Abstract
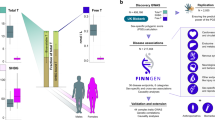
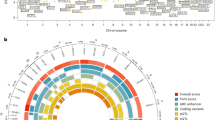
The genetic background of many female reproductive health diagnoses remains uncharacterized, compromising our understanding of the underlying biology. Here, we map the genetic architecture across 42 female-specific health conditions using data from up to 293,618 women from two large population-based cohorts, the Estonian Biobank and the FinnGen study. Our study illustrates the utility of genetic analyses in understanding women’s health better. As specific examples, we describe genetic risk factors for ovarian cysts that elucidate the genetic determinants of folliculogenesis and, by leveraging population-specific variants, uncover new candidate genes for uterine fibroids. We find that most female reproductive health diagnoses have a heritable component, with varying degrees of polygenicity and discoverability. Finally, we identify pleiotropic loci and genes that function in genital tract development (WNT4, PAX8, WT1, SALL1), hormonal regulation (FSHB, GREB1, BMPR1B, SYNE1/ESR1) and folliculogenesis (CHEK2), underlining their integral roles in female reproductive health.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Summary statistics from the meta-analyses described in this paper can be accessed from the GWAS Catalog (https://www.ebi.ac.uk/gwas/) (accession codes GCST90454202–GCST90454243). FinnGen summary statistics can be accessed as described at https://www.finngen.fi/en/access_results. The procedure to access EstBB individual-level data has been described at https://genomics.ut.ee/en/content/estonian-biobank#dataaccess.
The Trøndelag Health Study (HUNT) may be accessed by application to the HUNT Research Centre at https://www.ntnu.edu/hunt/data. The Trøndelag Health Study (HUNT) has invited persons aged 13–100 years to four surveys between 1984 and 2019. Comprehensive data from more than 140,000 persons who have participated at least once and biological material from 78,000 persons are collected. Data are stored in the HUNT databank, and biological material is stored in the HUNT biobank. The HUNT Research Centre has permission from the Norwegian Data Inspectorate to store and handle these data. The key identification in the database is the personal identification number given to all Norwegians at birth or immigration, whereas deidentified data are sent to researchers upon approval of a research protocol by the Regional Ethical Committee and the HUNT Research Centre. To protect participants’ privacy, the HUNT Research Centre aims to limit the storage of data outside the HUNT databank and cannot deposit data in open repositories. The HUNT databank has precise information on all data exported to different projects and is able to reproduce these on request. There are no restrictions regarding data export given approval of applications to the HUNT Research Centre. For more information, see http://www.ntnu.edu/hunt/data.
The following databases used in the analyses are freely available online:
1. To identify population-specific variants, we performed allele frequency look-up using the Open Targets Genetics v8 portal (https://genetics.opentargets.org/) and the gnomAD v2.1 database (https://gnomad.broadinstitute.org/).
2. To establish the potential biological role of candidate genes for the ovarian cysts phenotype, we queried the Mouse Genome Informatics database (https://www.informatics.jax.org/).
Other datasets (for example, the GRCh37 reference genome) used in the analyses are integrated components of the publicly available platforms and tools used for the data analysis.
Code availability
All software programs used for the analyses described in this paper are freely available online: GWAMA v2.1 (https://genomics.ut.ee/en/tools); REGENIE v2.0.2 and v2.2.4 (https://github.com/rgcgithub/regenie); UCSC liftOver command line tool version downloaded from the UCSC Genome Browser toolkit (http://genome.ucsc.edu/); FUMA v1.4.0 (https://fuma.ctglab.nl/); MAGMA v1.08 integrated in FUMA v1.4.0; LDSC v1.0.1 (https://github.com/bulik/ldsc); MiXeR v1.3 (https://github.com/precimed/mixer); Fuji plot (https://github.com/mkanai/fujiplot); PLINK2 (www.cog-genomics.org/plink/2.0/); PRS-CS (last version updated January 4, 2021, https://github.com/getian107/PRScs); PGS Catalog Calculator v2.0.0-beta.3 (https://github.com/PGScatalog/pgsc_calc); R package PheWAS v1.0. For FinnGen, code to perform GWAS analyses is available at the FinnGen GitHub (https://github.com/FINNGEN/). Individual plots were created using R v3.6.3, v4.2.2 and v4.3.2, including the R packages corrplot v0.92, ggplot2 v3.5.1 and tidyverse v2.0.0. The final figures were edited with Inkscape.
References
Mercuri, N. D. & Cox, B. J. The need for more research into reproductive health and disease. eLife 11, e75061 (2022).
A life-course approach to women’s health. Nat. Med. 30, 1 (2024).
Abdellaoui, A., Yengo, L., Verweij, K. J. H. & Visscher, P. M. 15 years of GWAS discovery: realizing the promise. Am. J. Hum. Genet. 110, 179–194 (2023).
Day, F. et al. Large-scale genome-wide meta-analysis of polycystic ovary syndrome suggests shared genetic architecture for different diagnosis criteria. PLoS Genet. 14, e1007813 (2018).
Moolhuijsen, L. M. E. et al. Genomic and proteomic evidence for hormonal and metabolic foundations of polycystic ovary syndrome. Preprint at medRxiv https://doi.org/10.1101/2024.04.18.24306020 (2024).
Rahmioglu, N. et al. The genetic basis of endometriosis and comorbidity with other pain and inflammatory conditions. Nat. Genet. 55, 423–436 (2023).
O’Mara, T. A. et al. Identification of nine new susceptibility loci for endometrial cancer. Nat. Commun. 9, 3166 (2018).
Phelan, C. M. et al. Identification of 12 new susceptibility loci for different histotypes of epithelial ovarian cancer. Nat. Genet. 49, 680–691 (2017).
Koel, M. et al. GWAS meta-analyses clarify the genetics of cervical phenotypes and inform risk stratification for cervical cancer. Hum. Mol. Genet. 32, 2103–2116 (2023).
Honigberg, M. C. et al. Polygenic prediction of preeclampsia and gestational hypertension. Nat. Med. 29, 1540–1549 (2023).
Tyrmi, J. S. et al. Genetic risk factors associated with preeclampsia and hypertensive disorders of pregnancy. JAMA Cardiol. 8, 674–683 (2023).
Pervjakova, N. et al. Multi-ancestry genome-wide association study of gestational diabetes mellitus highlights genetic links with type 2 diabetes. Hum. Mol. Genet. 31, 3377–3391 (2022).
Elliott, A. et al. Distinct and shared genetic architectures of gestational diabetes mellitus and type 2 diabetes. Nat. Genet. 56, 377–382 (2024).
Fejzo, M. S. et al. Placenta and appetite genes GDF15 and IGFBP7 are associated with hyperemesis gravidarum. Nat. Commun. 9, 1178 (2018).
Pujol Gualdo, N., Estonian Biobank Research Team, Mägi, R. & Laisk, T. Genome-wide association study meta-analysis supports association between MUC1 and ectopic pregnancy. Hum. Reprod. 38, 2516–2525 (2023).
Solé-Navais, P. et al. Genetic effects on the timing of parturition and links to fetal birth weight. Nat. Genet. 55, 559–567 (2023).
Kentistou, K. A. et al. Understanding the genetic complexity of puberty timing across the allele frequency spectrum. Nat. Genet. 56, 1397–1411 (2024).
Ruth, K. S. et al. Genetic insights into biological mechanisms governing human ovarian ageing. Nature 596, 393–397 (2021).
Xiao, B. et al. Inference of causal relationships between genetic risk factors for cardiometabolic phenotypes and female-specific health conditions. J. Am. Heart Assoc. 12, e026561 (2023).
Leitsalu, L. et al. Cohort profile: Estonian Biobank of the Estonian Genome Center, University of Tartu. Int. J. Epidemiol. 44, 1137–1147 (2015).
Milani, L. et al. From biobanking to personalized medicine: the journey of the Estonian Biobank. Preprint at medRxiv https://doi.org/10.1101/2024.09.22.24313964 (2024).
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613, 508–518 (2023).
Gallagher, C. S. et al. Genome-wide association and epidemiological analyses reveal common genetic origins between uterine leiomyomata and endometriosis. Nat. Commun. 10, 4857 (2019).
Dixon, P. H. et al. GWAS meta-analysis of intrahepatic cholestasis of pregnancy implicates multiple hepatic genes and regulatory elements. Nat. Commun. 13, 4840 (2022).
Pathare, A. D. S. et al. A large-scale genome-wide association study on female genital tract polyps highlights role of DNA repair, cell proliferation, and cell growth. Hum. Reprod. deaf025 (2025).
Ghoussaini, M. et al. Open Targets Genetics: systematic identification of trait-associated genes using large-scale genetics and functional genomics. Nucleic Acids Res. 49, D1311–D1320 (2021).
Mountjoy, E. et al. An open approach to systematically prioritize causal variants and genes at all published human GWAS trait-associated loci. Nat. Genet. 53, 1527–1533 (2021).
Buyukcelebi, K. et al. Integrating leiomyoma genetics, epigenomics, and single-cell transcriptomics reveals causal genetic variants, genes, and cell types. Nat. Commun. 15, 1169 (2024).
Laisk, T. et al. Large-scale meta-analysis highlights the hypothalamic–pituitary–gonadal axis in the genetic regulation of menstrual cycle length. Hum. Mol. Genet. 27, 4323–4332 (2018).
Tyrmi, J. S. et al. Leveraging Northern European population history: novel low-frequency variants for polycystic ovary syndrome. Hum. Reprod. 37, 352–365 (2022).
Sliz, E. et al. Evidence of a causal effect of genetic tendency to gain muscle mass on uterine leiomyomata. Nat. Commun. 14, 542 (2023).
Välimäki, N. et al. Genetic predisposition to uterine leiomyoma is determined by loci for genitourinary development and genome stability. eLife 7, e37110 (2018).
Bray, M. J. et al. Admixture mapping of uterine fibroid size and number in African American women. Fertil. Steril. 108, 1034–1042 (2017).
Ward, L. D. et al. Rare coding variants in DNA damage repair genes associated with timing of natural menopause. HGG Adv. 3, 100079 (2021).
Zhang, W. et al. Overexpression of myosin is associated with the development of uterine myoma. J. Obstet. Gynaecol. Res. 40, 2051–2057 (2014).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Pujol-Gualdo, N. et al. Advancing our understanding of genetic risk factors and potential personalized strategies for pelvic organ prolapse. Nat. Commun. 13, 3584 (2022).
Holland, D. et al. Beyond SNP heritability: polygenicity and discoverability of phenotypes estimated with a univariate Gaussian mixture model. PLoS Genet. 16, e1008612 (2020).
Venkatesh, S. S. et al. Genome-wide analyses identify 21 infertility loci and over 400 reproductive hormone loci across the allele frequency spectrum. Preprint at medRxiv https://doi.org/10.1101/2024.03.19.24304530 (2024).
Bulik-Sullivan, B. et al. An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Pan, M.-L., Chen, L.-R. & Chen, K.-H. Prepregnancy polycystic ovary syndrome as a risk factor of subsequent preterm labor: a national population-based cohort study. Int. J. Environ. Res. Public Health 19, 5470 (2022).
Roos, N. et al. Risk of adverse pregnancy outcomes in women with polycystic ovary syndrome: population based cohort study. BMJ 343, d6309 (2011).
Jin, L.-Y. et al. Overexpression of Pde4d in rat granulosa cells inhibits maturation and atresia of antral follicles to induce polycystic ovary. Biochim. Biophys. Acta Mol. Basis Dis. 1870, 166869 (2024).
Jensterle, M., Kocjan, T. & Janez, A. Phosphodiesterase 4 inhibition as a potential new therapeutic target in obese women with polycystic ovary syndrome. J. Clin. Endocrinol. Metab. 99, E1476–E1481 (2014).
Lim, E. T. et al. Distribution and medical impact of loss-of-function variants in the Finnish founder population. PLoS Genet. 10, e1004494 (2014).
Chheda, H. et al. Whole-genome view of the consequences of a population bottleneck using 2926 genome sequences from Finland and United Kingdom. Eur. J. Hum. Genet. 25, 477–484 (2017).
Kivisild, T. et al. Patterns of genetic connectedness between modern and medieval Estonian genomes reveal the origins of a major ancestry component of the Finnish population. Am. J. Hum. Genet. 108, 1792–1806 (2021).
Mitt, M. et al. Improved imputation accuracy of rare and low-frequency variants using population-specific high-coverage WGS-based imputation reference panel. Eur. J. Hum. Genet. 25, 869–876 (2017).
Day, F. R. et al. Causal mechanisms and balancing selection inferred from genetic associations with polycystic ovary syndrome. Nat. Commun. 6, 8464 (2015).
Varas Enriquez, P. J., McKerracher, L. J. & Elliot, M. G. Pre-eclampsia and maternal–fetal conflict. Evol. Med. Public Health 2018, 217–218 (2018).
Pavličev, M. et al. A common allele increases endometrial Wnt4 expression, with antagonistic implications for pregnancy, reproductive cancers, and endometriosis. Nat. Commun. 15, 1152 (2024).
Wilcox, N. et al. Exome sequencing identifies breast cancer susceptibility genes and defines the contribution of coding variants to breast cancer risk. Nat. Genet. 55, 1435–1439 (2023).
Breast Cancer Association Consortium et al. Breast cancer risk genes—association analysis in more than 113,000 women. N. Engl. J. Med. 384, 428–439 (2021).
Patel, A. P. et al. A multi-ancestry polygenic risk score improves risk prediction for coronary artery disease. Nat. Med. 29, 1793–1803 (2023).
Mars, N. et al. The role of polygenic risk and susceptibility genes in breast cancer over the course of life. Nat. Commun. 11, 6383 (2020).
Obstetrics Group of the Gynecology and Obstetrics Branch of Chinese Medical Association, Perinatal Medicine Branch of Chinese Medical Association, Yu, X., Yang, H. & Qi, H. Clinical management guidelines for intrahepatic cholestasis of pregnancy. Matern. Fetal Med. 6, 13–22 (2024).
Lennon, N. J. et al. Selection, optimization and validation of ten chronic disease polygenic risk scores for clinical implementation in diverse US populations. Nat. Med. 30, 480–487 (2024).
Khan, A. et al. Polygenic risk alters the penetrance of monogenic kidney disease. Nat. Commun. 14, 8318 (2023).
Fahed, A. C. et al. Polygenic background modifies penetrance of monogenic variants for tier 1 genomic conditions. Nat. Commun. 11, 3635 (2020).
Krebs, K. et al. Genome-wide study identifies association between HLA-B∗55:01 and self-reported penicillin allergy. Am. J. Hum. Genet. 107, 612–621 (2020).
Leitsalu, L., Alavere, H., Tammesoo, M.-L., Leego, E. & Metspalu, A. Linking a population biobank with national health registries—the Estonian experience. J. Pers. Med. 5, 96–106 (2015).
Mbatchou, J. et al. Computationally efficient whole-genome regression for quantitative and binary traits. Nat. Genet. 53, 1097–1103 (2021).
Kent, W. J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Mägi, R. & Morris, A. P. GWAMA: software for genome-wide association meta-analysis. BMC Bioinformatics 11, 288 (2010).
Watanabe, K., Taskesen, E., van Bochoven, A. & Posthuma, D. Functional mapping and annotation of genetic associations with FUMA. Nat. Commun. 8, 1826 (2017).
Lee, S. H., Wray, N. R., Goddard, M. E. & Visscher, P. M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Pasman, J. A. et al. GWAS of lifetime cannabis use reveals new risk loci, genetic overlap with psychiatric traits, and a causal influence of schizophrenia. Nat. Neurosci. 21, 1161–1170 (2018).
Romero, C. et al. Exploring the genetic overlap between twelve psychiatric disorders. Nat. Genet. 54, 1795–1802 (2022).
Watanabe, K. et al. A global overview of pleiotropy and genetic architecture in complex traits. Nat. Genet. 51, 1339–1348 (2019).
Kanai, M. et al. Genetic analysis of quantitative traits in the Japanese population links cell types to complex human diseases. Nat. Genet. 50, 390–400 (2018).
Ge, T., Chen, C.-Y., Ni, Y., Feng, Y.-C. A. & Smoller, J. W. Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nat. Commun. 10, 1776 (2019).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Carroll, R. J., Bastarache, L. & Denny, J. C. R PheWAS: data analysis and plotting tools for phenome-wide association studies in the R environment. Bioinformatics 30, 2375–2376 (2014).
Krokstad, S. et al. Cohort profile: the HUNT study, Norway. Int. J. Epidemiol. 42, 968–977 (2013).
Åsvold, B. O. et al. Cohort profile update: the HUNT study, Norway. Int. J. Epidemiol. 52, e80–e91 (2023).
Brumpton, B. M. et al. The HUNT study: a population-based cohort for genetic research. Cell Genom. 2, 100193 (2022).
McCarthy, S. et al. A reference panel of 64,976 haplotypes for genotype imputation. Nat. Genet. 48, 1279–1283 (2016).
Lambert, S. A. et al. Enhancing the Polygenic Score Catalog with tools for score calculation and ancestry normalization. Nat. Genet. https://doi.org/10.1038/s41588-024-01937-x (2024).
Lambert, S. A. et al. The Polygenic Score Catalog as an open database for reproducibility and systematic evaluation. Nat. Genet. 53, 420–425 (2021).
Acknowledgements
This study was funded by the Ministry of Education and Research Centres of Excellence grant TK214 Centre of Excellence for Personalised Medicine and by the Estonian Research Council grants PSG776, PRG1911 and PRG1076; and the European Union’s Horizon 2020 research and innovation program under the MATER Marie Skłodowska-Curie grant agreement no. 813707. The analyses were performed in the High-Performance Computing Center of the University of Tartu. We thank the participants of the FinnGen and EstBB studies. This work was partially written at writing retreats organized by the Institute of Genomics, University of Tartu.
The Trøndelag Health Study (HUNT) is a collaboration between the HUNT Research Centre (Faculty of Medicine and Health Sciences, Norwegian University of Science and Technology, NTNU), Trøndelag County Council, Central Norway Regional Health Authority and the Norwegian Institute of Public Health. The genotyping in HUNT was financed by the National Institutes of Health; the University of Michigan; the Research Council of Norway; the Liaison Committee for Education, Research and Innovation in central Norway; and the Joint Research Committee between St. Olavs Hospital and the Faculty of Medicine and Health Sciences, NTNU. Members of the HUNT All-In Research Team (in alphabetical order by surname): B. O. Åsvold, B. Brumpton, M. E. Gabrielsen, K. Hveem, I. Surakka, L. Thomas and W. Zhou.
Author information
Authors and Affiliations
Consortia
Contributions
N.P.G., R.M., P.P. and T.L. designed the study. The EstBB Research Team collected and provided the genotype and phenotype data used in the EstBB cohort-level analyses. B.M.B. provided the genotype and phenotype data used in HUNT analyses. N.P.G., J.D., V.R., F.-D.P., B.N.W., K.L., M.G., L.H. and T.L. performed and interpreted the analyses. N.P.G., J.D., V.R. and T.L. wrote the paper. M.V. performed final edits of the figures. All authors discussed the results and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Medicine thanks Peter Dixon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Sonia Muliyil and Liam Messin, in collaboration with the Nature Medicine team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Note, Supplementary Tables 2 and 6 and Supplementary Figs. 1 and 2.
Supplementary Tables (download XLSX )
Supplementary Tables 1, 3–5 and 7–17.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Pujol Gualdo, N., Džigurski, J., Rukins, V. et al. Atlas of genetic and phenotypic associations across 42 female reproductive health diagnoses. Nat Med 31, 1626–1634 (2025). https://doi.org/10.1038/s41591-025-03543-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41591-025-03543-8
This article is cited by
-
The "double-edged sword" effect of non-steroidal anti-inflammatory drugs (NSAIDs) in the treatment of endometriosis (EMS)
Reproductive Biology and Endocrinology (2025)
-
Intrahepatic cholestasis of pregnancy
Nature Reviews Disease Primers (2025)