Abstract
Neurons are best studied in their native states in which their functional and morphological dynamics support animals’ natural behaviors. Super-resolution microscopy can potentially reveal these dynamics in higher details but has been challenging in behaving animals due to severe motion artifacts. Here we report multiplexed, line-scanning, structured illumination microscopy, which can tolerate motion of up to 50 μm s−1 while achieving 150-nm and 100-nm lateral resolutions in its linear and nonlinear forms, respectively. We continuously imaged the dynamics of spinules in dendritic spines and axonal boutons volumetrically over thousands of frames and tens of minutes in head-fixed mouse brains during sleep–wake cycles. Super-resolution imaging of axonal boutons revealed spinule dynamics on a scale of seconds. Simultaneous two-color imaging further enabled analyses of the spatial distributions of diverse PSD-95 clusters and opened up possibilities to study their correlations with the structural dynamics of dendrites in the brains of head-fixed awake mice.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
All data are available from the corresponding author on reasonable request. Due to the substantial size of the image data (~4 TB), it is not currently feasible to deposit it into a central repository. Interested individuals can request all datasets directly from the corresponding author. Source data are provided with this paper.
Code availability
The codes for the whole pipeline of the MLS-SIM are deposited to an online GitHub repository at https://github.com/WKLabION/MLS-SIM/ under an MIT license.
References
Debanne, D. Information processing in the axon. Nat. Rev. Neurosci. 5, 304–316 (2004).
Scott, E. K. & Luo, L. How do dendrites take their shape? Nat. Neurosci. 4, 359–365 (2001).
Holtmaat, A. & Svoboda, K. Experience-dependent structural synaptic plasticity in the mammalian brain. Nat. Rev. Neurosci. 10, 647–658 (2009).
Fu, M. & Zuo, Y. Experience-dependent structural plasticity in the cortex. Trends Neurosci. 34, 177–187 (2011).
Wefelmeyer, W., Puhl, C. J. & Burrone, J. Homeostatic plasticity of subcellular neuronal structures: from inputs to outputs. Trends Neurosci. 39, 656–667 (2016).
Lendvai, B., Stern, E. A., Chen, B. & Svoboda, K. Experience-dependent plasticity of dendritic spines in the developing rat barrel cortex in vivo. Nature 404, 876–881 (2000).
Nägerl, U. V., Willig, K. I., Hein, B., Hell, S. W. & Bonhoeffer, T. Live-cell imaging of dendritic spines by STED microscopy. Proc. Natl Acad. Sci. USA 105, 18982–18987 (2008).
Berning, S., Willig, K. I., Steffens, H., Dibaj, P. & Hell, S. W. Nanoscopy in a living mouse brain. Science 335, 551–551 (2012).
Tønnesen, J., Katona, G., Rózsa, B. & Nägerl, U. V. Spine neck plasticity regulates compartmentalization of synapses. Nat. Neurosci. 17, 678–685 (2014).
Tønnesen, J., Inavalli, V. V. G. K. & Nägerl, U. V. Super-resolution imaging of the extracellular space in living brain tissue. Cell 172, 1108–1121 (2018).
Arizono, M., Idziak, A., Quici, F. & Nägerl, U. V. Getting sharper: the brain under the spotlight of super-resolution microscopy. Trends Cell Biol. 33, 148–161 (2023).
Sigal, Y. M., Speer, C. M., Babcock, H. P. & Zhuang, X. Mapping synaptic input fields of neurons with super-resolution imaging. Cell 163, 493–505 (2015).
Jacquemet, G., Carisey, A. F., Hamidi, H., Henriques, R. & Leterrier, C. The cell biologist’s guide to super-resolution microscopy. J. Cell Sci. 133, jcs240713 (2020).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Xu, K., Zhong, G. & Zhuang, X. Actin, spectrin, and associated proteins form a periodic cytoskeletal structure in axons. Science 339, 452–456 (2013).
Wu, Y. & Shroff, H. Faster, sharper, and deeper: structured illumination microscopy for biological imaging. Nat. Methods 15, 1011–1019 (2018).
Dombeck, D. A., Khabbaz, A. N., Collman, F., Adelman, T. L. & Tank, D. W. Imaging large-scale neural activity with cellular resolution in awake, mobile mice. Neuron 56, 43–57 (2007).
Griffiths, V. A. et al. Real-time 3D movement correction for two-photon imaging in behaving animals. Nat. Methods 17, 741–748 (2020).
Masch, J.-M. et al. Robust nanoscopy of a synaptic protein in living mice by organic-fluorophore labeling. Proc. Natl Acad. Sci. USA 115, E8047–E8056 (2018).
Wegner, W., Mott, A. C., Grant, S. G. N., Steffens, H. & Willig, K. I. In vivo STED microscopy visualizes PSD95 sub-structures and morphological changes over several hours in the mouse visual cortex. Sci. Rep. 8, 219 (2018).
Wegner, W., Steffens, H., Gregor, C., Wolf, F. & Willig, K. I. Environmental enrichment enhances patterning and remodeling of synaptic nanoarchitecture as revealed by STED nanoscopy. eLife 11, e73603 (2022).
Velasco, M. G. M. et al. 3D super-resolution deep-tissue imaging in living mice. Optica 8, 442–450 (2021).
Turcotte, R. et al. Dynamic super-resolution structured illumination imaging in the living brain. Proc. Natl Acad. Sci. USA 116, 9586–9591 (2019).
Heintzmann, R., Jovin, T. M. & Cremer, C. Saturated patterned excitation microscopy—a concept for optical resolution improvement. J. Opt. Soc. Am. A 19, 1599–1609 (2002).
Gustafsson, M. G. L. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl Acad. Sci. USA 102, 13081–13086 (2005).
Gustafsson, M. G. L. et al. Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970 (2008).
Mandula, O. et al. Line scan—structured illumination microscopy super-resolution imaging in thick fluorescent samples. Opt. Express 20, 24167–24174 (2012).
Müller, C. B. & Enderlein, J. Image scanning microscopy. Phys. Rev. Lett. 104, 198101 (2010).
Sheppard, C. J. R., Mehta, S. B. & Heintzmann, R. Superresolution by image scanning microscopy using pixel reassignment. Opt. Lett. 38, 2889–2892 (2013).
Mudry, E. et al. Structured illumination microscopy using unknown speckle patterns. Nat. Photon. 6, 312–315 (2012).
Schropp, M., Seebacher, C. & Uhl, R. XL-SIM: extending superresolution into deeper layers. Photonics 4, 33 (2017).
Hering, H. & Sheng, M. Dentritic spines: structure, dynamics and regulation. Nat. Rev. Neurosci. 2, 880–888 (2001).
Kasai, H., Ziv, N. E., Okazaki, H., Yagishita, S. & Toyoizumi, T. Spine dynamics in the brain, mental disorders and artificial neural networks. Nat. Rev. Neurosci. 22, 407–422 (2021).
Westrum, L. E. & Blackstad, T. W. An electron microscopic study of the stratum radiatum of the rat hippocampus (regio superior, CA 1) with particular emphasis on synaptology. J. Comp. Neurol. 119, 281–309 (1962).
Petralia, R. S., Wang, Y.-X., Mattson, M. P. & Yao, P. J. Structure, distribution, and function of neuronal/synaptic spinules and related invaginating projections. NeuroMolecular Med. 17, 211–240 (2015).
Zaccard, C. R. et al. Rapid 3D enhanced resolution microscopy reveals diversity in dendritic spinule dynamics, regulation, and function. Neuron 107, 522–537 (2020).
Qiao, Q. et al. Long-term stability of axonal boutons in the mouse barrel cortex. Dev. Neurobiol. 76, 252–261 (2016).
Tononi, G. & Cirelli, C. Sleep and the price of plasticity: from synaptic and cellular homeostasis to memory consolidation and integration. Neuron 81, 12–34 (2014).
Walker, M. P. & Stickgold, R. Sleep, memory, and plasticity. Annu. Rev. Psychol. 57, 139–166 (2006).
de Vivo, L. et al. Ultrastructural evidence for synaptic scaling across the wake/sleep cycle. Science 355, 507–510 (2017).
Yang, G. et al. Sleep promotes branch-specific formation of dendritic spines after learning. Science 344, 1173–1178 (2014).
Li, W., Ma, L., Yang, G. & Gan, W.-B. REM sleep selectively prunes and maintains new synapses in development and learning. Nat. Neurosci. 20, 427–437 (2017).
Maret, S., Faraguna, U., Nelson, A. B., Cirelli, C. & Tononi, G. Sleep and waking modulate spine turnover in the adolescent mouse cortex. Nat. Neurosci. 14, 1418–1420 (2011).
Kashiwagi, Y. et al. Computational geometry analysis of dendritic spines by structured illumination microscopy. Nat. Commun. 10, 1285 (2019).
Nair, D. et al. Super-resolution imaging reveals that AMPA receptors inside synapses are dynamically organized in nanodomains regulated by PSD95. J. Neurosci. 33, 13204–13224 (2013).
Gross, G. G. et al. Recombinant probes for visualizing endogenous synaptic proteins in living neurons. Neuron 78, 971–985 (2013).
Marrs, G. S., Green, S. H. & Dailey, M. E. Rapid formation and remodeling of postsynaptic densities in developing dendrites. Nat. Neurosci. 4, 1006–1013 (2001).
Weigert, M. et al. Content-aware image restoration: pushing the limits of fluorescence microscopy. Nat. Methods 15, 1090–1097 (2018).
Wu, Y. et al. Multiview confocal super-resolution microscopy. Nature 600, 279–284 (2021).
Hofmann, M., Eggeling, C., Jakobs, S. & Hell, S. W. Breaking the diffraction barrier in fluorescence microscopy at low light intensities by using reversibly photoswitchable proteins. Proc. Natl Acad. Sci. USA 102, 17565–17569 (2005).
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Rego, E. H. et al. Nonlinear structured-illumination microscopy with a photoswitchable protein reveals cellular structures at 50-nm resolution. Proc. Natl Acad. Sci. USA 109, E135–E143 (2012).
Li, D. et al. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science 349, aab3500 (2015).
Chen, Y.-H., Jin, S.-Y., Yang, J.-M. & Gao, T.-M. The memory orchestra: contribution of astrocytes. Neurosci. Bull. 39, 409–424 (2023).
Novotny, L. & Hecht, B. Principles of Nano-Optics (Cambridge Univ. Press, 2006).
Nakabayashi, T. et al. pH dependence of the fluorescence lifetime of enhanced yellow fluorescent protein in solution and cells. J. Photochem. Photobiol. Chem. 235, 65–71 (2012).
Lister, J. A., Robertson, C. P., Lepage, T., Johnson, S. L. & Raible, D. W. nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 126, 3757–3767 (1999).
Namikawa, K. et al. Modeling neurodegenerative spinocerebellar ataxia type 13 in zebrafish using a purkinje neuron specific tunable coexpression system. J. Neurosci. 39, 3948–3969 (2019).
Balciunas, D. et al. Harnessing a high cargo-capacity transposon for genetic applications in vertebrates. PLoS Genet. 2, e169 (2006).
Bensussen, S. et al. A viral toolbox of genetically encoded fluorescent synaptic tags. iScience 23, 101330 (2020).
Barger, Z., Frye, C. G., Liu, D., Dan, Y. & Bouchard, K. E. Robust, automated sleep scoring by a compact neural network with distributional shift correction. PLoS ONE 14, e0224642 (2019).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Tsai, J., Grutzendler, J., Duff, K. & Gan, W.-B. Fibrillar amyloid deposition leads to local synaptic abnormalities and breakage of neuronal branches. Nat. Neurosci. 7, 1181–1183 (2004).
Acknowledgements
We thank L. Shao for offering insightful critiques of our paper. We thank J. W. Zhou, H. T. Xu, Z. Y. Liu, D. Q. Liu, J. He and Y. Mu for valuable discussions on neural imaging and data analysis. We thank J. Cao for mouse care. We thank L. Cong, P. Yu and Z. Q. Shi for suggestions on data analysis. We thank H. T. Cao and Y. J. Xiu for suggestions on data analysis on sleep states. We thank T. L. Chen for initial experiments on zebrafish. This research was supported by STI2030-Major Projects (2021ZD0204503 to K.W.; 2021ZD0204502 to J.D.; 2022ZD0206100 to Z.Z.), National Natural Science Foundation of China (32125020 and 32321003 to K.W.; 82271269 to L.B.; U23A20433 to Z.Z.), Shanghai Committee of Science and Technology, China (22JC1403101 to K.W.), National Key R&D Program of China (2018YFA0801000 and 2018YFA0801001 to X.D.) and China Postdoctoral Science Foundation (2023M743609 and BX20240389 to L.B.).
Author information
Authors and Affiliations
Contributions
K.W. conceptualized the project. Y. Zhang built the MLS-SIM and wrote the code for image reconstructions and simulations. Y. Zhang and X.W. performed zebrafish-related experiments under the supervision of X.D. and J.D. Y. Zhang performed mouse-related experiments. Y. Zhang performed EEG and EMG recording and data analysis on sleep states of mice under the supervision of Z.Z. Y. Zhang, L.B., Y. Zhao, T.Z. and L.Y. contributed to data analysis and discussion. K.W. and Y. Zhang wrote the paper with inputs from all authors. K.W. and J.D. supervised the project.
Corresponding author
Ethics declarations
Competing interests
K.W., Y. Zhang and L.Y. are listed as inventors on the Chinese patent ‘Super-resolution imaging method and apparatus based on line scanning’ (no. 202310207189.0 registered on 6 March 2023; PCT no. PCT/CN2023/123332). The other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Thomas Huser, Valentin Nägerl and Jan Tonnesen for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nina Vogt, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental implementation of MLS-SIM.
a, Schematics of the optical system of MLS-SIM. DM, dichroic mirror; PBS, polarizing beam splitter; AOTF, acousto-optic tunable filter; RPG, rotation phase grating; PD, photodetector; TF, transmission filter; SM, scanning mirror; PM, phase mask; SF, spatial filter. PM and SF are absent in linear MLS-SIM imaging and present only in nonlinear MLS-SIM imaging. b, Schematics of device synchronization. During the capture of a single frame, the camera is exposed multiple times and each exposure corresponds to one line in the resulting imaging frame. Two acousto-optic tunable filters (AOTF1 and AOTF2 in a) switch two different patterns on and off alternatively. In two-color imaging, two different colors are interleaved before interleaving patterns to ensure co-alignment between the two colors. Y scanning mirror (SM in a) performs a single continuous scan in the y direction in each frame. At the same time, the phase grating (RPG in a) is continuously rotated to scan the second excitation pattern along the x direction. Due to the instability of the employed rotation stage, the rotational speed of the phase grating is monitored by a photodetector (PD in a) for phase estimation of the second illumination pattern in image reconstruction. c, Phase modulation mask for structured line illumination generation. Left, schematics of the circularly distributed grating (RPG in a). The blue line on the left part of the grating indicates the location where the line excitation laser is focused on. The green line on the magnified view of the grating indicates the location of the cross-section view shown at the bottom. It is a stepped transmission phase grating fabricated on the fused silica wafer. Right, a photo of the customized phase grating. The bottom right is a magnified photo taken on a differential interference contrast (DIC) microscope. d, Experimental implementation of MLS-SIM. Top, overview of the optical system. The yellow line indicates the light path of the excitation beam. Bottom, a magnified view of the multiplexing of two excitation patterns. The red line indicates the light path of the 635 nm laser for rotation phase grating monitoring. All labels of optical elements are the same as their counterparts described in a.
Extended Data Fig. 2 Schematics of image acquisition and reconstruction in MLS-SIM.
a, Simplified schematics of consecutive 10 camera snapshots (Snapshot1–Snapshot10) captured under alternative excitations of two different patterns during scanning in y direction at 10 different locations (y1-y10). Each camera snapshot has two rows and each row has 10 imaging pixels. To avoid confusion, imaging pixels are termed as detectors hereafter. The detector measurement M(i,j,k) can be indexed by the column number (i), row number (j) and snapshot number (k). Depending on the spatial relationships between excitation patterns and detectors, the detector measurements are divided into eight different groups. Each detector measurement belongs to a certain group indicated by the index on the pixel in the schematics. b, Schematics of super-resolution image reconstruction. Each group of detector measurements is captured under the same excitation pattern from the perspectives of these detectors. They share the same effective PSF (PSFEF), which depends on the spatial relationship between excitation PSF (PSFExt) and detection PSF (PSFDet). These detector measurements can be stitched into 2D images (CFImg) consisting of 10 rows by 10 columns of pixels. Not all pixels in the 2D array are provided as excitation patterns are interleaved in time and space. These pre-processed images CFImg are up-sampled first along the x direction by filling in unsampled pixels. The up-sampled images USCFImg are then jointly deconvolved using a customized algorithm derived from Richardson-Lucy deconvolution. Finally, pixel binning along the y axis is performed on the deconvolved super-resolution image to make the final image have the same scales along the x and y directions.
Extended Data Fig. 3 Numerical simulations of patterned line excitations.
Numerical simulations of line excitation patterns employed in linear and nonlinear MLS-SIM and their comparison with conventional diffraction-limited focus spot. First row, schematics of different patterns of excitations on the imaging objective’s pupil plane. Second row, 3D views of simulated intensity distributions of different excitation patterns near the focal plane. Third row, comparisons of intensity profiles at indicated locations in the excitation patterns. In the simulation, the wavelength of the excitation laser is 488 nm, and the numerical aperture of the objective is 1.15. Polarizations are indicated by “E” and full vector model was employed in the simulation55. Among all three patterns, line patterns 1 and 2 have the tightest focuses in the y and x directions, respectively. Line pattern 2 has the tightest focus in the z direction. The effective excitation profiles of y4 and x5 in nonlinear excitation conditions assume a fluorescence emission rate25 of 0.9.
Extended Data Fig. 4 Numerical simulations of resolution enhancement by MLS-SIM.
Comparison of excitation patterns, representative effective PSFs, and their OTFs of two patterned illuminations and wide-field illumination. White crosses “+” in the images of the first column indicate the position of the representative detector that collect images with the effective PSF and OTF shown in the seconds and third columns. Vertical and horizontal linecut profiles at center are shown for effective PSFs of pattern 1 and pattern 2, respectively. Horizontal and vertical linecut profiles at center are shown for effective OTFs of pattern 1 and pattern 2, respectively. All images are displayed with a linear scale colormap.
Extended Data Fig. 5 Multi-layer reconstruction in MLS-SIM.
a, Diversities of detector groups and their corresponding effective PSFs employed in simulations. Ten images were collected with 10 different PSFs (Supplementary Note 2). Top, 10 different spatial relationships between excitation patterns and detectors on the camera generate 10 different effective PSFs. Middle, effective PSFs at different depths from the focal plane. Bottom, the corresponding optical transfer functions (OTFs) of these effective PSFs. b, Simulations on noise amplification in multi-layer reconstruction. Based on a multi-layer reconstruction model in spatial frequency space, the reliability of reconstruction could be estimated by the noise amplification factors. The noise amplification factors in the spatial frequency space of two reconstructed layers in two configurations are presented on the left and right, respectively. Dashed circles indicate spatial frequencies supporting indicated lateral resolutions. c, Representative experimental super-resolution reconstruction using single-layer and two different multi-layer reconstruction strategies. Multi-layer reconstructions yielded images with reduced out-of-focus backgrounds. The performance of multi-layer reconstructions depended on the choices of reconstructed layers. Three-layer reconstruction at z = 0 and z = ±150 nm (middle) produce more reconstruction artifacts than the reconstruction at layers z = 0 and z = ±450 nm (right).
Extended Data Fig. 6 Characterization of axial resolution of MLS-SIM.
a, Representative confocal and MLS-SIM images of two-layer 100 nm diameter fluorescent beads. Depth information is color coded. b, Representative axial cross-section view at the position indicated by dotted lines in a. Yellow arrows indicate vertically spaced beads. White arrow indicates a single layer of beads. c, Intensity line profiles of reconstructed image at locations indicated by yellow arrows in b. Line profiles were fitted by two overlapping Gaussian functions to estimate the size of resolvable features. The FWHM of the fitted Gaussian functions and their separations are specified. d, Intensity line profiles of confocal (blue) and MLS-SIM (red) images at locations indicated by white arrows in b. Line profiles were fitted by Gaussian function to estimate the FWHM of the reconstructed feature. e, Representative image of neurons in the brain of anesthetized mouse. Depth information is color coded. f, Representative cross-section views of white boxes in e. The fitted axial separations between crossing axons are specified.
Extended Data Fig. 7 Characterization of achievable resolution under different levels of SNR.
a, Representative raw and reconstructed images of fluorescent beads captured at varying SNRs. Scale bar, 400 nm. b, Histograms of the FWHMs measured from confocal images of isolated 100 nm diameter fluorescent beads. c, Probability of successfully resolving bead-pairs with different inter-spaces under varying levels of SNR (n = 5 beads pairs for each condition, error bars indicate mean ± s.e.m.). d, FWHM of reconstructed beads in the axial and lateral directions at different levels of SNR (n = 10 beads for each condition, error bars indicate mean ± s.e.m.). e, Representative confocal and reconstructed MLS-SIM images of densely packed 100 nm diameter fluorescent beads (Methods). f, Magnified images of the indicated ROIs in e.
Extended Data Fig. 8 Achievable resolution under different levels of sample motions.
a, Resolution characterization of MLS-SIM on static fluorescent beads with diameters of 100 nm. b, Reconstructions of the same group of representative fluorescent beads moving at three different speeds (25, 50, and 100 micrometers per second) and in eight different directions. c, Statistical quantifications of experimentally obtained resolutions by resolving bead pairs with different orientations. A cubic spline function with 4 knots (at 0, 45, 90, and 135°) and periodic ending conditions was used to fit the measured interspaces at different angles.
Extended Data Fig. 9 Characterization of resolution enhancement of nonlinear MLS-SIM.
a, Representative images of 100 nm diameter beads imaged by nonlinear MLS-SIM with low (0.1 nJ pulse energy, corresponding to a fluorescence emission rate of 0.1) and high (1.2 nJ pulse energy, corresponding to a fluorescence emission rate of 0.9) saturation levels. b, Intensity line profiles at locations indicated by white arrows in a. Line profiles were fitted by Gaussian function to estimate the sizes of features. The FWHM of the fitted Gaussian functions are specified. c, Intensity line profiles at locations indicated by yellow arrows in a. Line profiles were fitted by two overlapping Gaussian functions to estimate the sizes and separation of features. The FWHM of the fitted Gaussian functions and their separations are specified. d, Representative images of neurons expressing membrane-targeted EGFP in mice brain imaged by nonlinear MLS-SIM and linear MLS-SIM. Images were displayed at the same scale for comparison.
Extended Data Fig. 10 Characterization of morphological dynamics of axonal boutons in mouse brains.
a-c, Overview of axons of somatostatin interneurons expressing membrane-targeted EGFP. Magnified and time-lapsed views of indicated regions in a are shown in b for the first day and in c for across 5 consecutive days. Gamma changes with a factor of 0.8 is applied to the image to enhance visualization. d, The number of days with observation of spinule dynamics on boutons. Experimental observation (blue) deviated significantly from shuffled data (orange), which assumes a uniform distribution of spinule dynamics on all boutons (p = 8.85×10−62, n = 475 boutons from 7 FOVs in 3 mice, one-sided Chi-square goodness of fit test). e, Relationship between the bouton’s area and its long term spinule dynamics. Red line is a linear fit of the raw data (circles). R and p values for Pearson correlation between two factors are indicated in the plot (p = 3.35×10−6, Pearson correlation, two-sided test for correlation, n = 535, 7 FOVs in 3 mice). f, Distributions of dynamic area changes over boutons (n = 535, 7 FOVs in 3 mice). The area fluctuation is defined as the standard deviation of a bouton’s area change normalized to its average area across 5 days. Scale bars: 10 µm in a; 1 µm in b and c.
Supplementary information
Supplementary Information (download PDF )
Supplementary Figs. 1–14, Table 1, and Notes 1 and 2.
Supplementary Software (download ZIP )
Software for image acquisition and processing in MLS-SIM.
Supplementary Video 1 (download MP4 )
MLS-SIM imaging in living zebrafish brain. Imaging of Purkinje cells expressing membrane-targeted EGFP in living larval zebrafish using MLS-SIM. Inset provides a magnified view.
Supplementary Video 2 (download MP4 )
MLS-SIM imaging in awake mouse brain. Imaging of neurons expressing membrane-targeted EGFP in awake mouse brain. LC, raw line confocal imaging; LCDeconv, line confocal imaging with deconvolution. Left, overview of the entire FOV; right, magnified views of representative ROIs processed as LC (first column), LCDeconv (second column) and MLS-SIM (third column). Moving bars in white boxes indicate the imaging depth of current views.
Supplementary Video 3 (download MP4 )
Comparison of MLS-SIM and two-photon imaging in the mouse brain. Visualization of a sparse population of neurons expressing membrane-targeted EGFP in the living mouse brain, imaged using TPM and MLS-SIM. In the MLS-SIM imaging display, each frame is a maximum intensity projection (MIP) of five reconstructed frames to enhance the depth of view for better comparison with TPM.
Supplementary Video 4 (download MP4 )
MLS-SIM imaging and motion correction in awake mouse brain. Example of motion distortion and correction of motions by image registrations. Fine structures under severe distortions could be resolved and tracked over time after motion correction.
Supplementary Video 5 (download MP4 )
Time-lapse volumetric imaging in awake mouse brain. Bottom, magnified views. Imaging volumes, consisting of 20 planes, were continuously captured at 5.0 frames per second and over 20 min.
Supplementary Video 6 (download MP4 )
Time-lapse volumetric imaging of spinules in awake mouse brain. Bottom, magnified views of spines with identified spinule emergence. Blink was caused by failures in image registration when animal motion was severe.
Supplementary Video 7 (download MP4 )
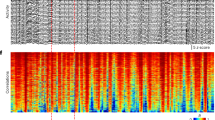
Time-lapse volumetric imaging in mouse brain during sleep–wake cycles. Top, brain states inferred from EEG and EMG recording. Gray, wake; yellow, NREM sleep; blue, REM sleep. Dark-red bar shows the recording time of the current imaging volume.
Supplementary Video 8 (download MP4 )
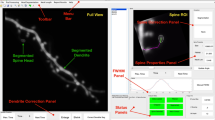
Two-color time-lapse imaging of PSD-95 and membrane of neurons in awake mouse brain. First part, scanning through an imaging stack. Second part, MIP views of two example imaging stacks during time-lapse imaging. Gray, membrane-targeted mRuby3; red, PSD95.FingR-EGFP. Right, magnified view of representative structures in the volume.
Supplementary Video 9 (download MP4 )
Schematics for image acquisition and super-resolution image reconstruction in MLS-SIM.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data for Fig. 1.
Source Data Fig. 2 (download XLSX )
Statistical source data for Fig. 2.
Source Data Fig. 3 (download XLSX )
Statistical source data for Fig. 3.
Source Data Fig. 4 (download XLSX )
Statistical source data for Fig. 4.
Source Data Fig. 5 (download XLSX )
Statistical source data for Fig. 5.
Source Data Extended Data Fig. 3 (download XLSX )
Simulation source data for Extended Data Fig. 3.
Source Data Extended Data Fig. 4 (download XLSX )
Simulation source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data for Extended Data Fig. 6.
Source Data Extended Data Fig. 7 (download XLSX )
Statistical source data for Extended Data Fig. 7.
Source Data Extended Data Fig. 8 (download XLSX )
Statistical source data for Extended Data Fig. 8.
Source Data Extended Data Fig. 9 (download XLSX )
Statistical source data for Extended Data Fig. 9.
Source Data Extended Data Fig. 10 (download XLSX )
Statistical source data for Extended Data Fig. 10.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, Y., Bai, L., Wang, X. et al. Super-resolution imaging of fast morphological dynamics of neurons in behaving animals. Nat Methods 22, 177–186 (2025). https://doi.org/10.1038/s41592-024-02535-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-024-02535-9