28 with or without 4-Hydroxytamoxifen (4-OHT). CD69, CD62L and Tomato expression on CD4+ T cells were measured over time. Pooled data from two experiments, n = 6. Data are presented as mean ± SEM. g, Similar to e, but T cells were stimulated with IFN-γ, IFN-β or IL-2 without CD3/CD28 stimulation. CD69, CD62L, CD44 and Tomato expression were measured on CD4+ T cells 24 hours post stimulation. Data from 4 mice. h, i, Analysis of CD4+CD44lowTomato+ within the spleen at 4, 8 or 13 weeks of age. Representative plot of minimum 3 independent experiments. For h, n = 3 in uninfected group and n = 4 in infected group. For I, n = 3 in 4 weeks and 13 weeks group. In the 8 weeks group, n = 3 in uninfected group and n = 4 in infected group. Two-tailed Student’s T-test. j, Analysis of CD69 and CD62L expression within the thymus in TRACK mice at 8 weeks of age. Pooled data from 2 independent experiments, n = 6. One-way analysis of variance (ANOVA) with Tukey’s multiple-comparison test.
Extended Data Fig. 2 Analysis of RNAseq data from influenza infected TRACK mice at day 9 and day 56 post infection.
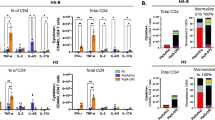
a and b, CD45.1 isogenic B6 mice were infected with influenza PR8 and treated with anti-CD62L at day 1 and 3. NP-specific naive 500.000 CD4 T cells were transferred to the recipients at day 2 post infection and the transferred T cells in the medLN and spleen were analyzed for activation markers at day 4 post infection. Statistical differences were determined with two-tailed Student’s t-test c, Analysis of CD4+CD44hiTomato+ T cells across timepoints and organs in TRACK mice post-influenza PR8 virus infection. Pooled data from 5 independent experiments n = 17. d, Cell counts across clusters divided by either organs (left) or mice (right) at day 9 post infection, n = 3. e, Cell counts across clusters divided by either organs (left) or mice (right) at day 56 post infection, n = 4. f, Gene signatures used for classifications in UMAP plots. g, All expanded clonotypes (clonal size >10) at day 9 post infection were analyzed to determine the Gini index per clonotype across medLN, spleen and lung. The top 25th and 75th quantile of the Gini index were determined and those selected clonotypes, that is low Gini index clonotypes ( < 0.26) or high Gini index clonotypes ( > 0.57) where pooled and analyzed for cluster distribution. Outer numbers: Cluster, Outer ring: organ origin, Inner ring: Cluster size, Inner number: Clone counts across organs. Pooled data from 3 mice.
Extended Data Fig. 3 Distinct clonal expansion and distribution across organs.
a, Clonotype size ranking from scRNAseq data across organs and timepoints. Dashed x-axis line indicates top 10 expanded clones. Pool ranked clonotypes from 3 mice for d9 p.i. and from 4 mice for d56 p.i. b, Cluster and organ distribution of top 5 expanded clones located in the lungs at day 9 post infection. Each dot represents one unique mouse. n = 3. c, Cluster and organ distribution of top 5 expanded clones located in the lungs at day 56 post infection. Each dot represents one unique mouse. n = 4.
Extended Data Fig. 4 Analysis of TCR specificity and functional avidity in fate-mapped T cells.
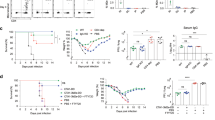
a, Experimental overview of testing TCR specificity with murine T cell hybridomas. The 15 most expanded TCRs across lungs, medLN and spleen from day 9 (b) and day 56 (d) post infection stimulated with anti-CD3 to determine functionality. TCRs are isolated from the scRNA-seq data in Fig. 3. c, TCR hybridomas from day 9 (c) or day 56 (e)were tested for viral specificity by co-culturing them with splenic DCs that had been pulsed with 105 plaque forming units (pfu) of PR8 particles for 24 hours. In day 9 group n = 14 and in day 56 group n = 15 unique T cell hybridomas. Representative data from 2 independent experiments. f, TRACK mice were infected with influenza PR8 and CD4+ T cells were enriched from the lungs, medLN and spleen, 9 or 56 days post infection. The isolated CD4 T cells were stained with CFSE and co-cultured with splenic DCs that were pulsed with various amounts of PR8 viral particles for 48 hours. Pooled data from 2 independent experiments, n = 5-7/group. One-way analysis of variance (ANOVA) with Tukey’s multiple-comparison test, per dilution point. Statistics determined at 104 PFU. g, Clonotype bias at day 9 (g) or day 56 (h) post infection (p.i.), analysis only considered expanded clonotypes with 5 or more cells and corrected for clonotype size by generating expected background distributions by random permutation (dotted line). Red circles indicate Z score > 1.97 and two-sided p < 0.05. Pooled RNAseq data from 3 individual (d9) or 4 individual mice (d56) mice. i, NP311-325-specific TCR hybridomas was co-cultured with splenic CD11c+ DCs and stimulated with various concentrations of NP311-325 peptide (10 µM, 5 µM, 2.5 µM, 1.25 µM, 0.63 µM and 0 µM) for 24 hours and NFAT-GFP expression was measured.
Supplementary information
Supplementary Information (download PDF )
Supplementary Fig. 1.
Supplementary Table 1 (download XLSX )
TCRs used for hybridomas, isolated from most expanded clones across organs.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Fig. 5 (download XLSX )
Statistical source data.
Source Data Fig. 6 (download XLSX )
Statistical source data.
Source Data Fig. 7 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Parsa, R., Assis, H.C., de Castro, T.B.R. et al. Tissue-specific clonal selection and differentiation of CD4⁺ T cells during infection. Nat Immunol 27, 725–737 (2026). https://doi.org/10.1038/s41590-026-02451-4
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41590-026-02451-4