Abstract
Nonparenchymal central nervous system (CNS)-associated macrophages (CAMs) mediate immune responses at brain boundaries. Perivascular and leptomeningeal CAMs are collectively termed subdural CAMs (sdCAMs). Both sdCAMs and juxtaneuronal microglia are derived from embryonic yolk sac precursors, long-living and maintain their populations through self-renewal. Following depletion, microglia autonomously repopulate from single surviving cells. In contrast, the course of sdCAM repopulation remains poorly understood. Here, by combining multilineage fate mapping, multiomic profiling and high-resolution imaging, we demonstrate divergent repopulation dynamics between sdCAMs and microglia. Unlike microglia, sdCAMs do not renew cell-autonomously, but become transiently accessible to CCR2+Ly6C+ monocyte engraftment after niche induction in an integrin-dependent manner. Moreover, replenished monocyte-derived sdCAMs remain transcriptomically, epigenetically and functionally distinct from their embryo-derived counterparts. Finally, we present a protocol enabling selective exchange of sdCAMs, modulating disease response without functionally affecting microglia. These new insights into CNS immune biology suggest new therapeutic avenues for neuroinflammatory and neurodegenerative diseases.
Similar content being viewed by others
Main
The central nervous system (CNS) hosts different tissue-resident macrophages. Microglia seed the parenchyma, whereas CNS-associated macrophages (CAMs)1, also known as border-associated macrophages (BAMs)2, are found at the CNS interfaces. CAMs have been further subdivided based on their anatomical location. The outermost meningeal layer of the CNS, the dura mater, is home to dural CAMs (dCAMs)3. Subdural CAMs (sdCAMs) include perivascular macrophages (pvMΦ), positioned in the Virchow-Robin space, as well as leptomeningeal macrophages (lmMΦ) and are thus situated at the parenchymal surface4, hence their alternative naming as parenchymal border macrophages5. CNS macrophages are involved in a plethora of perturbations such as inflammatory, degenerative or neurooncological disorders, where they shape the disease course1,6,7,8.
Microglia and lmMΦ originate during embryogenesis from a common c-Kit+ erythromyeloid progenitor in the extraembryonic YS4,9,10,11. PvMΦ are part of the lmMΦ pedigree, as they emerge during the early postnatal phase from descending lmMΦ that migrate into the developing Virchow-Robin space in a talin-1-mediated fashion12. Of note, and in contrast to other CAM subsets such as dCAMs, microglia, lmMΦ and pvMΦ are self-maintaining throughout life under homeostatic conditions and are not replaced by any circulating blood cells3,4,13,14,15.
For their survival, microglia and CAMs depend on constant signaling through the colony-stimulating factor 1 receptor (CSF-1R). Consequently, genetic deletion as well as blockade of CSF-1R signaling by inhibitors, such as PLX3397, PLX5622 or BLZ945, leads to effective depletion of both microglia and CAMs9,16,17,18,19. Previous studies employing various depletion strategies elucidated the repopulation features of microglia in mice20,21,22. Through fate mapping-based lineage-tracing, these studies demonstrated that, following a single depletion course, repopulated microglia emerge exclusively from the proliferation of surviving microglia without input from circulating blood cells18,22. Only if the endogenous microglial compartment is continuously or severely compromised, can monocytes replace microglia, representing an exception to the physiological rule23,24.
While depletion and repopulation of microglia is being actively tested as a therapeutic strategy for microgliopathies25,26,27,28, similar studies for CAMs are still very sparse18,29,30. Their low abundance, combined with the historical lack of tools to comprehensively and selectively investigate them, has hindered the study of the cellular sources, underlying molecular mechanisms and disease-related functional consequences of CAM repopulation.
Divergent spatiotemporal replenishment of CAMs and microglia
Following a single transient depletion, microglia rapidly repopulate, indicating a high demand for replenishing the juxtaneuronal niche20,21. To investigate the renewal kinetics of CAMs (Fig. 1a), we temporarily administered BLZ945, a brain penetrant CSF-1R inhibitor, to wild-type (WT) mice (Fig. 1b)16,17.
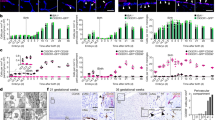
a, Overview of microglia (MG) and subsets of CAMs during homeostasis in the CNS. dCAMs, dural CAMs; sdCAMs, subdural CAMs, including leptomeningeal (lmMΦ) and perivascular (pvMΦ) macrophages. b, Left: Scheme of the experimental layout. Right: CNS macrophage repopulation in complementary myeloid-specific fate mapping mouse lines was analyzed by confocal microscopy, flow cytometry, RNA-seq and scATAC-seq, followed by systemic LPS challenge and a middle-cerebral artery occlusion (MCAO) model of thromboembolic stroke. c,d, Representative flow cytometry plots of MG and sdCAMs after depletion (c) and quantification of their numbers per brain hemisphere (d). Symbols represent individual mice, n = 3 (CTL and 1 d BLZ945), n = 4 (8w, 12w, 26w BLZ945), n = 5 (5 d BLZ945), n = 6 (2w BLZ945), mean ± s.e.m. Ordinary one-way analysis of variance (ANOVA) with Šidák-adjusted multiple-comparisons P values. e, Representative confocal images from cerebral cortex, cerebellum and olfactory tubercle at 1 d, 5 d and 8w after BLZ945 treatment in WT mice. Arrows point to myeloid cells in their distinct compartments. f, Proportion of MG, lmMΦ and pvMΦ density at different time points and in diverse brain regions after BLZ945 treatment compared to controls (CTL). Symbols represent mean depletion at each time point, calculated as proportion of CTL mean. Error bars indicate s.e. of the depletion estimate, propagated from the standard errors of BLZ945 and CTL densities. n = 3 (CTL and 1 d BLZ945), n = 4 (8w, 12w, 26w BLZ945), n = 5 (5 d BLZ945), n = 6 (2w BLZ945), logistic growth modeling, t90%: time point in days when the repopulation reaches 90% compared to controls. R2: coefficient of determination for model equation and data points). Unpaired two-tailed Welch’s t-test with Bonferroni-corrected P values.
Flow cytometry-based cell quantification of the whole brain revealed a robust reduction of microglia and sdCAM numbers compared to controls (Fig. 1c, d). While microglia had mostly recovered after 5 days (5 d) of repopulation, sdCAMs were still largely absent. At 8 weeks (8w) after depletion; however, both microglia and sdCAM numbers were fully restored. As microglial self-renewal kinetics differ between brain regions13,31, we examined the repopulation of IBA1+CD206− microglia as well as CD206+ lmMΦ and pvMΦ in cortex, cerebellum and olfactory tubercle (Fig. 1e, f). Histological assessment confirmed successful depletion of both microglia and sdCAMs at 1 d, followed by a swift re-emergence of microglia, whereas only few lmMΦ and pvMΦ were observed early during repopulation (Fig. 1e, f). An evaluation of the renewal kinetics revealed quick and uniform re-establishment of microglia (t90% of controls: 6.20 d in cerebral cortex, 6.00 d in cerebellum, 5.91 d in olfactory tubercle). In sharp contrast, repopulation of both lmMΦ and pvMΦ occurred at greatly reduced velocity and in a spatially distinct manner (t90% for lmMΦ, 15.44 d cerebral cortex, 12.29 d cerebellum, 19.28 d olfactory tubercle; and t90% for pvMΦ, 21.35 d, 15.66 d, 28.59 d, respectively).
Repopulating sdCAMs intensify cell–cell communications
CNS macrophages closely interact with diverse neighboring cells, forming a niche that provides essential signals shaping their phenotype, function, and survival1,32. To investigate the local tissue environments during repopulation, we performed single-cell RNA-sequencing (scRNA-seq) of CD11b+CD45int-hi myeloid cells together with CD31+ endothelial cells, nonhematopoietic PDPN+ as well as lineage negative (Lin−) cells from micro-dissected CNS compartments of control mice after depletion (Fig. 2a and Supplementary Fig. 1a). Analyzing 39,838 individual cells (Fig. 2b), we classified immune and niche cells according to canonical marker expression (Extended Data Fig. 1a), their tissue of origin (Extended Data Fig. 1b) and cell subtype markers previously identified3,33,34,35,36,37 (Supplementary Fig. 1b). We found that depletion and repopulation significantly affected macrophage abundance and composition with little impact on niche cells or other immune cells (Extended Data Fig. 1c). At 5 d after depletion, both homeostatic microglia and CAM populations were reduced, whereas repopulating macrophage subtypes (dividing Mac1, dividing Mac2, repop MG1 and repop MG2) arose, characterized by expression of Mki67 and other cell-cycle genes (Supplementary Fig. 1b). Differential expression analysis revealed only mild effects of CNS macrophage depletion and repopulation on gene expression in niche cells (Supplementary Fig. 2). Similarly, expression levels of factors known to regulate CNS macrophage survival, phenotype and function showed no clear differences (Extended Data Fig. 1d).
a, Experimental scheme: C57BL/6 mice were treated either with BLZ945 or kept as controls (CTL). Cranial dura mater, enriched cortical leptomeninges and cortex without meninges were isolated and CD11b+CD45int-hi, PDPN+, CD31+ and Lin− cells were sorted for scRNA-seq. n = 4 (dura 8w and 5 d), n = 3 (dura CTL), n = 8 (leptomeninges and cortex 8w and 5 d), n = 7 (leptomeninges and cortex CTL). b, Uniform Manifold Approximation and Projection (UMAP) of 39,838 individual cells with cell type annotation. c, Predicted incoming and outgoing interaction strengths for all cell types from leptomeninges and cortex were calculated using CellChat. Size of dots represents number of predicted interactions. d, Top ten predicted interaction partners by frequency of interaction between myeloid cell types and their surrounding niche in leptomeninges and cortex by time point (CTL, 5 d, 8w). e, Chord plot depicting predicted interactions between the two homeostatic CAM subtypes and the top interacting niche cell types. f,g, Chord plot depicting interactions between macrophages and either niche cells (f) or clMonocytes (g) that are differentially upregulated at 5 d after BLZ945. h, Proportion of macrophage clusters among all macrophages in leptomeninges and cortex samples. Proportions are derived from averages across leptomeningeal and cortical samples. ara, arachnoid; art, arterial; diff, differentiating; du, dural; EC, endothelial cells; FB, fibroblast; hom, homeostatic; ieg, immediate early genes; Mac, macrophage; myel, myelinating; nc, non-classical; Oligo, oligodendrocytes; PC, pericytes; repop, repopulating; SC, Schwann cells; ven, venous; VSMC, vascular smooth muscle cell; OPC, oligodendrocyte progenitor cell; cDC1/2, classical dendritic cell 1/2; migDC, migratory dendritic cell; NKT, natural killer T cell; Th2, T helper 2 cell.
To further explore local macrophage-niche interactions dynamics in the subdural compartment, we deployed CellChat. Assessing predicted outgoing versus incoming signaling strength, we visually identified three clusters of cells (Fig. 2c). Group 1, mainly composed of macrophages, were predicted to be the strongest signal receivers, with repopulating subtypes showing higher outgoing interaction strength than homeostatic counterparts. Group 2, including fibroblasts, endothelial cells and pericytes, were predicted to act as strong senders, whereas group 3 showed low interaction strength (Fig. 2c). The overall interaction strength showed a spike at 5 d post-depletion and a return to baseline 8w after depletion (Extended Data Fig. 1e). When investigating the contribution of niche-macrophage–monocyte interactions to this phenomenon in the subdural compartment, we found the highest interaction frequencies between repopulating macrophage subtypes and the vascular and leptomeningeal stroma (Fig. 2d). In control animals, signaling pathways from endothelial, mural and fibroblastic stromal cells contributed in a balanced way to sdCAM niche homeostasis (Fig. 2e). Assessing pathways upregulated during repopulation, we identified perivascular fibroblast-derived Il34 as the most prominent signal received by repopulating sdCAMs (Fig. 2f).
By examination of subclustered subdural macrophage populations we found early repopulating CNS macrophages to express Nestin, in agreement with previous findings22 (Extended Data Fig. 1f, g). Repopulating subdural macrophage subsets and classical monocytes (clMonocyte) were among the cell types with highest interaction frequencies at 5 d (Fig. 2d). Between these cells, we found CCR2-mediated signaling, a canonical monocyte attraction pathway, to be the most prominent predicted interaction (Fig. 2g). Furthermore, repopulating CNS macrophages highly expressed Tnf and chemokines, including Ccl3, Ccl4, Ccl6, Ccl7 and Ccl9, whereas the corresponding receptors were found on monocytes and endothelial cells (Extended Data Fig. 1h). Notably, the relative abundance of sdCAM subtypes had changed 8w after depletion compared to controls (Fig. 2h). Taken together, transcriptional profiling revealed that CNS macrophage-niche cells were largely unaffected; however, the change in sdCAM composition, combined with the upregulation of monocyte-attracting chemokines, led us to investigate the cellular origins of sdCAM repopulation.
sdCAMs and microglia renew from distinct cellular sources
To determine whether sdCAMs repopulate completely cell-autonomously, similarly to microglia18,22, we employed several myeloid fate mapping mouse models. First, we performed depletion experiments in Cx3cr1CreERT2R26tdT mice (Extended Data Fig. 2a). As expected4,33,38, short living CX3CR1+Ly6Chi monocytes quickly lost labeling after tamoxifen (TAM) injection (Extended Data Fig. 2b). In contrast, flow cytometry-based (Extended Data Fig. 2c) and histological analyses (Extended Data Fig. 2d, e) revealed constant high labeling in fully renewed microglia at all time points after depletion (Extended Data Fig. 2c). Conversely, sdCAMs showed a clear reduction in tdTomato+ (tdT+) proportion, with many sdCAMs remaining tdT− 12w after depletion, suggesting noncell-autonomous sources of their repopulation.
To decipher the cellular source of replenished microglia and sdCAMs, we next took advantage of myeloid subset-specific inducible CreER lines that selectively label either microglia (HexbCreERT2R26YFP)18, CAMs (Mrc1CreERT2R26tdT)3,12 or hematopoietic stem cell (HSC)-derived cells (Cxcr4CreERT2R26tdT)12,39,40 (Fig. 3a, b and Extended Data Fig. 3a). While Ly6Chi or Ly6Clo monocytes were not labeled in HexbCreERT2R26YFP and Mrc1CreERT2R26tdT mice, these cells were efficiently labeled in Cxcr4CreERT2R26tdT animals (Extended Data Fig. 3b–d). Notably, microglia showed consistent labeling in the cortex of HexbCreERT2R26YFP mice across all time points, whereas only single lmMΦ and pvMΦ were YFP+ (Fig. 3c, d). In contrast, labeling of renewed cortical lmMΦ and pvMΦ in Mrc1CreERT2R26tdT mice dropped compared to controls, whereas virtually no tdT+ microglia were observed (Fig. 3e, f). Of note, besides some consistent labeling in nonimmune cells (Supplementary Fig. 3), brains of Cxcr4CreERT2R26tdT animals showed substantial tdT labeling in lmMΦ and pvMΦ after depletion, but not in microglia (Fig. 3g, h and Extended Data Fig. 3e–k). After depletion, both resident yolk sac (YS)- as well as HSC-derived sdCAMs were positive for the proliferation marker Ki-67 (Extended Data Fig. 3l). In summary, our data demonstrate cell-autonomous repopulation of microglia. In stark contrast, sdCAMs utilize both surviving remnants as well as peripheral HSC-derived cells to refill their niches in a heteronomous manner.
a,b, Experimental scheme. c, Representative images of YFP labeling in cerebral cortex from HexbCreERT2R26YFP mice at 8w after CTL or BLZ945 treatment. Arrowheads depict CD206+YFP− cells. d, Quantification of YFP labeled MG, lmMΦ and pvMΦ in HexbCreERT2R26YFP mice at different time points post CTL or BLZ945 treatment. Symbols represent individual mice, n = 2 (2w CTL pvMΦ), n = 3 (2w CTL MG and lmMΦ, 26w), n = 4 (2w BLZ945, 8w), mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. e, Typical immunofluorescence images of tdT labeling in cerebral cortex from Mrc1CreERT2R26tdT mice at 8w after treatment. Arrowheads, blank indicate CD206+tdT+ cells; orange show CD206+tdT− cells. f, Quantification of tdT labeled MG, lmMΦ and pvMΦ in Mrc1CreERT2R26tdT mice at different time points after treatment. Symbols represent individual mice, n = 3 (2w CTL), n = 4 (2w BLZ945 and 26w CTL), n = 5 (8w BLZ945 and 26w BLZ945) n = 6 (8w CTL), mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. g, Representative pictures of tdT labeling in the cerebral cortex from Cxcr4CreERT2R26tdT mice at 8w after treatment. Arrowheads, blank show CD206+tdT− cells; orange show CD206+tdT+ cells. h, Quantification of tdT labeled proportions of MG, lmMΦ and pvMΦ in Cxcr4CreERT2R26tdT mice at different time points after treatment in cerebral cortex, cerebellum and olfactory tubercle. Symbols represent individual mice, n = 3 (5 d CTL, 2w CTL, 8w CTL and 12w CTL), n = 4 (2w BLZ945, 8w BLZ945 and 12w BLZ945), n = 5 (5 d BLZ945, 26w), mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values.
Adhesion molecules shape transient infiltration of monocytes
Having identified descendants of CXCR4+ HSCs as a partial source of replenished sdCAMs, we next aimed to uncover their direct progenitor. As Ly6ChiCCR2hi monocytes are known to differentiate into brain macrophages upon perturbation23,41, we labeled them during the first two weeks of repopulation using Ccr2CreERT2R26tdT mice (Fig. 4a and Extended Data Fig. 4a–c). We detected a significant increase of replenished tdT+CD206+ sdCAMs, whereas microglia consistently remained unlabeled (Fig. 4a, b). Conversely, labeling Ly6ChiCCR2hi monocytes 8w post-depletion, when microglia and sdCAM numbers were fully restored, did not lead to an increase in tdT+ sdCAM proportion (Fig. 4c), revealing only transient monocyte engraftment after niche induction.
a,b, Experimental scheme (top). Recombination rates in MG and sdCAMs were determined by flow cytometry (FC) (a) or immunofluorescence (IF) (b). Typical immunofluorescence pictures of cortices from Ccr2CreERT2R26tdT mice 8w after treatment and quantification of IBA1+tdT+ MG and CD206+tdT+ CAMs. Arrowheads, yellow show CD206+tdT− CAMs; blank show CD206+tdT+ CAMs. Symbols represent individual mice, n = 4 (14 d), n = 6 (8w CTL), n = 9 (8w BLZ945), mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. c, Experimental scheme (top). FC quantification of tdT+ MG and sdCAMs. Symbols represent individual mice, n = 3 per group, mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. d, Diagram of integrins and their receptors in cellular extravasation. e, Experimental scheme (top). FC quantification of tdT+ MG or sdCAMs. n = 5 per group. Symbols represent individual mice, mean ± s.e.m. Two-tailed t-test. f, Experimental scheme (top left). Representative pictures of anti-ICAM-1 immunofluorescence (right) and quantification of ICAM-1+CD31+ cerebral cortex vasculature 5 d after treatment (bottom left). Symbols represent individual mice, n = 3 per group, mean ± s.e.m. Unpaired two-tailed t-test. g, Experimental scheme (top left). Representative MRI images (right) and quantification thereof (bottom left). Symbols represent individual mice, n = 5 per group, mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. MPIO, microparticles of iron oxide. h, Experimental scheme (top left). FC quantification of tdT+ MG or CAMs (bottom left). Typical immunofluorescence pictures of cortices from Ccr2CreERT2R26tdT mice 21 d after treatment (right). n = 9 (isotype), n = 8 (anti-VCAM-1/ICAM-1). Symbols represent individual mice, mean ± s.e.m. Unpaired two-tailed t-test.
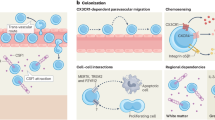
To explore the underlying mechanisms of this monocytic influx, we first checked the gene expression of adhesion molecules and integrins guiding monocytic extravasation42 (Fig. 4d) in our scRNA-seq dataset (Extended Data Fig. 4d). While monocytes expressed high levels of integrin α-4 (Itga4) as well as Itgb1 and Itgb2, CNS endothelial cells showed expression of Icam1, Vcam1, Sele and Selp (Extended Data Fig. 4d). Notably, no differential expression of the endothelial adhesion molecules P-Selectin (CD62P) and E-Selectin (CD62E) was detectable at protein level 5 d after depletion (Extended Data Fig. 4e). To test which molecules mediate monocyte infiltration, we next targeted integrin receptors. Injection of very late antigen-4 (VLA-4) neutralizing antibodies during the first two weeks of repopulation in Ccr2CreERT2R26tdT mice, did not significantly decrease the proportion of tdT+ monocyte-derived sdCAMs at 21 d after depletion (Fig. 4e). Similarly, CD45.2+ ITGB2-deficient monocytes did not show impaired engraftment into the sdCAM compartment (Extended Data Fig. 4f–i).
Therefore, we next assessed the integrin receptors intercellular adhesion molecule 1 (ICAM-1; Fig. 4f) and vascular cell adhesion molecule 1 (VCAM-1; Fig. 4g) on brain endothelial cells. Both proteins were found to be upregulated 5 d after sdCAM depletion (Fig. 4f, g). To evaluate their functional relevance, we injected Ccr2CreERTR26tdT mice with ICAM-1 and VCAM-1 blocking antibodies during the first 2 weeks of repopulation and found reduced amounts of tdT+ monocyte-derived sdCAMs (Fig. 4h). Taken together, our data show that sdCAM repopulation is partially dependent on an early but transient engraftment of circulating monocytes, which is mediated by interactions with VCAM-1/ICAM-1 on endothelial cells.
dCAMs repopulate through amplified monocyte infiltration
In contrast to sdCAM niches and the CNS parenchyma, the healthy dura mater is readily accessible to monocytes, making dCAMs a heterogenous population of mostly HSC origin3,29. We therefore compared the repopulation kinetics and cellular origin of dCAMs to those observed for sdCAMs and microglia (Extended Data Fig. 5a, b). While dCAMs were strongly reduced at 1 d after depletion, they reached normal numbers by 5 d (Extended Data Fig. 5c). Analysis of their cellular origin under control conditions clearly revealed continuous exchange of dCAMs by HSC-derived cells, with CD206lo dCAMs showing a higher rate of homeostatic exchange compared to CD206hi dCAMs (Extended Data Fig. 5d–h). After depletion, both CD206hi and CD206lo dCAMs were replaced to similar near-complete rates, exceeding those seen in sdCAMs (Extended Data Fig. 5d–h). We did not detect significant differences in the amount of monocyte-derived dCAMs between extrasinusoidal and sinusoidal compartments3 8w after depletion (Extended Data Fig. 5i, j). We found a comparable ratio of tdT+ dCAMs in both Ccr2CreERT2R26tdT and Cxcr4CreERT2R26tdT mice, suggesting that repopulated dCAMs are mostly derived from monocytes (Extended Data Fig. 5k, l). In line with our observations for sdCAMs, there was no reduction of monocyte-derived dCAMs following injection of VLA-4-neutralizing antibodies or after transplantation of CD45.2+Itgb2−/− bone marrow. In contrast to sdCAMs, injection of anti-VCAM-1 and anti-ICAM-1 antibodies did not affect the infiltration of monocyte-derived dCAMs (Extended Data Fig. 5m–o). Therefore, dCAMs repopulate rapidly through an enhancement of physiological monocyte engraftment.
Long-term transcriptional alterations in repopulated sdCAMs
To characterize comprehensively the replenished microglia and sdCAMs at the molecular level, we performed scRNA-seq at 8 weeks post-depletion (Fig. 5a and Supplementary Fig. 4a). A total of 21,402 individual immune cells were annotated according to their cell-type-specific markers3,33 (Fig. 5b and Extended Data Fig. 6a). Microglia were virtually unaffected by the treatment, with only three genes slightly regulated, whereas 648 differentially expressed genes (DEGs) were identified for repopulated sdCAMs (Fig. 5c). Further subclustering resulted in five microglia clusters (Fig. 5d and Supplementary Fig. 4b–d). Microglia clusters 1–4 (MG1–MG4) had a typical homeostatic signature expressing Sparc, Hexb, P2ry12 and Siglech, whereas MG5 contained few cells with subtle induction of Spp1, Cst7 and Itgax (Extended Data Fig. 6b and Supplementary Fig. 4e, f). In contrast, sdCAMs split into four distinct clusters (sdCAM1–sdCAM4) (Fig. 5d and Supplementary Fig. 4b, d) with sdCAM1, 3 and 4 displaying high expression of Mrc1, Pf4 and Lyve1, whereas sdCAM2 instead expressed higher levels of MHC class II-related genes such as H2-Aa and Cd74 (Extended Data Fig. 6c and Supplementary Fig. 4g, h). Of note, all sdCAM and microglia clusters were found in control and repopulated mice; however, we found significant alterations in the composition of repopulated sdCAMs, but not in microglia. After repopulation, sdCAM2 made up 40.07% of sdCAMs (21.18% in control), accompanied by a significant decrease of sdCAM1 to 32.83% (39.90% in control) and sdCAM3 to 21.33% (27.85% in control) (Fig. 5e). DEG analysis between the affected sdCAM clusters showed increased expression levels of MHC class II-related genes, Ccr2 and Fxyd5 in sdCAM2, whereas Lyve1, Folr2 and Cd163 were up in sdCAM1 + 3 (Fig. 5f and Extended Data Fig. 6d, e). In summary, microglia regained their initial transcriptional profile after repopulation, whereas sdCAM composition remained altered, shifting toward the immunologically primed sdCAM2 phenotype.
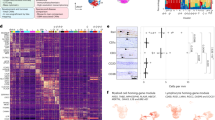
a, Experimental scheme. TAM, tamoxifen; CP, choroid plexus. n = 4 mice per treatment. b, Annotated UMAP of 21,402 individual cells. c, Reclustered UMAPs of 11,147 MG and 8,683 sdCAMs. d, Volcano plots comparing MG and sdCAMs from 8w after BLZ945 to CTL. Bonferroni-corrected P values. NS, not significant, P > 0.05. e, Cluster proportions for MG or sdCAMs at 8w after CTL or BLZ945 treatment. n = 4 mice per treatment, arithmetic mean across biological replicates, arcsine transformation with t-test and Benjamini–Hochberg false discovery rate (FDR) correction. f, Volcano plot comparing DEGs between sdCAM2 to sdCAM1 and sdCAM3 (c). Bonferroni-corrected P values, NS: P > 0.05. g,h, Experimental scheme. CPs were removed and CD206+tdT+ and CD206+tdT− sdCAMs were sorted for bulk RNA-seq (g). PCA (h). n = 4 mice per treatment. i, Volcano plot depicting DEGs between CD206+tdT+ sdCAMs and CD206+tdT− sdCAMs. Benjamini–Hochberg-adjusted P values. NS, P > 0.05. j, Genes found to be upregulated in CD206+tdT+ sdCAMs or CD206+tdT− sdCAMs in the bulk RNA-seq dataset (i) were plotted onto the scRNA-seq sdCAM dataset (c) as module scores for feature plots. k, Radar chart depicting relative expression levels of selected genes across sdCAM populations from bulk RNA-seq. Per gene, highest expression level is scaled to 1. l, Immunofluorescence based quantification of MHC class II expression in brains of Cxcr4CreERT2R26tdT mice. Arrowheads, white show MHC II−CD206+tdT− cell; blank show MHC II−CD206+ tdT+ cell; orange show MHC II+CD206+tdT+ cell. Symbols represent individual mice, n = 3 (CTL), n = 4 (BLZ945), mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. DAPI, 4,6-diamidino-2-phenylindole. m, Representative immunofluorescence image of MHC II expression in whole mount leptomeninges of Ccr2CreERT2R26tdT mice. Arrowheads, blank show MHC II−CD206+tdT+ cell; orange show MHC II+CD206+tdT+ cell. Representative of three mice.
Ontogeny drives phenotypic changes in repopulated sdCAMs
As scRNA-seq revealed long-term transcriptional changes only in sdCAMs, we next investigated how developmental origin contributes to sdCAM phenotypes after repopulation. We thus performed bulk RNA-seq of YS-derived CD206+tdT+ and HSC-derived CD206+tdT− sdCAMs from Mrc1CreERT2R26tdT mice 8w after depletion (Fig. 5g). Principal-component analysis (PCA) showed a close association of control tdT+ sdCAMs with re-emerged tdT+ sdCAMs, whereas renewed tdT− sdCAMs were clearly distinct (Fig. 5h). Accordingly, DEG analysis revealed only minor differences between tdT+ sdCAMs from repopulated and control mice, but major changes compared to tdT− sdCAMs (Fig. 5i and Extended Data Fig. 6f). tdT+ sdCAMs highly expressed the typical sdCAM markers Cd163, Lyve1 and C4b, whereas tdT− sdCAMs showed stronger expression of MHC class II-related genes as well as Fxyd5 and Ccr2 (Extended Data Fig. 6f). The differences closely resembled those identified between sdCAM1 + 3 and sdCAM2 in the scRNA-seq analysis (Fig. 5a–f). Gene Ontology (GO) term analysis revealed enrichment of a ‘chemotactic’ phenotype in tdT+ sdCAMs and a ‘leukocyte’ and ‘regulation of cell adhesion’ pattern in HSC-derived tdT− sdCAMs (Extended Data Fig. 6g). We then investigated whether sdCAM phenotypes found in our bulk RNA-seq dataset would align with sdCAM clusters detected by scRNA-seq (Fig. 5a–f). Indeed, the genes upregulated in tdT+ sdCAMs were highly and selectively expressed in sdCAM1 and sdCAM3, whereas the genes upregulated in tdT− sdCAMs were nearly solely found in sdCAM2 (Fig. 5j and Extended Data Fig. 6h). Assessment of the most consistent genes from scRNA-seq and bulk RNA-seq established a common sdCAM gene signature (Clec12a, Lyz2, Ms4a7 and Apoe) as well as unique signatures for YS-derived tdT+ and HSC-derived tdT− sdCAMs (Fig. 5k).
In Cxcr4CreERT2R26tdT fate mapping mice, HSC-derived CD206+tdT+ sdCAMs were found in the leptomeninges and perivascular space in repopulated animals. Consistent with the observed transcriptomic changes, they were frequently positive for MHC class II (Fig. 5l). In addition, replenished CD206+tdT+ lmMΦ coexpressed MHC class II in repopulated Ccr2CreERT2R26tdT mice (Fig. 5m). In summary, during repopulation, HSC-derived sdCAMs preferentially gain an MHC IIhi phenotype and thus shape the CNS border macrophage transcriptome long-lastingly.
Ontogeny and repopulation-associated chromatin patterns
As engrafted HSC-derived sdCAMs showed a distinct transcriptomic pattern compared to YS-derived sdCAMs, we wondered to what extent this could be explained by epigenetic differences. Therefore, we performed single-cell assay for transposase-accessible chromatin-sequencing (scATAC-seq) of dCAMs, sdCAMs, microglia and Ly6Chi blood monocytes from control Mrc1CreERT2R26tdT mice or 8w post-depletion. The tdT reporter thereby allowed to distinguish CAMs based on their ontogeny (Fig. 6a). Cell subtypes were defined by transferring annotations from our scRNA-seq data (Fig. 2b) and cross-verification with our sorting strategy (Fig. 6a, b). Analysis revealed specific chromatin-accessibility profiles for microglia, sdCAMs, dCAMs and classical Ly6Chi monocytes (Extended Data Fig. 7a, b). While microglia showed high peak enrichment at the Sall1 locus, all CAM populations showed high accessibility of Ms4a7. All dCAMs showed enrichment at the Cd83 locus and Ly6Chi monocytes showed an enrichment at the Ly6c2 locus. We validated the relevance of these differences in chromatin accessibility on RNA level (Extended Data Fig. 7a). In summary, these data provide a framework for the identification of CNS macrophage subtypes at the chromatin level.
a, Experimental scheme. Mrc1CreERT2R26tdT mice were injected with TAM and either treated with BLZ945 or CTL. CPs were removed and MG, CD206+tdT+ and CD206+tdT− sdCAMs, CD206+tdT+ and CD206+tdT− dCAMs as well as Ly6Chi blood monocytes were sorted for scATAC-seq. Cells pooled from n = 3 (CTL) and n = 4 (BLZ945) mice. b, UMAP based on called peaks of 10,224 individual nuclei with cell type annotation. c, Bar graph depicting all significant differentially accessible peaks between BLZ945 and CTL for each cell type. d, Volcano plots comparing accessible peaks in MG and sdCAMs from 8w after BLZ945 to CTL. Bonferroni-corrected P values. NS, P > 0.05. e, Feature plots depicting module scores derived from selected differential gene activities between hom_ys_sdCAM and hsc_sdCAM. f, Coverage plots of Colec12 and H2-Aa loci depicting the aggregated ATAC reads for each cell subtype. Differentially accessible peaks are highlighted in gray. g, Venn diagram showing overlap of differentially accessible peaks in the BLZ945 compared to CTL condition for the depicted cell types. h,i, Feature plot displaying a repopulation module score calculated from differential gene activities that are commonly and significantly enriched in macrophages from BLZ945-treated animals compared to CTL (i). Coverage plot of the Zfp932 locus depicting the aggregated ATAC reads for each cell cluster. Differentially accessible peak is highlighted in gray.
As all macrophage populations displayed differences in chromatin accessibility following repopulation, we investigated the effects of treatment and ontogeny by comparing repopulated cells to control cells within each population (Extended Data Fig. 7d). Blood monocytes showed only four differentially accessible loci. In contrast, sdCAMs displayed 3,215 alterations in chromatin accessibility, about twofold more than found for microglia or dCAMs (Fig. 6c, d and Extended Data Fig. 7e). Both YS- and HSC-derived sdCAMs exhibited high accessibility at genes that were linked to their origins by RNA-seq (Figs. 6e and 5f, i). Colec12 accessibility was increased in homeostatic and repopulated YS-derived sdCAMs, whereas H2-Aa accessibility was heightened in HSC-derived sdCAMs (Fig. 6f). Of note, some homeostatic dCAMs were covered by the YS-derived CAM module, whereas the majority of the population was covered by the HSC-derived CAM module (Fig. 6b, e). Next, we performed motif analysis to infer transcription factor activity in YS- versus HSC-derived sdCAMs. It revealed significant enrichment of Arid3a, Hic1 and NFATC2 motifs in YS-derived sdCAMs. Consistent with their recent emergence from monocytic progenitors, HSC-derived sdCAMs were enriched for the RUNX1 motif, a transcription factor known to interact with other lineage-defining factors during monocyte differentiation43 (Extended Data Fig. 7f, g).
We next compared peaks with higher accessibility in repopulated macrophages between CNS macrophage populations (Fig. 6g). Hereby, we identified a conserved repopulation-associated signature (Fig. 6h), including highly enriched accessibility to Zfp932, a zinc finger protein linked to repression of the differentiation associated hedgehog signaling pathway44 (Fig. 6d,i and Extended Data Fig. 7e). Taken together, all macrophages underwent chromatin remodeling following repopulation, with the most pronounced changes observed in sdCAMs, reflecting a switch in ontogeny during repopulation.
Ontogeny impacts transcriptional response to endotoxemia
After repopulation, monocyte engraftment into the sdCAM compartment generates an immunoreactive transcriptomic profile and additional chromatin-accessibility changes at the Tlr4 locus in repopulated CAMs (Fig. 6d and Extended Data Fig. 7e). We therefore investigated cellular responses to systemic lipopolysaccharide (LPS) in CNS macrophage populations with distinct ontogeny. We exposed repopulated Ccr2CreERT2R26tdT mice to LPS and profiled microglia, tdT+ and tdT− sdCAMs as well as tdT+ and tdT− dCAMs by RNA-seq 12 h after treatment (Fig. 7a, Extended Data Fig. 8a and Supplementary Fig. 5a, b). PCA and analysis of the most variable genes revealed clustering according to cell types and treatment paradigms (Fig. 7b, c and Extended Data Fig. 8b, c). Known LPS-inducible genes, including Saa3, Il1rn, Spp1, Lcn2, Mt1, Ccl5 or Ccl13, were commonly found across all macrophage populations (Extended Data Fig. 8d–f). In line with our preceding results, the GO terms ‘leukocyte differentiation’ and ‘cell adhesion’ were enriched in HSC-derived tdT+ sdCAMs, whereas tdT− sdCAMs showed enrichment for gene sets related to ‘RNA generation and splicing’ (Extended Data Fig. 8d). Comparison of DEGs across macrophage populations revealed partial overlap of induced genes (n = 211), including Saa3, Lcn2, Ccl5 and Cxcl13 (Supplementary Fig. 5c–e), alongside population-specific responses (Fig. 7d, e).
a, Experimental scheme: Following BLZ945 treatment and tamoxifen injections, Ccr2CreERT2R26tdT mice were intraperitoneally (i.p.) injected with PBS or LPS and MG and sdCAMs were sorted 12 h later. n = 3 mice per condition. b, PCA of MG, CD206+tdT− sdCAMs and CD206+tdT+ sdCAMs isolated from BLZ945-treated PBS or LPS injected mice. c, Heatmap of the top 100 most variable genes. Scale bar indicates z-score normalized expression. d, Venn diagram depicting the overlapping and distinct number of upregulated DEGs, defined as Benjamini–Hochberg-adjusted P < 0.05 and |log2 fold change| > 1. e, Linear scaled radar chart depicting mean relative expression levels of selected genes within the individual CNS macrophage populations following LPS stimulation. f, Volcano plot showing DEGs between tdT+ sdCAMs and tdT− sdCAMs after LPS stimulation (left). Benjamini–Hochberg-adjusted P values. NS, P > 0.05. The top five GO terms for each population are shown (right). g, Setup of the experiment is displayed. MG and sdCAMs were sorted from either WT or Ccr2−/− mice that were either kept untreated (∅) or treated with CTL or BLZ945 followed by i.p. injection with PBS or LPS 12 h before termination. n = 3 mice per condition. h, PCA of MG (squares) and sdCAMs (circles) isolated from PBS or LPS injected WT and Ccr2−/− mice. i, Bar plots comparing the number of DEGs, defined as Benjamini–Hochberg-adjusted P < 0.05 and |log2 fold change| > 1, for BLZ945 versus CTL between WT or Ccr2−/− mice for MG or sdCAMs. Chi-squared test of independence for DEGs per genotype out of expressed genes.
Focusing on sdCAM responses, tdT− and tdT+ sdCAMs showed clear separation following LPS stimulation (Extended Data Fig. 8g). tdT+ sdCAMs were enriched for Il10ra, Irf4 and Hdac11, resulting in the GO terms ‘proliferation of immune cell subsets’ and ‘wound healing’. In contrast, tdT− sdCAMs expressed numerous inflammatory and activation-related genes such as Aif1, Wfdc17, Saa3, Atox1, Mt1/2, Slirp, Sem1 or Ubl5, indicating ‘major cellular activation’ and ‘cell stress’ (Fig. 7f). Notably, sdCAM subsets diverged while dCAMs converged upon LPS stimulation (Extended Data Fig. 8b, g). This effect was also apparent when comparing common LPS-induced genes, which revealed no expression difference between tdT+ and tdT− dCAMs (Extended Data Fig. 8h, i).
To determine whether HSC-derived sdCAMs alter overall sdCAM LPS responses, we applied the same model in either WT or in Ccr2-deficient (Ccr2−/−) mice, which lack circulating monocytes impairing the establishment of repopulated HSC-derived sdCAMs (Fig. 7g). Microglia and sdCAM samples segregated after LPS exposure, independent of the treatment group and genotype (Fig. 7h). The top 250 most variable genes revealed common as well as unique modules of regulated genes between microglia and sdCAMs (Extended Data Fig. 9a). Msr1, Saa3, Ccl5, Il1rn, Il15ra, Spp1, Il1a/b and Tnfaip3 were found among the commonly upregulated genes across CNS macrophage moieties. While Tnf and Il12b were increased in microglia, Saa3, Lcn2, Cxcl13 and Ccl5 were more strongly induced in sdCAMs (Extended Data Fig. 9a and Supplementary Fig. 5f–i). Independent of the genotype, repopulated microglia showed an almost identical LPS response compared to controls, indicating that their reaction was not altered by depletion and repopulation (Fig. 7i and Extended Data Fig. 9b). In contrast, the LPS response of sdCAMs differed between repopulated and control WT animals, with those differences largely disappearing between repopulated and control Ccr2−/− animals (Fig. 7i and Extended Data Fig. 9b).
Collectively, these results reveal compartment- and origin-specific signatures of CNS macrophages upon systemic immunological challenge. Monocyte engraftment into the sdCAM compartment shapes the functional response of CNS border macrophages to endotoxin, while leaving microglia unaffected in phenotype and immune responsiveness.
Selective sdCAM replacement shapes ischemic stroke outcome
Recent evidence highlights sdCAMs as key modulators of neuroinflammation and tissue repair after stroke45,46,47, yet the impact of repopulation and replacement on these functions remains unclear. To further validate potential differences between YS- and HSC-derived sdCAMs, we examined their roles in cerebral ischemia, the second-leading cause of death and primary driver of neurological disabilities in humans.
Thus, we applied a model of thromboembolic stroke47,48 in WT mice after repopulation (Fig. 8a). Notably, repopulated mice performed worse at 1 day post stroke (+1 d) and +5 d in two independent neurological function paradigms compared to control animals (Fig. 8b, c). No differences in lesion volume, angiographic score, hemorrhage score, histological tissue examination or IBA1 immunoreactivity in the CNS parenchyma were apparent at +1 d (Extended Data Fig. 10a–c and Supplementary Fig. 6a–c). Moreover, the lesion sites were not infiltrated by neutrophils, T cells or B cells at +1 d (Supplementary Fig. 6d).
a, Experimental scheme. IHC, immunohistochemistry; i.v., intravenous; (m)MRI, (molecular) magnetic resonance imaging; MPIO, Superparamagnetic iron oxide microparticles. b,c, Quantifications of strength deficit (b) and corridor test (c). Symbols represent individual mice, n = 10 (CTL), n = 8 (BLZ945). Mean ± s.e.m. with unpaired two-tailed t-test (b). Median ± interquartile range with two-sided Brunner–Munzel test with Benjamini–Hochberg FDR adjustment (c). d, DEGs between ipsilateral and contralateral tissue slices. n = 4 biological replicates per group. Benjamini–Hochberg-adjusted P values. NS, P > 0.05. e,f, Overlap (e) and number of stroke response genes with higher mean expression in either CTL or BLZ945 (f). Chi-squared test of goodness of fit. g, Expression levels of selected stroke response genes. n = 4 biological replicates per group. Ordinary one-way ANOVA with Tukey’s honest significant difference (HSD) test. h, Experimental scheme. n = 5 mice (CTL), n = 5 biological replicates (BLZ945, pooled from two mice each). i,j, Number of stroke response genes with higher mean expression in MG and sdCAMs from no stroke (∅) or stroke mice (i) and overlap of stroke response genes (j). Benjamini–Hochberg-adjusted P values, P < 0.05, log2fold change > 0.59. k, DEGs between tdT+ sdCAMs from stroke versus ∅ mice overlayed with DEGs between tdT− sdCAMs from stroke versus ∅ mice. Benjamini–Hochberg-adjusted P values. l, Selection of significant stroke response genes. Symbols represent individual biological replicates, n = 3 (tdT+ sdCAMs BLZ945), n = 5 (all other groups). Ordinary one-way ANOVA with Tukey’s HSD test. m,n, Representative mMRI of anti-VCAM-1 (m) or anti-P-selectin (n) MPIOs and quantification. Symbols represent individual mice, n = 10 (CTL), n = 8 (BLZ945), mean ± s.e.m. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons P values. o, Correlation between P-selectin mMRI signal and grip strength deficit. Confidence and prediction intervals are shown. PS, Spearman correlation coefficient.
To explore the underlying mechanism of the neurological impairment, we investigated the transcriptional reaction to stroke at +1 d in whole CNS tissue. Immune response genes were strongly induced in the ipsilateral hemisphere, accounting for most upregulated DEGs (Extended Data Fig. 10d). We identified 115 upregulated DEGs in controls between ipsi- and contralateral hemispheres and 274 upregulated DEGs in repopulated mice (Fig. 8d, e). Of note, 266 of the 279 total upregulated DEGs showed higher mean expression in repopulated mice, including genes involved in macrophage activation, inflammation, reactive oxygen species (ROS) production and blood–brain barrier or extracellular matrix modification (Fig. 8f, g and Extended Data Fig. 10e).
Similar to the whole-tissue analysis, examination of transcriptional changes in sdCAMs at +1 d revealed a substantial induction of genes involved in immune responses (Extended Data Fig. 10f). Comparison to the contralateral hemispheres showed pronounced differences in the stroke response. We found 425 upregulated DEGs in sdCAMs from repopulated mice and 225 upregulated DEGs in control mice, of which 74 DEGs overlapped (Extended Data Fig. 10g, h). GO term analysis suggested a strong shift toward a proliferative and phagocytic phenotype in repopulated sdCAMs (Extended Data Fig. 10i). Gene-level analysis further supported the notion and suggested an increase in activation markers and ROS production (Extended Data Fig. 10j, k).
To disentangle treatment- and origin-dependent responses, we induced strokes in Ccr2CreERT2R26tdT mice (Fig. 8h). Immunofluorescence revealed pronounced engraftment of HSC-derived tdT+ lmMΦ and pvMΦ, both in the cortical lesion as well as in the corresponding contralateral region in BLZ945-treated mice (Supplementary Fig. 6e). RNA-seq of tdT+ sdCAMs, tdT− sdCAMs and microglia revealed upregulation of previously identified stroke response genes and downregulation of homeostatic markers (Extended Data Fig. 10l) compared to nonstroke-induced repopulated controls (Fig. 7a). Consistent with our previous results (Fig. 8f), repopulated sdCAM populations showed the highest expression of most stroke response genes, with tdT+ sdCAMs exceeding their tdT− counterparts (Fig. 8i). The sdCAM populations from control and BLZ945-treated mice showed both common as well as subset-specific responses (Fig. 8j), with each gene set being associated with distinct GO terms (Extended Data Fig. 10m). Overlaying the stroke reactions of repopulated tdT+ and tdT− sdCAMs revealed genes specifically regulated in tdT+ sdCAMs (Fig. 8k). In contrast, microglia showed no such discrepancy. DEG count in control microglia even slightly outnumbered that of repopulated microglia (Extended Data Fig. 10n). Supporting our previous analysis, we found increased expression of macrophage activation markers and chemokines, as well as tdT (Gt(ROSA)26Sor), in tdT+ sdCAMs (Fig. 8l and Extended Data Fig. 10o).
Finally, we performed molecular magnetic resonance imaging42,47 to determine whether the elevated cytokine and ROS production and macrophage activation translated into enhanced endothelial activation. While VCAM-1 signal intensity did not differ (Fig. 8m), the signal intensity of P-selectin, known to be upregulated under highly inflammatory conditions42, was significantly stronger in the ipsilateral hemisphere of repopulated mice and positively correlated with grip strength deficit (Fig. 8n, o).
Collectively, these results demonstrate that selective replacement of sdCAMs by HSC-derived macrophages influences immune response, cytokine production and functional outcome in a thrombin model of ischemic stroke.
Discussion
In this study, we demonstrate that repopulated sdCAMs partially derive from transiently infiltrating but permanently engrafting HSC-derived CCR2hiLy6Chi monocytes. The resulting changes in the overall sdCAM transcriptome led to an altered reaction of sdCAMs to peripheral inflammatory stimuli and worsened the outcome following ischemic stroke.
Earlier studies had worked toward a method to efficiently and permanently replace CNS macrophages, microglia and sdCAMs, by HSC-derived cells. This was achieved through a combination of whole body irradiation (WBI) and bone-marrow transplantation (BMT) followed by CSF-1R-mediated depletion of CNS macrophages24. Other reports claimed to selectively replace sdCAMs with HSC-derived cells, at least temporarily, using WBI and BMT alone26,27; however, WBI has been shown to artificially prime the microglia niche for subsequent myeloid cell engraftment23. Deploying depletion and repopulation, we herein present a broadly applicable niche-specific permanent replacement strategy for directed and selective monocytic engraftment into CNS border niches.
While sdCAMs were strongly affected by their depletion and repopulation, we did not detect substantial changes in subdural or dural vascular niche cell types by scRNA-seq analysis. The predicted transient changes in the interactions between niche cells and macrophages, including increased CSF-1R signaling together with production of monocyte-attractant chemokines in repopulating macrophages, probably reflect the effort to restore homeostatic macrophage density within the CNS niches. These changes align with the infiltration of circulating monocytes. In accordance with known mechanisms of extravasation, the monocytic engraftment depended on integrin-mediated adhesion to ICAM-1 and VCAM-1, which were upregulated on vascular endothelial cells following macrophage depletion. Increased Tnf production by repopulating CNS macrophages might contribute to this vascular activation. Of note, microglia and sdCAMs remained separate during repopulation, with neither leaving its developmentally established niche.
Single-cell profiling revealed repopulated sdCAMs fall into the range of homeostatic YS-derived sdCAM subsets, but preferentially adopt an MHC class IIhi phenotype reminiscent of a previously described sdCAM state5,49,50,51. Bulk RNA-seq furthermore enabled us to define pan sdCAM markers (Mrc1, Clec12a, Lyz2, Ms4a7 and Apoe) and unique signatures of HSC-derived (MHC class II-related genes, Ccr2 and Fxyd5) and YS-derived (Lyve1, Cd163 and Folr2) sdCAMs. This proposed set of markers allows the capture of sdCAM subtypes with implications for ontogeny. Chromatin accessibility was altered in all repopulated CNS macrophage populations; however, those alterations associated with proliferation or differentiation in response to depletion do not seem to be the main driver of the transcriptional changes seen in sdCAMs. Instead, ontogeny-related epigenetic imprinting remains the factor deciding the phenotype of sdCAMs.
While microglia exhibit a time-dependent transcriptional response to LPS52,53, sdCAM gene expression changes have only been studied in a Lyve1-expressing subset54. We therefore comprehensively analyzed overall and origin-specific responses of sdCAMs, dCAMs and microglia after repopulation. While many genes were commonly upregulated, we also identified cell-type-specific patterns. Notably, microglia and dCAMs remained unaffected by repopulation, whereas sdCAM responses were markedly altered. Experiments using Ccr2−/− animals strongly suggested that these differences were mainly resulting from the engraftment of HSC-derived sdCAMs. It should be noted that due to the systemic route of LPS delivery, effects mediated by peripheral immune reactions cannot be fully ruled out as a confounding variable.
sdCAMs have been implicated in many CNS diseases, such as cerebral aneurysm formation55, ischemic stroke45,47,51 or neurodegenerative disorders26,27,56,57. Ischemic stroke induces a neuroinflammatory reaction that is thought to strongly contribute to the resulting neurological disabilities58,59. Accordingly, the increased expression levels of many cytokines, chemokines such as Cxcl10, and macrophage activation markers could explain the greater neurological impairment observed in repopulated mice. Notably, CXCL10 has been shown to directly impair synaptic plasticity via neuronal CXCR3, inducing sickness behavior in mice60. sdCAMs react rapidly to cerebral ischemia and switching their gene expression profile to an activated phenotype46,51,61. This immunological cascade can shape the outcome in either direction, worsening neurological deficits or dampening damage and aiding in resolution47,62. Age-related changes in sdCAM phenotype were recently shown to counteract increased ischemic stroke severity in aging mice47. Absence of sdCAMs has also been shown to increase stroke severity45. Our data further strengthen this theory by pointing to a protective role for YS-derived sdCAMs in stroke. Of note, CAMs are replaced long-lastingly by HSC-derived cells in models of traumatic brain injury and infection30. Consequently, we propose that such HSC-derived sdCAMs could be important players in case of a secondary insult.
In summary, the dual origin of replenished sdCAMs based on a single short-term depletion paradigm renders sdCAMs uniquely accessible for HSC-linked replacement strategies in CNS diseases.
Methods
Mice
All animal experiments were performed in compliance with respective national and federal regulations and approved by animal welfare authorities (Regierungspräsidium Freiburg, French Ministry of Higher Education and Research). C57BL/6JCrl (Charles River) served as WT mice. Ccr2-KO mice were from The Jackson Laboratory (B6.129S4-Ccr2tm1Ifc/J, Jax #004999). Cx3cr1CreERT2 (ref.63), Cxcr4CreERT2 (ref. 39), HexbCreERT2 (ref. 18), Mrc1CreERT2 (ref. 12) and Ccr2CreERT2 (ref. 24) (Jax #035229) lines were crossed with R26tdT (ref. 64) (Jax #007914) or R26YFP (ref. 65) (Jax #006148) reporter lines. Mice were bred under specific-pathogen-free conditions and housed under a regular light–dark cycle at 20–22 °C and 50–60% humidity. Both sexes were included in each group. Age of first gavage was 10 ± 3 weeks.
Tamoxifen treatment
TAM in corn oil (both Sigma-Aldrich) was injected i.p. using Cre-driver line-adjusted protocols: 1 mg TAM per 100 μl oil for 5 days (Cxcr4CreERT2R26tdT), 4 mg 200 μl−1 twice 48 h apart (Cx3cr1CreERT2R26tdT, Mrc1CreERT2R26tdT) in 5–9-week-old animals, 0.4 mg 20 µl−1 on P7 and P9 (HexbCreERT2R26yfp) or 2 mg 200 μl−1 three times 48 h apart (Ccr2CreERT2R26tdT).
BLZ treatment
BLZ945-HCl (Novartis) at 27.28 mg ml−1 in aqueous 20% hydroxypropyl-β-cyclodextrin (CTL; Sigma-Aldrich) solution was applied through oral gavage (200 mg kg−1 BW) daily for 7 days.
Antibodies
Information about all antibodies used throughout the study can be found in Supplementary Table 1.
Antibody injections
For adhesion blockade in vivo, antibodies or isotype controls were mixed in InVivoPure, pH 7.0 Dilution Buffer (BioXCell) injected i.p. (200 µg per antibody in 100 µl, every other day for seven injections) starting on the last day of gavage.
Bone-marrow transplantations
Recipient mice were anesthetized with 20 µl of a 1:1 medetomidin (Domitor, Orion Pharma)/ketamine (Ketamin 10%, Serumwerk) mixture, positioned under a ~1-cm thick lead shield covering the head, and irradiated with 11 Gy (X-ray irradiator, Rad Source RS2000). After 24 h, femurs and tibias from sex- and age-matched donor mice (1:2 donor:recipient ratio) were flushed with ice-cold PBS, filtered (100 µm) and centrifuged (4 °C, 300g, 5 min). Red blood cells were lysed (1 ml eBioscience RBC lysis buffer, Thermo Fisher), washed and resuspended at 1 × 10⁵ cells per µl in PBS, filtered (100 µm), and ~15 × 106 cells were injected intravenously into recipients. Mice were housed in IVC cages and received neomycin-sulfate drinking water (1.1 g l−1; Sigma-Aldrich) for 14 days. Blood chimerism was assessed 4 weeks post-transplantation.
LPS injections
Mice were i.p. injected with LPS (Sigma-Aldrich, L2880) in PBS at 2.5 mg kg−1 bodyweight or PBS alone 12 h before killing.
Thromboembolic stroke model
Purified mouse α-thrombin was injected into the middle-cerebral artery (MCA) to induce occlusion as previously described48. In brief, mice were anesthetized, fixed, craniotomized, the dura was removed and a glass micropipette was inserted into the MCA to inject 1 μl thrombin (2 IU μl−1 for WT). The pipette was withdrawn after 10 min for thrombus stabilization. Cerebral blood flow was monitored by laser Doppler up to 20 min after occlusion. Lesion volume was assessed by MRI at +1 d. Experiments followed STAIR recommendations. Group sizes were based on previous studies47,48. RNA-seq experiments were performed at two independent centers. Animals were excluded if they died perioperatively or had infarcts <5 mm³ (2 of 34 mice).
MRI procedure
MRI was performed as previously described48. Mice were anesthetized and imaging was performed on a 7T Pharmascan scanner (Paravision v.6.0.1). T2-weighted multislice multiecho images (TE/TR 33/2,500 ms) were used to quantify lesion volumes with ImageJ. T2*-weighted sequences assessed hemorrhage (0–2 scale) and two-dimensional time-of-flight angiography (TE/TR 12/7 ms) evaluated MCA recanalization using a thrombolysis in cerebral infarction-like scoring system (0–3).
Molecular magnetic resonance imaging
Molecular MRI was performed as previously described42,47. MPIOs were conjugated to anti-P-selectin or anti-VCAM-1 antibodies. Conjugation was verified by intrastriatal LPS injection followed by 3D T2*-weighted imaging. MRI was performed immediately after intravenous injection of conjugated MPIOs (200 μl, 2 mg Fe per kg) at 70-μm isotropic resolution (TE/TR 13.2/200 ms; flip angle 21°). Images were displayed as minimum intensity projections of six slices, and signal voids were quantified in ImageJ using automated thresholding and expressed as the percentage of void area within the region of interest.
Corridor test
Sensorimotor performance was assessed using a corridor exploration task. A black PVC corridor (150 × 6 × 16 cm) with five objects (2 cm) on each side, spaced 18.5 cm apart, 1 cm above the floor was used. Object shape and color varied between test days. After 15 min habituation, mice explored the corridor freely for 1 min, and object explorations were recorded.
Grip strength test
Forepaw grip strength was measured using a BIOSEB grip strength meter. Mice were placed at a T-bar and gently pulled by the tail. Measurements were taken at baseline (day before MCAO), and at 24 h and 5 days post-MCAO. Each session consisted of five trials with 1-min rest intervals, and mean force was calculated. Mean strength deficit relative to baseline was calculated for each mouse.
Immunofluorescence
After transcardial perfusion with PBS, brains were fixed for 5–6 h in 4% PFA, put in 30% sucrose in PBS until saturation, embedded in Tissue-Tek O.C.T. (Sakura Finetek), frozen at −80 °C, sliced (Leica CM1900) at 20 µm (analysis) or 50 µm (representative images), rehydrated in PBS (5 min) and permeabilized for 1 h (5% bovine serum albumin and 0.5% Triton X-100 in PBS). Brain sections were processed free floating or on slides in upright racks. Primary antibodies were added (overnight, 4 °C) in blocking buffer. Secondary antibodies were applied (2 h). Coverslips were mounted with ProLong Diamond (Thermo Fisher Scientific) or with Mowiol.
Immunohistochemistry and H&E staining
Chromogenic IHC66 and hematoxylin and eosin (H&E) staining were conducted on 3-µm-thick FFPE sections. IHC used the labeled streptavidin–biotin technique. The slides were deparaffinized in xylene, subjected to heat-induced antigen retrieval in EnVision low pH buffer (40 min) and treated with 3% hydrogen peroxide (Carl Roth, 8070.1) for 10 min. Blocking solution (10% normal goat serum (SouthernBiotec, 0060-01), 1% Triton X-100 (Sigma, T8787-100ML) and TRIS buffer (EnVision Flex Wash Buffer, DAKO, K8000), at pH 7.2, 1 h) was applied. The primary antibody was incubated overnight. Slides were washed and treated with secondary antibody (45 min) and washed. Streptavidin-HRP (7105-05, SouthernBiotech, 1:1,000 dilution in Tris buffer, 45 min) was applied. Sections were washed, exposed to DAB solution (one drop of EnVision Flex DAB Chromogen per 1 ml of EnVision Flex Substrate Buffer) and counterstained (Gill’s hematoxylin solution (Sigma, 1051750500). For H&E staining, slides were deparaffinized, rehydrated through graded ethanol to water and stained with Gill’s hematoxylin. After rinsing, sections were differentiated in 0.3% acid alcohol (1% HCl in 70% ethanol, 5–10 s), rinsed and blued (0.1% sodium bicarbonate, 1 min). Eosin Y (0.5% in ethanol) staining was performed (1 min), followed by dehydration through ascending ethanol and clearing in xylene (2 × 5 min). Coverslips were mounted with Vitro-Clud (R. Langenbrinck, 04-0001).
Wholemounts
Leptomeninges were sliced off from fixed brains, cut into 3 × 3-mm pieces and processed as described under the immunofluorescence section.
Microscopy
Imaging was conducted with a conventional fluorescence microscope (Keyence BZX or Leica Thunder Imager, quantification) or with a TCS SP8 X confocal (Leica, LAS X 3.5.7.23225, representative images) using a ×20 0.75 NA objective (HC PL APO 20x/0.75 IMM CORR CS2).
Flow cytometry
Flow cytometry was conducted as described previously3. In brief, around 100 µl of blood were collected in PBS with 0,5 mol l−1 EDTA at pH 8.0 and lysed three times with Red Blood Cell Lysis Buffer (Thermo Fisher Scientific) before the staining procedure. Animals were then perfused transcardially with ice-cold PBS. MG and sdCAMs were isolated from whole brain for the experiments in Figs. 1 and 3, otherwise choroid plexuses were removed under a stereomicroscope. For digestion, brains were homogenized in HBSS containing 0.1 mg ml−1 DNase I (Roche) and 0.2 mg ml−1 Liberase (Roche) and put in a shaker at (37 °C, 1,000 rpm, 15 min). Cells were filtered, density centrifuged, cleaned of myelin, washed and used for the staining procedure. For dCAM data, cranial dura mater was removed from the cranial bone under a stereomicroscope and digested in 0.5 mg ml−1 Collagenase IV (Sigma-Aldrich) in PBS (30 min, 37 °C). Cell suspensions were mechanically dissociated, filtered, washed and used for the staining procedure. To analyze niche cells from different compartments, cranial dura, choroid plexus, leptomeninges and the cortical parenchyma were separated and enzymatically digested. Dura and the parenchyma were put into 0.2 mg ml−1 collagenase P (Sigma), 60 U ml−1 DNase I (Applichem), 0.3 U ml−1 Dispase I (Roche), 2% FCS and 20 mM HEPES in RPMI 1640 (all three Lonza) and digested (pH 7.2, 37 °C, 15 min, three cycles). For leptomeningeal dissociation, 0.2 mg ml−1 collagenase type 2, with two 5-min digestion steps was applied. Cells were then washed. From cortical samples, myelin was removed by density centrifugation. For the staining procedure, first, an Fc Block was applied (5 min, 4 °C) and samples were incubated with antibodies for 45 min (for blood 20 min, 4 °C). After washing, cells were sorted using an Aria III or analyzed using a LSRFortessa (Becton Dickinson). Dead cells were defined by labeling with Fixable Viability Dye (65-0866, eBioscience) or DAPI. Data were acquired with FACSDiva (Becton Dickinson). Post-acquisition analysis was performed using FlowJo v.10.10.
Single-cell RNA-sequencing
After isolating whole brain and removing choroid plexuses under a stereomicroscope, for Fig. 2, cerebral cortex, leptomeninges and dural immune and niche cells were sorted by gating DAPI− cells and then for CD11b+CD45+ (immune), CD31+ (endothelial), PDPN+ (fibroblasts) and marker-negative (other cell types) cells as described in the flow cytometry section and as seen in the gating strategies (Supplementary Fig. 1). For Fig. 5, only CD45+CD11b+Gr-1−CD11c− cells were sorted from brain devoid of choroid plexus. Afterwards, the 10x scRNA-seq platform using the Chromium controller with the Chromium Next GEM Single Cell 3’ kit v.3.1 (Fig. 5) or Next GEM Single Cell 3’ HT kit v.3.1 (Fig. 2) (10x Genomics) was applied. Amplification of complementary DNA and library preparation were conducted following the manufacturer’s instructions. Libraries were sequenced on a NextSeq1000 Sequencer (Illumina) appropriate for obtaining 20,000 reads per cell. The resulting fastq files were further processed using the Cell Ranger v.7.1.0 pipeline (10x Genomics) for demultiplexing, read alignment to the mouse genome (GRCm39 or mm10; Cre, tdTomato, WPRE-QY added as chromosomes) and gene count determination.
Doublet detection, quality control and analysis of the single-cell transcriptomic data
Transcriptomic data was analyzed in RStudio (build 764) with R programming language (v.4.4.0). The feature bc matrices were loaded with Seurat (v.5.0.3)67.
For Fig. 2, count matrices were filtered for low-quality cells (features 200–6,000, mitochondrial RNA < 5%) and doublets were removed using scDblFinder (v.1.16.0) after conversion with as.SingleCellExperiment68. Singlets were retained and merged into one Seurat object. Standard Seurat preprocessing was applied (NormalizeData, FindVariableFeatures (5,000), ScaleData, RunPCA, ElbowPlot, RunUMAP, FindNeighbors and FindClusters). Cells were manually inspected with CellSelector and cells expressing multiple lineage markers were removed12,18,29. Visualization used UMAP/DimPlot69. Batch correction was performed using Harmony70 (Split and IntegrateLayers), followed by the standard Seurat workflow using the Harmony reduction. The integrated object was converted to SingleCellExperiment and annotated with SingleR71. Major cell classes were subset (fibroblasts, endothelial, vascular smooth muscle cells, pericytes, immune and others) and subtypes manually annotated using FindAllMarkers with established literature3,22,33,34,35,36,37 and previous biological knowledge. CellChat72 was used to infer cell–cell interactions within the dura and leptomeninges/cortex. For macrophage-niche communication, CellChat objects were split into CTL, 5 d and 8w, top interaction partners were identified (excluding self-loops and niche–niche interactions) and interactions with homeostatic and repopulating macrophages were visualized.
For Fig. 5, processing was carried out as for Fig. 2, but cells were filtered for features 200–3,000 and mitochondrial RNA < 5% and Seurat preprocessing differed for FindVariableFeatures (2,000; excluding artificial, mitochondrial and ribosomal genes). A metadata column for tdTomato+WPRE-QY expression was added. Batch correction was performed using CCA73 (SplitObject, FindIntegrationAnchors and IntegrateData), followed by the standard Seurat workflow. Two additional low-quality clusters were removed based on low nCountRNA/nFeatureRNA and low gene expression. The standard Seurat workflow was applied to the updated object. Cell types were annotated based on known cell-type-specific marker genes, shown in Fig. 6a.
CAM and MG subset analysis
CAMs and MG were subsetted, reintegrated by CCA, and processed using the standard Seurat workflow. The integrated data slot was used for downstream analysis. Clustering resolution was set to 0.25 for both. Heatmap genes were filtered for adjusted P < 0.05, log2fold change > 0.59, pct.1 > 0.2 and pct.1–pct.2 > 0.15, showing up to 30 genes per cluster. Volcano plots (BLZ945 versus CTL) were generated using FindMarkers (pct.min = 0.2). Cluster proportion changes were assessed using the propeller workflow (speckle)74. For CAMs, CAM2 was compared to CAM1 + 3 using FindMarkers (pct.min = 0.2). Overlapping DEGs (adj. P < 0.05, pct.1 > 0.2) between treatment and cluster comparisons were visualized using VennDiagram75; top ten genes up/down by log2fold change are shown. GO analysis was performed with EnrichGO (clusterProfiler) using significant cluster 2 versus 1 + 3 genes and nonzero genes as background, followed by simplify (cutoff of 0.7)76. YS-derived and HSC-derived gene modules were derived from bulk RNA-seq (adj. P < 0.05; log2fold change > 1 or <−1) and visualized in CAM subsets using AddModuleScore with FeaturePlot or DoHeatmap.
Bulk RNA-seq of CNS macrophages and stroke tissue
CAMs, sdCAMs, dCAMs and MG were isolated by FACS from dissociated mouse brains (processed as described in the flow cytometry section) and sorted into RNA Protect Cell Reagent (QIAGEN); total RNA was extracted using the PicoPure RNA Isolation kit. First-strand cDNA was generated from 150–600 pg RNA using the SMARTer Ultra Low Input RNA kit v.4, followed by long-distance-PCR amplification (12–14 cycles), bead purification, Nextera XT library preparation, equimolar pooling and 50-cycle single-read sequencing on Illumina HiSeq 1000; reads were aligned to Gencode M36/mm10 or GRcm39 with STAR v.2.7.11a and counted with FeatureCounts v.2.0.8. In the post-repopulation experiment (Fig. 5), CAMs from Mrc1CreERT2R26tdT mice were sorted (tdT⁺ and tdT⁻ CD206⁺). Lowly expressed genes (sum counts <10 in ≥75% of samples) were removed before performing DESeq2 differential expression, lfcShrink(apeglm), heatmaps of significant DEGs (Padj ≤ 0.05, |log2fold change | ≥ 1; n = 380), volcano plots and enrichGO analysis. For LPS stimulation, MG and (all or tdT⁺/tdT⁻) sdCAMs were isolated from Ccr2CreERT2R26tdT, WT or Ccr2−/− mice 12 h after LPS (2.5 mg kg−1) or PBS. dCAMs were assessed in a separate experiment. Lowly expressed genes (sum count < sample number/2) were filtered, ComBat-seq batch correction was applied to WT/Ccr2−/− datasets (batch = batch, group = treatment; covariates = genotype, cell type). To ensure the quality of RNA-seq analysis and focus on immune response-related gene expression, genes in the KEGG_ribosome pathway, KEGG_oxidative_phosphorylation pathway (msigdbr), biomaRt pseudogenes, predicted genes, novel or uncharacterized genes as well as biomaRt noncoding genes (lincRNA, miRNA, snRNA and snoRNA) and genes with substrings associated with such pathways (‘Mrp’, ‘Rpl’, ‘Rps’, ‘Sn’, ‘Sf’, ‘Gtf’, ‘Med’, ‘Rbm’, ‘Rik’, ‘Atp’, ‘Nduf’, ‘Tomm’, ‘Pol’, ‘Anap’, ‘Rna’, ‘Dna’, ‘Gm’, ‘Ddx’, ‘Smim’, ‘Bola’, ‘Timm’ and ‘Micos’) were removed. DESeq2 was used for differential expression (MG excluded for CAM-specific analyses), with PCA and heatmaps on variance-stabilized top variable genes, apeglm-shrunken volcano plots, enrichGO on DEGs (Padj ≤ 0.05) and ggradar plots of selected LPS-responsive genes. In the stroke experiment (Fig. 8), at 24 h post-MCAO, brains from WT or Ccr2CreERT2R26tdT mice were sectioned, choroid plexus removed, and CAMs sorted as Ly6C⁻Ly6G⁻CD11c⁻CD45⁺CD11b⁺CD206⁺ (tdT⁺/tdT⁻ in reporter mice) or MG as CD206⁻ counterparts (cells of two BLZ945 mice were pooled per replicate); whole-tissue RNA was extracted using RNeasy Micro kit with DNase and Bioanalyzer quality control. Lowly detected genes (sum count < sample number/2) were filtered, DESeq2 performed differential expression, PCA, apeglm-based volcano plots and GO analysis on upregulated genes (Padj ≤ 0.05) in ipsilateral hemisphere (CTL or BLZ945); additional visualizations included Venn diagrams of overlapping upregulated genes, bar plots of gene counts with higher stroke-induced expression, box plots of selected immune genes. Volcano plots for Ccr2CreERT2R26tdT data were generated by plotting DEGs detected in repopulated stroke MG versus CTL MG and overlaying DEGs detected in vehicle stroke MG versus CTL MG or by plotting DEGs detected in tdT+ stroke CAMs versus tdT+ CTL CAMs and overlaying DEGs detected in tdT− stroke CAMs versus tdT− CTL CAMs.
scATAC-seq
MG, sdCAMs, dCAMs and Ly6Chi monocytes were isolated from brains or blood of Mrc1CreERT2R26tdT mice 8 weeks after depletion as described under flow cytometry. Then, 3–4 biological replicates were pooled, MG and monocytes from the same condition were pooled. Cells were sorted and processed using the Chromium Single Cell ATAC-seq platform (10x Genomics). Nuclei were isolated following the 10x Genomics protocol (CG000366, Rev A) with minor adaptations. In brief, sorted cells were lysed in diluted lysis buffer, quenched with wash buffer, centrifuged, resuspended and filtered (10-µm strainer). Nuclei integrity was assessed by DAPI staining and counted using Trypan Blue on a Countess automated cell counter. Encapsulation and library preparation were performed using Chromium Next GEM Single Cell ATAC Reagent Kits v.2 (10x Genomics). Library quality was assessed by Qubit and Bioanalyzer. Raw reads were processed using Cell Ranger ATAC (v.2.2.0) with alignment to the mm10 genome and generation of single-cell accessibility matrices. Downstream analysis was performed in R using Signac (v.1.16). Feature-barcode matrices were loaded, and quality control was performed based on transcription start site enrichment and nCount_peaks. Samples were merged based on a common GRanges peak set. Standard Signac workflow (RunTFIDF, FindTopFeatures, RunSVD, RunUMAP, FindNeighbors and FindClusters) was applied. Gene activity scores were calculated. Cell type annotation was transferred from scRNA-seq data (Fig. 2; macrophages and monocytes from CTL and 8w samples) using TransferData and refined using predicted identities and sorting metadata. Peak calling was performed using CallPeaks (MACS2; group.by = cell type, combine.peaks = TRUE), followed by removal of nonstandard chromosomes, quantification (FeatureMatrix) and annotation (RegionStats). A standard Signac workflow and gene activity calculation were repeated on peak assay. The bar graph of differential accessibility includes increased and decreased peaks. Module contents were based on (sc)RNA-seq analysis and FindAllMarkers results. Motif enrichment was performed using JASPAR2020, and transcription factor activity was assessed using chromVAR. Visualizations were generated with SCpubR77, Signac, EnhancedVolcano78 and ggplot2 (ref. 79).
Analysis and experimental information
Analysis, surgeries and outcome assessments were conducted blinded to treatment conditions. Animals were randomly assigned to treatment groups, with a similar number of males and females per group. Sample sizes were not predetermined, but are similar to those reported previously3,12,47.
Histological analysis
All data points for histological analyses in Figs. 1 and 3 and associated Extended Data derive from manual quantification (ImageJ Fiji and Cell Counter plug-in) of ten regions of interest for cerebral cortex, at least six for cerebellum and at least four for olfactory tubercle per biological replicate. In brief, the laminin channel was activated and parenchyma area measurements and leptomeningeal length measurements were performed, the IBA1 channel was used to count the number of MG. Next, the CD206 channel was used to quantify the number of lmMΦ and pvMΦ. Last, the reporter channel was activated and tdT+ cells were quantified. For Fig. 4, images of whole-brain sagittal sections were semi-automatically analyzed using QuPath v.4.4 software (≥1,000 CAMs per biological replicate). For Fig. 5, whole-brain sagittal sections were manually counted using ImageJ Fiji (≥1,000 CAMs per biological replicate). For analysis of CD62P+, CD62E+ or ICAM-1+ vessels, Cell Profiler v.4.2.5 (Crop, Threshold, IdentifyPrimaryObjects, MaskObjects and MeasureImageAreaOccupied) was deployed.
Statistical analysis
Normality and homoscedasticity were generally assumed; equality of variances was assessed for all GraphPad-based analyses. A significance level of 5% was used (P < 0.05). Histological and flow cytometry data were analyzed in GraphPad Prism v.11 using unpaired two-tailed t-tests or one-way ANOVA with Šidák’s or Tukey’s post hoc tests. For Fig. 1f, cell density data were analyzed in R using biological replicate-level counts. Data were converted to long format and Welch’s two-sample t-tests were performed for each day (CTL versus BLZ), followed by Bonferroni correction across six comparisons. A three-parameter logistic growth model was fitted to mean depletion over time using nonlinear least squares:
where L is the asymptote, k is the growth rate and t₀ is the inflection point. Convergence failures were handled with tryCatch. Model fit was evaluated (R²) and time to 90% depletion (t₉₀) was derived analytically. Sequencing data were analyzed in R using methods described in the respective RNA-seq sections. Cluster composition in Figs. 2 and 5 was assessed with the propeller workflow (speckle), applying arcsine square-root transformation followed by a t-test (Fig. 5e, two conditions) or ANOVA (Extended Data Fig. 1c; three conditions) with Benjamini–Hochberg FDR correction. Corridor test discrete number statistics were performed with gstat and significance was tested with the Brunner–Munzel test. P-selectin MPIO signal and strength deficit correlation was evaluated using Spearman correlation and linear regression (lawstat). Box plots were analyzed by ANOVA with Tukey’s test. Discrete number graphs were analyzed by chi-squared testing.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Code availability
Code for statistical analyses is available at https://github.com/fliegaufm/Repopulation_of_CAMs.git.
References
Kierdorf, K., Masuda, T., Jordão, M. J. C. & Prinz, M. Macrophages at CNS interfaces: ontogeny and function in health and disease. Nat. Rev. Neurosci. 20, 547–562 (2019).
Mrdjen, D. et al. High-dimensional single-cell mapping of central nervous system immune cells reveals distinct myeloid subsets in health, aging, and disease. Immunity 48, 380–395.e6 (2018).
Amann, L. et al. Extrasinusoidal macrophages are a distinct subset of immunologically active dural macrophages. Sci. Immunol. 9, eadh1129 (2024).
Goldmann, T. et al. Origin, fate and dynamics of macrophages at central nervous system interfaces. Nat. Immunol. 17, 797–805 (2016).
Drieu, A. et al. Parenchymal border macrophages regulate the flow dynamics of the cerebrospinal fluid. Nature 611, 585–593 (2022).
Colonna, M. & Butovsky, O. Microglia function in the central nervous system during health and neurodegeneration. Annu. Rev. Immunol. 35, 441–468 (2017).
Prinz, M., Jung, S. & Priller, J. Microglia biology: one century of evolving concepts. Cell 179, 292–311 (2019).
Prinz, M., Masuda, T., Wheeler, M. A. & Quintana, F. J. Microglia and central nervous system–associated macrophages—from origin to disease modulation. Annu. Rev. Immunol. 39, 251–277 (2021).
Ginhoux, F. et al. Fate mapping analysis reveals that adult microglia derive from primitive macrophages. Science 330, 841–845 (2010).
Kierdorf, K. et al. Microglia emerge from erythromyeloid precursors via Pu.1- and Irf8-dependent pathways. Nat. Neurosci. 16, 273–280 (2013).
Schulz, C. et al. A lineage of myeloid cells independent of Myb and hematopoietic stem cells. Science 336, 86–90 (2012).
Masuda, T. et al. Specification of CNS macrophage subsets occurs postnatally in defined niches. Nature 604, 740–748 (2022).
Tay, T. L. et al. A new fate mapping system reveals context-dependent random or clonal expansion of microglia. Nat. Neurosci. 20, 793–803 (2017).
Füger, P. et al. Microglia turnover with aging and in an Alzheimer’s model via long-term in vivo single-cell imaging. Nat. Neurosci. 20, 1371–1376 (2017).
Ajami, B., Bennett, J. L., Krieger, C., Tetzlaff, W. & Rossi, F. M. V. Local self-renewal can sustain CNS microglia maintenance and function throughout adult life. Nat. Neurosci. 10, 1538–1543 (2007).
Elmore, M. R. P. et al. Colony-stimulating factor 1 receptor signaling is necessary for microglia viability, unmasking a microglia progenitor cell in the adult brain. Neuron 82, 380–397 (2014).
Hagemeyer, N. et al. Microglia contribute to normal myelinogenesis and to oligodendrocyte progenitor maintenance during adulthood. Acta Neuropathol 134, 441–458 (2017).
Masuda, T. et al. Novel Hexb-based tools for studying microglia in the CNS. Nat. Immunol. 21, 802–815 (2020).
Buttgereit, A. et al. Sall1 is a transcriptional regulator defining microglia identity and function. Nature Immunology 17, 1397–1406 (2016).
Parkhurst, C. N. et al. Microglia promote learning-dependent synapse formation through brain-derived neurotrophic factor. Cell 155, 1596–1609 (2013).
Bruttger, J. et al. Genetic cell ablation reveals clusters of local self-renewing microglia in the mammalian central nervous system. Immunity 43, 92–106 (2015).
Huang, Y. et al. Repopulated microglia are solely derived from the proliferation of residual microglia after acute depletion. Nat. Neurosci. 21, 530–540 (2018).
Mildner, A. et al. Microglia in the adult brain arise from Ly-6C hi CCR2+ monocytes only under defined host conditions. Nat. Neurosci. 10, 1544–1553 (2007).
Xu, Z. et al. Efficient strategies for microglia replacement in the central nervous system. Cell Rep. 32, 108041 (2020).
Gerganova, G., Riddell, A. & Miller, A. A. CNS border-associated macrophages in the homeostatic and ischaemic brain. Pharmacol. Ther. 240, 108220 (2022).
Anfray, A. et al. A cell-autonomous role for border-associated macrophages in ApoE4 neurovascular dysfunction and susceptibility to white matter injury. Nat. Neurosci. 27, 2138–2151 (2024).
Uekawa, K. et al. Border-associated macrophages promote cerebral amyloid angiopathy and cognitive impairment through vascular oxidative stress. Mol. Neurodeg. 18, 73 (2023).
Frosch, M. & Prinz, M. Niche-specific therapeutic targeting of myeloid cells in the central nervous system. Immunity 58, 1101–1119 (2025). S1074761325001359.
Van Hove, H. et al. A single-cell atlas of mouse brain macrophages reveals unique transcriptional identities shaped by ontogeny and tissue environment. Nat. Neurosci. 22, 1021–1035 (2019).
Wang, L. et al. CCR2+ monocytes replenish border-associated macrophages in the diseased mouse brain. Cell Rep. 43, 114120 (2024).
Askew, K. et al. Coupled proliferation and apoptosis maintain the rapid turnover of microglia in the adult brain. Cell Rep. 18, 391–405 (2017).
Amann, L., Masuda, T. & Prinz, M. Mechanisms of myeloid cell entry to the healthy and diseased central nervous system. Nat. Immunol. 24, 393–407 (2023).
Jordão, M. J. C. et al. Single-cell profiling identifies myeloid cell subsets with distinct fates during neuroinflammation. Science 363, eaat7554 (2019).
Vanlandewijck et al. A molecular atlas of cell types and zonation in the brain vasculature. Nature 554, 475–480 (2018).
Pietilä, R. et al. Molecular anatomy of adult mouse leptomeninges. Neuron 111, 3745–3764.e7 (2023).
Rustenhoven, J. et al. Functional characterization of the dural sinuses as a neuroimmune interface. Cell 184, 1000–1016.e27 (2021).
Smyth, L. C. D. et al. Identification of direct connections between the dura and the brain. Nature 627, 165–173 (2024).
Goldmann, T. et al. A new type of microglia gene targeting shows TAK1 to be pivotal in CNS autoimmune inflammation. Nat. Neurosci. 16, 1618–1626 (2013).
Werner, Y. et al. Cxcr4 distinguishes HSC-derived monocytes from microglia and reveals monocyte immune responses to experimental stroke. Nat. Neurosci. 23, 351–362 (2020).
Ydens, E. et al. Profiling peripheral nerve macrophages reveals two macrophage subsets with distinct localization, transcriptome and response to injury. Nat. Neurosci. 23, 676–689 (2020).
Mildner, A. et al. CCR2+Ly-6Chi monocytes are crucial for the effector phase of autoimmunity in the central nervous system. Brain 132, 2487–2500 (2009).
Gauberti, M., Fournier, A. P., Docagne, F., Vivien, D. & Martinez de Lizarrondo, S. Molecular magnetic resonance imaging of endothelial activation in the central nervous system. Theranostics 8, 1195–1212 (2018).
Hu, Z. et al. RUNX1 regulates corepressor interactions of PU.1. Blood 117, 6498–6508 (2011).
Huang, G. J., He, Z. & Ma, L. ZFP932 suppresses cellular Hedgehog response and Patched1 transcription. Vitam. Horm. 88, 309–332 (2012).
Pedragosa, J. et al. CNS-border associated macrophages respond to acute ischemic stroke attracting granulocytes and promoting vascular leakage. Acta Neuropathol. Commun. 6, 76 (2018).
Rajan, W. D. et al. Defining molecular identity and fates of CNS-border associated macrophages after ischemic stroke in rodents and humans. Neurobiol. Dis. 137, 104722 (2020).
Levard, D. et al. Central nervous system-associated macrophages modulate the immune response following stroke in aged mice. Nat. Neurosci. 27, 1721–1733 (2024).
Orset, C. et al. Mouse model of in situ thromboembolic stroke and reperfusion. Stroke 38, 2771–2778 (2007).
Dick, S. A. et al. Three tissue resident macrophage subsets coexist across organs with conserved origins and life cycles. Sci. Immunol. 7, eabf7777 (2022).
Brioschi, S. et al. A Cre-deleter specific for embryo-derived brain macrophages reveals distinct features of microglia and border macrophages. Immunity 56, 1027–1045.e8 (2023).
Garcia-Bonilla, L. et al. Analysis of brain and blood single-cell transcriptomics in acute and subacute phases after experimental stroke. Nat. Immunol. 25, 357–370 (2024).
Shemer, A. et al. Interleukin-10 Prevents pathological microglia hyperactivation following peripheral endotoxin challenge. Immunity 53, 1033–1049.e7 (2020).
Holtman, I. R. et al. Induction of a common microglia gene expression signature by aging and neurodegenerative conditions: a co-expression meta-analysis. Acta Neuropathol. Commun. 3, 31 (2015).
Kim, J.-S. et al. A binary Cre transgenic approach dissects microglia and CNS border-associated macrophages. Immunity 54, 176–190.e7 (2021).
Glavan, M. et al. CNS-associated macrophages contribute to intracerebral aneurysm pathophysiology. Acta Neuropathol. Commun. 12, 43 (2024).
Hawkes, C. A. & McLaurin, J. Selective targeting of perivascular macrophages for clearance of β-amyloid in cerebral amyloid angiopathy. Proc. Natl Acad. Sci. USA 106, 1261–1266 (2009).
Schonhoff, A. M. et al. Border-associated macrophages mediate the neuroinflammatory response in an alpha-synuclein model of Parkinson disease. Nat. Commun. 14, 3754 (2023).
Lambertsen, K. L., Biber, K. & Finsen, B. Inflammatory cytokines in experimental and human stroke. J. Cereb. Blood Flow Metab. 32, 1677–1698 (2012).
Iadecola, C. & Anrather, J. The immunology of stroke and dementia. Immunity 58, 18–39 (2025).
Blank, T. et al. Brain endothelial- and epithelial-specific interferon receptor chain 1 drives virus-induced sickness behavior and cognitive impairment. Immunity 44, 901–912 (2016).
Zheng, K. et al. Single-cell RNA-seq reveals the transcriptional landscape in ischemic stroke. J. Cereb. Blood Flow Metab. 42, 56–73 (2022).
Zeyen, T. et al. Microglial-specific depletion of TAK1 is neuroprotective in the acute phase after ischemic stroke. J. Mol. Med. 98, 833–847 (2020).
Yona, S. et al. Fate mapping reveals origins and dynamics of monocytes and tissue macrophages under homeostasis. Immunity 38, 79–91 (2013).
Madisen, L. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat. Neurosci. 13, 133–140 (2010).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Schwabenland, M. et al. Analyzing microglial phenotypes across neuropathologies: a practical guide. Acta Neuropathol. 142, 923–936 (2021).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Amezquita, R. A. et al. Orchestrating single-cell analysis with Bioconductor. Nat. Methods 17, 137–145 (2020).
McInnes, L., Healy, J., Saul, N. & Groβberger, L. UMAP: Uniform Manifold Approximation and Projection. J. Open Source Softw. 3, 861 (2018).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol 20, 163–172 (2019).
Jin, S., Plikus, M. V. & Nie, Q. CellChat for systematic analysis of cell–cell communication from single-cell transcriptomics. Nat. Protoc 20, 180–219 (2025).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol 36, 411–420 (2018).
Phipson, B. et al. propeller: testing for differences in cell type proportions in single cell data. Bioinformatics 38, 4720–4726 (2022).
Chen, H. & Boutros, P. C. VennDiagram: a package for the generation of highly-customizable Venn and Euler diagrams in R. BMC Bioinform. 12, 35 (2011).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Blanco-Carmona, E. Generating publication ready visualizations for single cell transcriptomics using SCpubr. Preprint at bioRxiv https://doi.org/10.1101/2022.02.28.482303 (2022).
Blighe, K., Rana, S. & Lewis M. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. R package version 1.28.2 https://bioconductor.org/packages/EnhancedVolcano (2025).
Wickham, H. Ggplot2. (Springer International Publishing, 2016).
Acknowledgements
We thank K. Gerber, S. Wundt, M. Ditter, J. Bodinek and U. Gopakumar for excellent technical assistance. We thank F. Elsayed for her establishment of immunofluorescence protocols. We thank C. Möhle and T. Stempfl for their contributions to the sequencing experiments. We thank D. Shimshek, F. Gasparini and I. Galimberti for the provision of BLZ945. We thank G. Monaco and C. Chhatbar for assistance with alignment of bulk RNA-seq data and for their suggestions for the scRNA-seq experiment. We thank the Lighthouse Core Facility, University of Freiburg, for their support with FACS-based cell sorting. The Lighthouse Core Facility is funded in part by the Medical Faculty, University of Freiburg (project numbers 2023/A2-Fol; 2021/B3-Fol), the DKTK, and the DFG (project number 450392965). We thank the Bioinformatics and Sequencing facilities at the MPI-IE; and in particular M. Navandar and W. Deboutte for demultiplexing and preliminary analysis of the 10x scATAC-seq dataset; the Imaging facility and Fly facility at the MPI-IE. Cell sorting for RNA-seq data related to stroke was performed on a MACSQuant Tyto Cell Sorter in the InnovaSEQ – Cell sorting and NGS @ Caen-Normandie Facility. The European project EquipInnovCaen2022-RNA-seq is funded by the European Union within the framework of the Operationnal Programme ERDF/ESF 2014-2020. The authors thank the Core Facility AMIRCF (DFG-RIsources no. RI_00052) for support in MR Imaging. A.A.D. is supported by the Hans A. Krebs Medical Scientist Program, Faculty of Medicine, University of Freiburg. M.S. is supported by the Berta-Ottenstein-Programme for Clinician Scientists, Faculty of Medicine, University of Freiburg, and the IMM-PACT-Programme for Clinician Scientists, Department of Medicine II, Medical Center—University of Freiburg and Faculty of Medicine, University of Freiburg, funded by the German Research Foundation (DFG; project number 413517907). M.D.B. is a member of the excellence cluster ImmunoSensation2 (EXC2151 project number 390873048). This work was funded by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany’s Excellence Strategy within the framework of the Munich Cluster for Systems Neurology (EXC 2145 SyNergy – ID 390857198). M.R. and L.A. are founded by the Procope Mobility Grant, supported by Campus France and the DAAD (project IDs 52948ZD or 57756192, respectively). L.A. and M.P. are supported by the German Research Foundation (TRR 167 project ID 259373024). M.P. is supported by the Faber Foundation, the Novo Nordisk Prize and the German Research Foundation (SFB 1160, SFB 1479, TRR 359, Gottfried-Wilhelm-Leibniz-Prize). This study was supported by the DFG under Germany’s Excellence Strategy (CIBSS – EXC-2189, project ID 390939984).
Funding
Open access funding provided by Albert-Ludwigs-Universität Freiburg im Breisgau.
Author information
Authors and Affiliations
Contributions
M. Fliegauf, L.A., D.L., F.C., M.R., J.C., M. Frosch, M.S. and R.S. conducted experiments and analyzed the data. S.K., A.A.D., A.D.G., R.S., M.D.B., M.K., D.B., W.R. and D.E. supported experiments. P.A. and A.O. provided reagents or materials. D.V., J.J.N. and N.I. contributed experimental suggestions and supported experiments. L.A. and M.P. supervised the project. M. Fliegauf, L.A. and M.P. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Peer review
Peer review information
Nature Immunology thanks Zsuzsanna Fabry, Guoping Peng and the other anonymous reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: L. A. Dempsey, in collaboration with the Nature Immunology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Single-cell RNA-sequencing of the CNS macrophage niche.
(a) Dot plot depicting canonical cell-type-defining markers. Color scale: Relative expression level. (b) Bar graph showing the proportion of each cluster within the different tissues (Related to Fig. 2). (c) Marimekko chart showing the proportion of each cluster within each treatment group. CTL: control; 5d: five days after BLZ945; 8w: eight weeks after BLZ945. n = 4 (Dura 8w and 5d), n = 3 (Dura CTL), n = 8 (Leptomeninges and Cortex 8w and 5 d), n = 7 (Leptomeninges and Cortex CTL), arcsine transformation followed by ordinary one-way ANOVA combined with Benjamini–Hochberg FDR correction. (d) Violin plots showing expression levels of selected macrophage-niche factors and chemokines across niche cells. (e) Bar plot showing aggregated strength of all interactions at the different analysis time points. (f, g) Reclustered UMAP of all macrophage subsets in leptomeninges and cortex (f). Feature plots of condition (g, left) and Nes (Nestin) expression levels (g, right) are shown. Scale bar shows z-score of minimum to maximum scaled gene expression. (h) Dot plot depicting gene expression of a selection of tumor necrosis factor (TNF) signaling components and chemokines predicted to mediate interactions between myeloid and niche cells. Color scale: Relative expression levels.
Extended Data Fig. 2 Non-cell-autonomous repopulation of sdCAMs but not MG throughout the brain.
(a) Experimental scheme: Tamoxifen (TAM)-induced Cx3cr1CreERT2R26tdT mice were treated with control (CTL) or BLZ945 and analyzed at different time points thereafter. (b) TdTomato (tdT) expression in Ly6Chi and Ly6Clo blood monocytes from Cx3cr1CreERT2R26tdT mice. Symbols represent individual mice, n = 3 (8w, 12w CTL), n = 4 (2w, 12w BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (c) Flow cytometry-based quantification of labeling in microglia (MG) or sdCAMs. Symbols represent individual mice, n = 3 (8w, 12w CTL), n = 4 (2w, 12w BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (d, e) Representative images from cerebral cortex, cerebellum and olfactory tubercle at 1d and 8w post CTL or BLZ945 treatment, respectively. Arrowheads: blank = tdT+ lmMΦ or pvMΦ; orange = tdT− lmMΦ or pvMΦ (d). Histological quantification of the proportion of tdT+ MG, lmMΦ and pvMΦ at different time points and in different brain regions (e). Symbols represent individual mice, n = 3 (8w, 12w CTL), n = 4 (2w, 12w BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values.
Extended Data Fig. 3 Repopulation dynamics of CNS macrophages throughout the brain.
(a) Experimental scheme. HSC: hematopoietic stem cell; MG: microglia, Mono: monocyte, TAM: Tamoxifen. (b-d) Flow cytometric quantification of reporter gene expression in Ly6Chi and Ly6Clo blood monocytes from HexbCreERT2R26YFP (b), Mrc1CreERT2R26tdT (c) and Cxcr4CreERT2R26tdT (d). Symbols represent individual mice, HexbCreERT2R26YFP: n = 3 (2w CTL), n = 4 (2w BLZ945, 8w, 26w), Mrc1CreERT2R26tdT: n = 3 (2w CTL, 26w CTL), n = 4 (2w BLZ945, 26w BLZ945), n = 6 (8w), Cxcr4CreERT2R26tdT: n = 3 (2w CTL, 8w CTL, 12w CTL, 26w CTL), n = 4 (8w BLZ945, 12w BLZ945), n = 5 (26w BLZ945), n = 6 (5d CTL), n = 8 (5d BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (e-f) Representative images (e) and quantifications (f) of labeled MG, leptomeningeal (lmMΦ) and perivascular (pvMΦ) macrophages. Symbols represent individual mice, HexbCreERT2R26YFP: n = 4; Mrc1CreERT2R26tdT: n = 4 (olfactory tubercle BLZ945), n = 5 (all other groups), mean ± SEM. Unpaired two-tailed t-test with Welch’s correction. Arrowheads: magenta = YFP+ MG; red = YPF− MG; blue = YFP− leptomeningeal (lmMΦ) and perivascular (pvMΦ) macrophages; open = tdT+ lmMΦ and pvMΦ; orange = tdT− lmMΦ and pvM. (g) Representative images from Cxcr4CreERT2R26tdT mice. Arrowheads: open = tdT− lmMΦ and pvMΦ; orange = tdT+ lmMΦ and pvMΦ. (h-k) Flow cytometric analysis. Representative histograms (h) and quantification of reporter gene expression in MG and sdCAMs from HexbCreERT2R26YFP (i), Mrc1CreERT2R26tdT (j) and Cxcr4CreERT2R26tdT (k). Symbols represent individual mice, HexbCreERT2R26YFP: n = 3 (2w CTL), n = 4 (2w BLZ945, 8w, 26w); Mrc1CreERT2R26tdT: n = 3 (2w CTL, 26w CTL), n = 4 (2w BLZ945, 26w BLZ945), n = 6 (8w); Cxcr4CreERT2R26tdT: n = 3 (5d CTL, 2w CTL, 8w CTL, 12w CTL, 26w CTL), n = 4 (8w BLZ945, 12w BLZ945), n = 5 (5d BLZ945, 26w BLZ945), n = 6 (2w BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (l) Representative images showing Ki-67 immunofluorescence in the cortex of Cxcr4CreERT2R26tdT mice 5d post CTL or BLZ945 treatment, respectively and quantification. Arrowheads: blue = Ki-67− lmMΦ and pvMΦ; red = Ki-67+ lmMΦ and pvMΦ. Symbols represent individual mice, n = 3 per group, mean ± SEM. Unpaired two-tailed t-test with Welch’s correction (MG) or ordinary one-way ANOVA with Šidák’s multiple comparison test, multiplicity adjusted p values (lmMΦ and pvMΦ).
Extended Data Fig. 4 Modulation of monocyte infiltration into CNS interfaces by integrins.
(a) Representative gating strategy for blood. (b) Experimental scheme and quantification of tdTomato (tdT) reporter expression. Symbols represent individual mice, n = 4 (17d–56d), n = 6 (11d BLZ945, 14d), n = 8 (11d CTL, 0d–7d), mean ± SEM. (c) Representative images of isolated choroid plexus (CP) from lateral ventricles (LV), 3rd ventricle (V) or 4th V. (d) Gene expression of selected cell adhesion molecules and integrins in leptomeninges and cortex (related to Fig. 1). Color scale: Relative expression levels. (e) Experimental scheme, representative images and quantification of CD62P+Laminin+ or CD62E+Laminin+ cerebral cortex vasculature. Symbols represent individual mice, n = 3 (CD62P), n = 8 (CD64E BLZ945), n = 9 (CD64E CTL), mean ± SEM, unpaired two-tailed t-test. Positive control for CD62P: Spinal cord of an experimental autoimmune encephalomyelitis (EAE) mouse at peak (Score 3). (f-i) Experimental scheme (left) and representative photography of a mouse 8 weeks after whole bone marrow (BMT) transplantation (right) (f). Flow cytometric quantification of CD45.2+ immune cells (g) and validation of successful ITGB2 knock-out in CD45.2+ Ly6Chi monocytes (h) in recipient blood at 21d. Flow cytometric quantification of CD45.2+ microglia (MG) and sdCAMs (i). Symbols represent individual mice, n = 3 (BLZ945), n = 4 (CTL), mice pooled for figure (h) Mean ± SEM, unpaired two-tailed t-test in (i). CP: Choroid plexus; FC: Flow cytometry.
Extended Data Fig. 5 Repopulation of macrophages in the dura mater.
(a) Experimental scheme. CTL: control; HSC: hematopoietic stem cell; Mono: monocyte; Tamoxifen: TAM (TAM); dCAMs: dural CAMs. (b) Representative gating strategy. (c) dCAM count per cranial dura mater quantified by flow cytometry. Symbols represent individual mice, n = 3 (1d, 2w, 8w, 12W, 26w all CTL), n = 4 (1d BLZ945, 8w BLZ945, 12w BLZ945), n = 5 (26w BLZ945), n = 6 (5d CTL, 2w BLZ945), n = 7 (5d BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (d) Flow cytometry-based quantification of tdTomato+ (tdT+) dCAMs from Cx3cr1CreERT2R26tdT mice. Symbols represent individual mice, n = 2 (2w BLZ945), n = 3 (8w, 26w CTL), n = 4 (2w CTL, 26w BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (e-h) Flow cytometric analysis. Representative histograms (e) and quantification of reporter gene expression in dCAMs from HexbCreERT2R26YFP (f), Mrc1CreERT2R26tdT (g) and Cxcr4CreERT2R26tdT (h) mice. Symbols represent individual mice, HexbCreERT2R26YFP: n = 1 (2w CTL), n = 2 (2w BLZ945), n = 3 (8w CTL, 26w CTL), n = 4 (8w BLZ945, 26w BLZ945), Mrc1CreERT2R26tdT: n = 3 (2w CTL, 26w CTL), n = 4 (2w BLZ945, 26w BLZ945), n = 6 (8w), Cxcr4CreERT2R26tdT: n = 3 (2w, 8w, 12w, 26w all CTL), n = 4 (8w BLZ945, 12w BLZ945), n = 5 (26w BLZ945), n = 6 (5d CTL, 2w BLZ945), n = 7 (5d BLZ945), mean ± SEM. Ordinary one-way ANOVA with Šidák-adjusted multiple-comparisons p values. (i, j) Representative images (i) and histological quantification (j) of tdT+ dCAMs in extrasinus and sinus area of cranial dura mater. Arrowheads: open = tdT− dCAMs; orange = tdT− dCAMs. Symbols represent individual mice, n = 3 (CTL), n = 4 (BLZ945), mean ± SEM. Ordinary one-way ANOVA with Tukey HSD-adjusted multiple-comparisons p values. (k-n) Experimental schemes and quantification of tdT+ dCAMs by flow cytometry (FC). Symbols represent individual mice; k: n = 4 (21d, 8w CTL), n = 9 (8w BLZ945); l: n = 3 per group; m: n = 5 per group; n: n = 9 (CTL), n = 8 (BLZ945); mean ± SEM. Ordinary one-way ANOVA with Tukey’s multiple comparison test, multiplicity adjusted p values (k) or Unpaired two-tailed t-test (l, m, n). (o) Experimental scheme and quantification CD45.2+ dCAMs by FC. Symbols represent individual mice, n = 3 (CTL), n = 4 (BLZ945), mean ± SEM. Unpaired two-tailed t-test.
Extended Data Fig. 6 Cluster markers of microglia and sdCAMs.
Dot plot depicting selected cell-type-defining markers. Color scale: Relative expression levels. (b, c) Feature plots of selected cluster defining markers for MG clusters 1–5 (MG1-5) (b) and sdCAM clusters 1–4 (sdCAM1-4) (c) (Related to Fig. 5c). Scale bars show z-scores of minimum to maximum scaled gene expression. (d) Venn diagram depicting overlay between DEGs (Bonferroni adjusted p < 0.05 and |log2 fold change| > 0.59) between all sdCAMs from BLZ945 vs CTL mice and DEGs between CAM1 and 3 vs CAM2. Top 10 overlapping up (left) and down (right) regulated genes are shown. (e) Gene ontology (GO) term analysis showing upregulated terms from sdCAM1 and 3 compared to sdCAM2. p values are Benjamini–Hochberg adjusted. (f) Heatmap depicting the top 472 DEGs (Benjamini–Hochberg adjusted p < 0.05 and |log2 fold change| > 1) from the bulk RNA-seq analysis (Related to Fig. 5g). n = 4 mice per group, scale bar indicates z-score normalized expression. (g) GO term analysis of CD206+tdT+ CAMs and CD206+tdT− CAMs. p values are Benjamini–Hochberg adjusted. (h) Genes found to be upregulated in either CD206+tdT+ CAMs or CD206+tdT− CAMs in the bulk RNA-seq dataset (Fig. 5i) were provided as input to the scRNA-seq CAM dataset to generate cluster-wise heatmaps.
Extended Data Fig. 7 scATAC-seq of repopulated CNS macrophages.
(a) Feature plots displaying module scores calculated from gene activities for the distinct cell populations based on scATAC-seq (top) or for the same genes based on gene expression levels derived from scRNA-seq from Fig. 2 (bottom). Scale bars show z-scores of minimum to maximum scaled gene activity (top) or gene expression (bottom). (b) Heatmap depicting gene activities of identified scATAC-seq marker genes for CNS macrophage populations. Scale bar shows z-score of gene activity. (c) Coverage plots of Sall1, Ms4a7, Cd83 and Ly6c2 loci depicting the aggregated ATAC reads for each cell cluster. Differentially accessible peaks are highlighted in gray. (d) Feature plot overlaying the treatment (top) and bar plot depicting the number of nuclei derived from either of the two treatments within each cell cluster (bottom). (e) Volcano plots comparing accessible peaks in dCAMs from 8 weeks (8w) post BLZ945 to control (CTL). Bonferroni-corrected p values, NS: p > 0.05. (f) Bar plots showing inferred transcription factor activities in hom_ysCAM and hscCAM computed from chromVAR deviation z-scores using accessible transcription factor motifs. Scale bar shows fold enrichment over accessibility of respective motifs in all cells. p values are Benjamini–Hochberg adjusted. (g) Position weight matrix logos of enriched transcription factor motifs for hom_ysCAM and hscCAM. The height of each nucleotide letter is proportional to its frequency at that position, reflecting sequence conservation.
Extended Data Fig. 8 Response of repopulated CNS macrophages to LPS challenge.
(a) Experimental overview is displayed. Following BLZ945 treatment and tamoxifen injections, Ccr2CreERT2R26tdT mice were i.p. injected with PBS or LPS dural macrophages (dCAMs) were sorted 12h later for bulk RNA-sequencing. n = 5 mice per condition. (b) Principal-component analysis (PCA) of tdT− dCAMs and tdT+ dCAMs isolated from BLZ945-treated PBS or LPS injected mice. (c) Heatmap of all 715 differentially expressed genes (DEGs; Benjamini–Hochberg adjusted p < 0.05 and |log2 fold change| > 1) detected between groups. n = 5 mice per group, scale bar indicates z-score normalized expression. (d) Volcano plots showing DEGs between BLZ945-treated PBS or LPS injected mice for MG, tdT− sdCAMs and tdT+ sdCAMs. The top upregulated GO terms following LPS treatment are shown. Benjamini–Hochberg adjusted p values, NS: p > 0.05. (e) Volcano plots showing DEGs between tdT− dCAMs and tdT+ dCAMs in BLZ945-treated PBS (left) or BLZ945 LPS injected (right) mice. Benjamini–Hochberg adjusted p values, NS: p > 0.05. (f) Volcano plots showing DEGs between BLZ945 LPS injected versus PBS injected mice for tdT− dCAMs (top) and tdT+ dCAMs (bottom). The top upregulated GO terms following LPS treatment are shown. Benjamini–Hochberg adjusted p values, NS: p > 0.05. (g) Reclustered PCA of tdT− CAMs and tdT+ CAMs. (h, i) Logarithmic scaled radar charts depicting normalized expression levels of selected genes for tdT− dCAMs (top) as well as tdT+ dCAMs MG tdT− as well as tdT+ dCAMs following systemic LPS challenge (h) and for CTL versus LPS challenge conditions for the individual populations (i).
Extended Data Fig. 9 Ccr2 deficiency prevents differential responsiveness of repopulated sdCAMs.
(a) Heatmap of the top 250 most variable genes. Scale bar indicates z-score normalized expression. (b) Volcano plots showing differentially expressed genes (DEGs) between MG or sdCAMs isolated from CTL or BLZ945-treated mice following LPS treatment, respectively. Benjamini–Hochberg adjusted p values, NS: p > 0.05.
Extended Data Fig. 10 sdCAM replacement influences stroke without affecting structural characteristics.
(a-c) Representative magnetic resonance imaging (MRI) one day after (+1d) MCAO. Quantification of lesion volume (a), angiographic score (b) and hemorrhage score (c). Symbols represent individual mice, n = 10 (CTL), n = 8 (BLZ945). Mean ± SEM and unpaired two-tailed t-test (a). Median ± interquartile range, two-tailed Mann-Whitney test (b, c). (d, e) Heatmap of all 353 differentially expressed genes (DEGs; Benjamini–Hochberg adjusted p < 0.05 and |log2 fold change| > 0.59) between ipsilateral and contralateral brain tissue (d) and additional selected stroke response genes. n = 4 mice per group. Scale bar indicates z-score normalized expression (d). Median ± interquartile range, ordinary one-way ANOVA with Tukey’s HSD test (e). (f-k) Transcriptional analysis of sdCAMs from WT stroke mice. Heatmap of all 2,346 DEGs (f) and volcano plots (g) of DEGs between ipsilateral and contralateral hemisphere, Benjamini–Hochberg adjusted p < 0.05 and |log2 fold change| > 0.59, NS: p > 0.05. Overlap of stroke response genes (h). Gene ontology (GO) terms derived from treatment-specific sdCAM stroke response genes (i) and selected stroke response genes (j, k). sdCAMs from two mice each were pooled; n = 3 (ispilateral), n = 4 (contralateral) biological replicates. Scale bar indicates z-score normalized expression (f). Median ± interquartile range and ordinary one-way ANOVA with Tukey’s HSD test (j, k). (l-o) Transcriptional analysis of microglia (MG) and sdCAMs from Ccr2CreERT2R26tdT no stroke (∅) and stroke mice. Heatmap of all 642 DEGs between MG, tdT− sdCAMs and tdT+ sdCAMs (l). GO terms derived from sdCAM subtype specific stroke response genes (m). DEGs in MG with Benjamini–Hochberg adjusted p values (n). Boxplots of selected genes (o). n = 3 (tdT+ sdCAMs BLZ945), n = 5 (all other groups). Scale bar indicates z-score normalized expression (l). Median ± interquartile range and ordinary one-way ANOVA with Tukey’s HSD test (o).
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–6 and Table 1
Supplementary Source Data (download XLSX )
Source Data for Supplementary Figures.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5.
Source Data Fig. 8 (download XLSX )
Statistical source data for Fig. 8.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2 (download XLSX )
Statistical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 3 (download XLSX )
Statistical source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Statistical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data for Extended Data Fig. 10.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fliegauf, M., Levard, D., Cardamone, F. et al. Distinct origins and niches determine the cellular responsiveness of CNS macrophages after repopulation. Nat Immunol (2026). https://doi.org/10.1038/s41590-026-02457-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41590-026-02457-y