Abstract
Transposon (IS200/IS605)-encoded TnpB proteins are predecessors of class 2 type V CRISPR effectors and have emerged as one of the most compact genome editors identified thus far. Here, we optimized the design of Deinococcus radiodurans (ISDra2) TnpB for application in mammalian cells (TnpBmax), leading to an average 4.4-fold improvement in editing. In addition, we developed variants mutated at position K76 that recognize alternative target-adjacent motifs (TAMs), expanding the targeting range of ISDra2 TnpB. We further generated an extensive dataset on TnpBmax editing efficiencies at 10,211 target sites. This enabled us to delineate rules for on-target and off-target editing and to devise a deep learning model, termed TnpB editing efficiency predictor (TEEP; https://www.tnpb.app), capable of predicting ISDra2 TnpB guiding RNA (ωRNA) activity with high performance (r > 0.8). Employing TEEP, we achieved editing efficiencies up to 75.3% in the murine liver and 65.9% in the murine brain after adeno-associated virus (AAV) vector delivery of TnpBmax. Overall, the set of tools presented in this study facilitates the application of TnpB as an ultracompact programmable endonuclease in research and therapeutics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All ωRNA and HTS primer sequences used for this study are provided in Supplementary Data 1. Deep amplicon sequencing data files are available from the National Center for Biotechnology Information’s Sequence Read Archive (accession PRJNA1019264). Plasmid sequences are provided at https://benchling.com/marquark7/f_/FOdfdV1v-tnpb/. Additionally, key plasmids from this work are available from Addgene. All data are freely accessible to the public.
Code availability
Computer code for the analysis of the pooled libraries is available at https://github.com/Schwank-Lab/tnpb. The code for training the machine learning models is available on GitHub (https://github.com/uzh-dqbm-cmi/Tnpb). In addition, we have developed a publicly available web application (https://go.tnpb.app or https://www.tnpb.app) for predicting TnpB ωRNA efficiencies using our trained models. HTS data were collected and demultiplexed by Illumina NovaSeq Control software version 1.7 and MiSeq Control software (versions 3.1 and 4.0). Pooled library analysis was performed using Python 3.9. Cutadapt (3.5) was used to trim sequencing reads. For characterization of indels and base edits at single sites (endogenous), CRISPResso2 (2.2.7) was used. For statistical analysis, SciPy (1.10.1) and Prism (9.0.0) were used.
References
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Altae-Tran, H. et al. The widespread IS200/IS605 transposon family encodes diverse programmable RNA-guided endonucleases. Science 374, 57–65 (2021).
Karvelis, T. et al. Transposon-associated TnpB is a programmable RNA-guided DNA endonuclease. Nature 599, 692–696 (2021).
Nakagawa, R. et al. Cryo-EM structure of the transposon-associated TnpB enzyme. Nature 616, 390–397 (2023).
Sasnauskas, G. et al. TnpB structure reveals minimal functional core of Cas12 nuclease family. Nature 616, 384–389 (2023).
Schmidheini, L. et al. Continuous directed evolution of a compact CjCas9 variant with broad PAM compatibility. Nat. Chem. Biol. 20, 333–343 (2023).
Koblan, L. W. et al. Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction. Nat. Biotechnol. 36, 843–846 (2018).
Suzuki, K. et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Xiang, G. et al. Evolutionary mining and functional characterization of TnpB nucleases identify efficient miniature genome editors. Nat. Biotechnol. 42, 745–757 (2023).
Saito, M. et al. Fanzor is a eukaryotic programmable RNA-guided endonuclease. Nature 620, 660–668 (2023).
Koblan, L. W. et al. Efficient C•G-to-G•C base editors developed using CRISPRi screens, target-library analysis, and machine learning. Nat. Biotechnol. 39, 1414–1425 (2021).
Komor, A. C. et al. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016).
Gaudelli, N. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017).
Richter, M. F. et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat. Biotechnol. 38, 883–891 (2020).
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015).
Walton, R. T., Hsu, J. Y., Joung, J. K. & Kleinstiver, B. P. Scalable characterization of the PAM requirements of CRISPR–Cas enzymes using HT-PAMDA. Nat. Protoc. 16, 1511–1547 (2021).
Marquart, K. F. et al. Predicting base editing outcomes with an attention-based deep learning algorithm trained on high-throughput target library screens. Nat. Commun. 12, 5114 (2021).
Clement, K. et al. CRISPResso2 provides accurate and rapid genome editing sequence analysis. Nat. Biotechnol. 37, 224–226 (2019).
Vriend, L. E. M., Jasin, M. & Krawczyk, P. M. Assaying break and nick-induced homologous recombination in mammalian cells using the DR-GFP reporter and Cas9 nucleases. Methods Enzymol. 546, 175–191 (2014).
Tsai, S. Q., Topkar, V. V., Joung, J. K. & Aryee, M. J. Open-source guideseq software for analysis of GUIDE-seq data. Nat. Biotechnol. 34, 483 (2016).
Turchiano, G. et al. Quantitative evaluation of chromosomal rearrangements in gene-edited human stem cells by CAST-seq. Cell Stem Cell 28, 1136–1147 (2021).
Klermund, J. et al. On- and off-target effects of paired CRISPR–Cas nickase in primary human cells. Mol. Ther. 32, 1298–1310 (2024).
Srivastava, N., Hinton, G., Krizhevsky, A., Sutskever, I. & Salakhutdinov, R. Dropout: a simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 15, 1929–1958 (2014).
Kingma, D. P. & Ba, J. Adam: a method for stochastic optimization. Preprint at https://arxiv.org/abs/1412.6980 (2017).
Acknowledgements
We thank the Functional Genomics Center Zurich for technical support and access to instruments at the University of Zurich and ETH Zürich, the mRNA platform at UZH–USZ and S. Pascolo, J. Frei and C. Wyss for the production and purification of RNA, the Viral Vector Facility of UZH and J.-C. Paterna and M. Rauch for production of AAVs, G. Andrieux for bioinformatic analysis of CAST-seq data and O. Melkonyan for HT-TAMDA analysis as well as J. Häberle and N. Rimann for measurements of blood LDL levels. We thank I. Querques, M. Jinek, M. Pacesa, L.-M. Koch, Lotti and members of the Schwank laboratory for valuable discussions, feedback and help throughout the study. This work was supported by the University Research Priority Programs ‘Human Reproduction Reloaded’ (to G.S.) and ‘ITINERARE’ (to G.S. and M. Krauthammer), the ProMedica Foundation (to G.S.), the Swiss National Science Foundation grant numbers 185293 and 214936 (to G.S.) and grant number 201184 (to M. Krauthammer), a UZH PhD fellowship (to T.R.), ETH PhD fellowships (to L.S. and K.F.M.) and the German Research Foundation (CRC 1597-A05 to T.C.).
Author information
Authors and Affiliations
Contributions
K.F.M. performed numerous biological experiments throughout the study, analyzed data and prepared figures. N.M. performed bioinformatic analysis of all target-matched library experiments, prepared figures, curated data for the machine learning models and contributed to XGBoost model design. A.M. designed and developed machine learning models and implemented the web app for TEEP. S.M. prepared plasmids for TnpB and Fanzor and ωRNA expression, performed and analyzed endogenous DNA-editing experiments, conducted HT-TAMDA assays and performed western blotting experiments. L.K. and T.R. performed in vivo experiments, including intracerebroventricular and stereotactic injections and brain and hepatocyte isolation. L.S. prepared plasmids for ωRNA expression and conducted HT-TAMDA assays. P.I.K. performed and analyzed GUIDE-seq experiments. A.A. contributed to the design and development of machine learning models. M.M.K. performed CAST-seq experiments. M.M. assessed inflammation-linked cytokines. T.H. contributed to western blotting experiments. T.C., M. Kopf, M. Krauthammer and G.S. supervised the research and provided field-specific expertise. K.F.M. and G.S. designed the study and wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.F.M. and G.S. are co-inventors on a patent application filed by the University of Zurich relating to the work described in this paper. G.S. is an advisor to Prime Medicine. The other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Lei Tang, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Benchmarking of TnpB and Fanzor architectures in HEK293T cells.
(a) Schematic representation of experimental workflow and designs. NLS, nuclear localization sequence; BPNLS, bipartite NLS; SRAD, Serine-Arginine-Alanine-Aspartic acid; GS, Glycine-Serine; PuroR, Puromycin resistance; d, days; HTS, high-throughput sequencing; a codon-optimization and design from Xiang et al.11 and Saito et al.12 (b–d) Benchmarking of different architectures of ISDra2, ISAam1 and ISYmu1 TnpBs. Number of analyzed endogenous targets: ISDra2 TnpB, N = 7; ISAam1 TnpB, N = 7; ISYmu1 TnpB, N = 8. Each dot represents the mean of n = 3 independent biological replicates; the black bar represents the mean of all target sites tested for the respective design. Means were compared by two-tailed t-test. (e) Benchmarking of SpuFz1-v2 Fanzor embedded in various designs tested at one endogenous locus (B2M). Each bar represents the mean ± s.d. of n = 3 independent biological replicates and a two-tailed t-test was used to calculate variance. Indel frequencies are shown in Datafile S1.
Extended Data Fig. 2 High-throughput TAM determination assay (HT-TAMDA) of TnpBmax and variants thereof.
The log10 (rate constant) represents the mean of two replicates against two distinct target sequences.
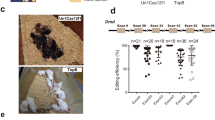
Extended Data Fig. 3 Direct intracortical injection of scAAV-TnpB-Dnmt1.
a) Schematic representation of stereotactic scAAV injection. (b, c) TnpBmax mediated editing at the Dnmt1 locus determined by deep amplicon sequencing in separated brain regions of mice treated with 5.0 × 1013 vg/kg scAAV. CTX, cortex; BS, brain stem; Hipp, hippocampus; Hypo, hypothalamus; MB, midbrain; OB, olfactory bulb; ST, striatum; TM, thalamus; CTRL, control. Each dot represents data from one animal; bar represents the mean ± s.d. of n = 3 animals.
Extended Data Fig. 4 Detailed protocol for ωRNA cloning.
Step 1: Digest and purify the ωRNA acceptor plasmid with BbsI. Step 2: Perform ligation or Golden-Gate-Assembly of phosphorylated and annealed oligonucleotides into the digested pωRNA-acceptor.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–12 and Note 1.
Supplementary Data 1 (download XLSX )
Supplementary dataset with DNA sequences, indel/editing rates and features for ML.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Marquart, K.F., Mathis, N., Mollaysa, A. et al. Effective genome editing with an enhanced ISDra2 TnpB system and deep learning-predicted ωRNAs. Nat Methods 21, 2084–2093 (2024). https://doi.org/10.1038/s41592-024-02418-z
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-024-02418-z
This article is cited by
-
Harnessing artificial intelligence to advance CRISPR-based genome editing technologies
Nature Reviews Genetics (2026)
-
Coiled-coil heterodimer-mediated split base editing systems enable flexible and robust nucleotide substitutions
Nature Communications (2026)
-
Engineering eukaryotic transposon-encoded Fanzor2 system for genome editing in mammals
Nature Chemical Biology (2026)
-
Molecular effects of transposable element sequences in mammalian cells
Genome Biology (2025)
-
Therapeutic in vivo genome editing: innovations and challenges in rAAV vector-based CRISPR delivery
Gene Therapy (2025)