Abstract
Most biological processes, from development to pathogenesis, span multiple time and length scales. While light-sheet fluorescence microscopy has become a fast and efficient method for imaging organisms, cells and subcellular dynamics, simultaneous observations across all these scales have remained challenging. Moreover, continuous high-resolution imaging inside living organisms has mostly been limited to a few hours, as regions of interest quickly move out of view due to sample movement and growth. Here, we present a self-driving, multiresolution light-sheet microscope platform controlled by custom Python-based software, to simultaneously observe and quantify subcellular dynamics in the context of entire organisms in vitro and in vivo over hours of imaging. We apply the platform to the study of developmental processes, cancer invasion and metastasis, and we provide quantitative multiscale analysis of immune–cancer cell interactions in zebrafish xenografts.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
Test data for all our analysis code is available on Zenodo at https://zenodo.org/records/12791724 (ref. 89). All raw imaging data (several TB of data) are available on Synapse90,91, the official storage space of the MC2 center, supported by the National Institutes of Health: test data collection (same as Zenodo) at https://doi.org/10.7303/syn61795850.2 (ref. 91) and experiment datasets collection at https://doi.org/10.7303/syn61795837.2 (ref. 90). Source data are provided with this paper.
Code availability
All algorithms, code and software used in this study are available on GitHub. The microscope control code and all image processing software is available at https://github.com/DaetwylerStephan/self_driving_multiscale_control and https://github.com/DaetwylerStephan/multiscale-image-analysis.
References
Daetwyler, S. & Huisken, J. Fast fluorescence microscopy with light sheets. Biol. Bull. 231, 14–25 (2016).
Power, R. M. & Huisken, J. A guide to light-sheet fluorescence microscopy for multiscale imaging. Nat. Methods 14, 360–373 (2017).
Stelzer, E. H. K. et al. Light sheet fluorescence microscopy. Nat. Rev. Methods Primers 1, 73 (2021).
Chen, B.-C. et al. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346, 1257998 (2014).
Chang, B.-J. et al. Universal light-sheet generation with field synthesis. Nat. Methods https://doi.org/10.1038/s41592-019-0327-9 (2019).
Wu, Y. et al. Spatially isotropic four-dimensional imaging with dual-view plane illumination microscopy. Nat. Biotechnol. 31, 1032–1038 (2013).
Dean, K. M., Roudot, P., Welf, E. S., Danuser, G. & Fiolka, R. Deconvolution-free subcellular imaging with axially swept light sheet microscopy. Biophys. J. 108, 2807–2815 (2015).
Dean, K. M. et al. Isotropic imaging across spatial scales with axially swept light-sheet microscopy. Nat. Protoc. https://doi.org/10.1038/s41596-022-00706-6 (2022).
Huisken, J., Swoger, J., Del Bene, F., Wittbrodt, J. & Stelzer, E. H. K. Optical sectioning deep inside live embryos by selective plane illumination microscopy. Science 305, 1007–1009 (2004).
Daetwyler, S., Günther, U., Modes, C. D., Harrington, K. & Huisken, J. Multi-sample SPIM image acquisition, processing and analysis of vascular growth in zebrafish. Development https://doi.org/10.1242/dev.173757 (2019).
McDole, K. et al. In toto imaging and reconstruction of post-implantation mouse development at the single-cell level. Cell 175, 859–876.e33 (2018).
Glaser, A. K. et al. A hybrid open-top light-sheet microscope for versatile multi-scale imaging of cleared tissues. Nat. Methods 19, 613–619 (2022).
Daetwyler, S. & Fiolka, R. P. Light-sheets and smart microscopy, an exciting future is dawning. Commun. Biol. 6, 502 (2023).
Zhang, Y. & Gross, H. Systematic design of microscope objectives. Part I: system review and analysis. Adv. Opt. Technol. 8, 313–347 (2019).
Almada, P. et al. Automating multimodal microscopy with NanoJ-fluidics. Nat. Commun. 10, 1223 (2019).
Alvelid, J., Damenti, M., Sgattoni, C. & Testa, I. Event-triggered STED imaging. Nat. Methods 19, 1268–1275 (2022).
Shi, Y. et al. Smart lattice light-sheet microscopy for imaging rare and complex cellular events. Nat. Methods https://doi.org/10.1038/s41592-023-02126-0 (2024).
Huisken, J. & Stainier, D. Y. R. Even fluorescence excitation by multidirectional selective plane illumination microscopy (mSPIM). Opt. Lett. 32, 2608–2610 (2007).
Ershov, D. et al. TrackMate 7: integrating state-of-the-art segmentation algorithms into tracking pipelines. Nat. Methods 19, 829–832 (2022).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Royer, L. A. et al. Adaptive light-sheet microscopy for long-term, high-resolution imaging in living organisms. Nat. Biotechnol. 34, 1267–1278 (2016).
Chiron, L. et al. CyberSco.Py an open-source software for event-based, conditional microscopy. Sci. Rep. 12, 11579 (2022).
Voigt, F. F. et al. The mesoSPIM initiative: open-source light-sheet microscopes for imaging cleared tissue. Nat. Methods 16, 1105–1108 (2019).
Bradski, G. The OpenCV library. Dr Dobbs J. 120, 122–125 (2000).
Thayer, N. H. & York, A. G. GitHub Repository: Tools. https://github.com/AndrewGYork/tools (2024)
Ahlers, J. et al. napari: a multi-dimensional image viewer for Python. Zenodo https://doi.org/10.5281/zenodo.8115575 (2023).
Liu, T.-L. et al. Observing the cell in its native state: imaging subcellular dynamics in multicellular organisms. Science 360, eaaq1392 (2018).
Mondal, C. et al. A proliferative to invasive switch is mediated by srGAP1 downregulation through the activation of TGF-β2 signaling. Cell Rep. 40, 111358 (2022).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Westcott, J. M. et al. An epigenetically distinct breast cancer cell subpopulation promotes collective invasion. J. Clin. Invest. 125, 1927–1943 (2015).
Flanagan, L., Weelden, K. V., Ammerman, C., Ethier, S. P. & Welsh, J. SUM-159PT cells: a novel estrogen independent human breast cancer model system. Breast Cancer Res. Treat. 58, 193–204 (1999).
Lacroix, M. & Leclercq, G. Relevance of breast cancer cell lines as models for breast tumours: an update. Breast Cancer Res. Treat. 83, 249–289 (2004).
Chi, N. C. et al. Foxn4 directly regulates tbx2b expression and atrioventricular canal formation. Genes Dev. 22, 734–739 (2008).
Link, B. A. & Megason, S. G. in Sourcebook of Models for Biomedical Research (ed. Conn, P. M.) 103–112 (Humana Press, 2008).
White, R. M. et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2, 183–189 (2008).
Scheele, C. L. G. J. et al. Multiphoton intravital microscopy of rodents. Nat. Rev. Methods Primers 2, 89 (2022).
Pauls, S., Geldmacher-Voss, B. & Campos-Ortega, J. A. A zebrafish histone variant H2A.F/Z and a transgenic H2A.F/Z:GFP fusion protein for in vivo studies of embryonic development. Dev. Genes Evol. 211, 603–610 (2001).
Jin, S.-W., Beis, D., Mitchell, T., Chen, J.-N. & Stainier, D. Y. R. Cellular and molecular analyses of vascular tube and lumen formation in zebrafish. Development 132, 5199–5209 (2005).
Valastyan, S. & Weinberg, R. A. Tumor metastasis: molecular insights and evolving paradigms. Cell 147, 275–292 (2011).
Talmadge, J. E. & Fidler, I. J. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res. 70, 5649–5669 (2010).
Osmani, N. & Goetz, J. G. Multiscale imaging of metastasis in zebrafish. Trends Cancer 5, 766–778 (2019).
White, R., Rose, K. & Zon, L. Zebrafish cancer: the state of the art and the path forward. Nat. Rev. Cancer 13, 624–636 (2013).
Segal, D. et al. In vivo 3D profiling of site-specific human cancer cell morphotypes in zebrafish. J. Cell Biol. 221, e202109100 (2022).
Asokan, N. et al. Long-term in vivo imaging reveals tumor-specific dissemination and captures host tumor interaction in zebrafish xenografts. Sci. Rep. 10, 13254 (2020).
Cailleau, R., Young, R., Olivé, M. & Reeves, W. J. Jr. Breast tumor cell lines from pleural effusions. J. Natl Cancer Inst. 53, 661–674 (1974).
Belin, B. J., Goins, L. M. & Mullins, R. D. Comparative analysis of tools for live cell imaging of actin network architecture. BioArchitecture 4, 189–202 (2014).
Póvoa, V. et al. Innate immune evasion revealed in a colorectal zebrafish xenograft model. Nat. Commun. 12, 1156 (2021).
Ponten, J. & Saksela, E. Two established in vitro cell lines from human mesenchymal tumours. Int. J. Cancer 2, 434–447 (1967).
Subach, O. M. et al. A photoswitchable orange-to-far-red fluorescent protein, PSmOrange. Nat. Methods 8, 771–777 (2011).
Ellett, F., Pase, L., Hayman, J. W., Andrianopoulos, A. & Lieschke, G. J. mpeg1 promoter transgenes direct macrophage-lineage expression in zebrafish. Blood 117, e49–e56 (2011).
Giard, D. J. et al. In vitro cultivation of human tumors: establishment of cell lines derived from a series of solid tumors. J. Natl Cancer Inst. 51, 1417–1423 (1973).
Haase, R. et al. CLIJ: GPU-accelerated image processing for everyone. Nat. Methods 17, 5–6 (2020).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 9, 62–66 (1979).
Zhou, F. Y. et al. A general algorithm for consensus 3D cell segmentation from 2D segmented stacks. Preprint at bioRxiv https://doi.org/10.1101/2024.05.03.592249 (2024).
Stringer, C., Wang, T., Michaelos, M. & Pachitariu, M. Cellpose: a generalist algorithm for cellular segmentation. Nat. Methods 18, 100–106 (2021).
Hopkins, B. & Skellam, J. G. A new method for determining the type of distribution of plant individuals. Ann. Bot. 18, 213–227 (1954).
Lawson, R. G. & Jurs, P. C. New index for clustering tendency and its application to chemical problems. J. Chem. Inf. Comput. Sci. 30, 36–41 (1990).
Rousseeuw, P. J., Ruts, I. & Tukey, J. W. The bagplot: a bivariate boxplot. Am. Stat. 53, 382–387 (1999).
Zhang, Z. et al. Multi-scale light-sheet fluorescence microscopy for fast whole brain imaging. Front. Neuroanat. https://doi.org/10.3389/fnana.2021.732464 (2021).
S R, R. et al. Simultaneous multiple-level magnification selective plane illumination microscopy (sMx-SPIM) imaging system. J. Opt. 24, 024010 (2022).
Icha, J., Weber, M., Waters, J. C. & Norden, C. Phototoxicity in live fluorescence microscopy, and how to avoid it. BioEssays 39, 1700003 (2017).
Reynaud, E. G., Kržič, U., Greger, K. & Stelzer, E. H. K. Light sheet‐based fluorescence microscopy: more dimensions, more photons, and less photodamage. HFSP J. 2, 266–275 (2008).
Nazib, A., Fookes, C. & Perrin, D. A comparative analysis of registration tools: traditional vs deep learning approach on high resolution tissue cleared data. Preprint at https://arxiv.org/abs/1810.08315 (2018).
Maška, M. et al. The cell tracking challenge: 10 years of objective benchmarking. Nat. Methods 20, 1010–1020 (2023).
Shi, W. et al. Curvature sensing as an emergent property of multiscale assembly of septins. Proc. Natl Acad. Sci. USA 120, e2208253120 (2023).
Weems, A. D. et al. Blebs promote cell survival by assembling oncogenic signalling hubs. Nature 615, 517–525 (2023).
Yuan, J. et al. Osteoblastic and osteolytic human osteosarcomas can be studied with a new xenograft mouse model producing spontaneous metastases. Cancer Invest. 27, 435–442 (2009).
Rozenberg, G. I., Monahan, K. B., Torrice, C., Bear, J. E. & Sharpless, N. E. Metastasis in an orthotopic murine model of melanoma is independent of RAS/RAF mutation. Melanoma Res. https://doi.org/10.1097/cmr.0b013e328336ee17 (2010).
Banerjee, S. et al. The Vps13-like protein BLTP2 is pro-survival and regulates phosphatidylethanolamine levels in the plasma membrane to maintain its fluidity and function. Preprint at bioRxiv https://doi.org/10.1101/2024.02.04.578804 (2024).
Daetwyler, S. et al. Mesoscopic oblique plane microscopy via light-sheet mirroring. Optica 10, 1571–1581 (2023).
Sapoznik, E. et al. A versatile oblique plane microscope for large-scale and high-resolution imaging of subcellular dynamics. eLife 9, e57681 (2020).
Matthews, C. & Cordelieres, F. P. MetroloJ: an ImageJ plugin to help monitor microscopes’ health. In ImageJ User & Developer Conference Proc. 1–6 (2010).
Dahm, R. & Nüsslein-Volhard, C. Zebrafish: A Practical Approach (Oxford University Press, 2002).
Westerfield, M. The Zebrafish Book: A Guide for the Laboratory Use of Zebrafish (Danio rerio) 4th edn (Univ. Oregon Press, 2000).
Kaufmann, A., Mickoleit, M., Weber, M. & Huisken, J. Multilayer mounting enables long-term imaging of zebrafish development in a light sheet microscope. Development 139, 3242–3247 (2012).
Cross, G. R. & Jain, A. K. Measurement of clustering tendency. In Theory and Application of Digital Control. Proc. IFAC Symposium (ed. Mahalanabis, A. K.) 315–320 (Pergamon, 1982).
Wright, K. Will the real Hopkins statistic please stand up? R J. 14, 282–292 (2022).
Liao, P.-S., Chen, T.-S. & Chung, P.-C. A fast algorithm for multilevel thresholding. J. Inf. Sci. Eng. 17, 713–727 (2001).
van der Walt, S. et al. scikit-image: image processing in Python. PeerJ 2, e453 (2014).
Thevenaz, P., Ruttimann, U. E. & Unser, M. A pyramid approach to subpixel registration based on intensity. IEEE Trans. Image Process. 7, 27–41 (1998).
Semechko, A. Exact minimum bounding spheres and circles. GitHub https://github.com/AntonSemechko/Bounding-Spheres-And-Circles (2019).
Ritter, J. in Graphics Gems (ed. Glassner, A. S.) 301–303 (Academic Press Professional, 1990).
G. Taubin. Curve and surface smoothing without shrinkage. In Proc. IEEE International Conference on Computer Vision. 852–857 (ICCV, 1995). doi: 10.1109/ICCV.1995.466848.
S. Rusinkiewicz. Estimating curvatures and their derivatives on triangle meshes. In Proc. 2nd International Symposium on 3D Data Processing, Visualization and Transmission, 2004. 486–493 (IEEE, 2004).
Ben Shabat, Y. & Fischer, A. Design of porous micro-structures using curvature analysis for additive-manufacturing. Procedia CIRP 36, 279–284 (2015).
Driscoll, M. K. et al. Robust and automated detection of subcellular morphological motifs in 3D microscopy images. Nat. Methods 16, 1037–1044 (2019).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Daetwyler, S., Modes, C. D. & Fiolka, R. Fiji plugin for annotating movies with custom arrows. Biol. Open 9, bio056200 (2020).
Daetwyler, S. et al. Test data for ‘Imaging of cellular dynamics in vitro and in situ: from a whole organism to sub-cellular imaging with self-driving, multi-scale microscopy’. Zenodo https://doi.org/10.5281/zenodo.12791724 (2024).
Daetwyler, S. & Fiolka, R. P. CA268072-msls-6-2024-experiments. Synapse https://doi.org/10.7303/syn61795837.2 (2024).
Daetwyler, S. & Fiolka, R. P. CA268072-msls-6-2024-code-example-datasets. Synapse https://doi.org/10.7303/syn61795850.2 (2024).
Acknowledgements
We thank A. York for discussion on the control software, B.-J. Chang, J. Keth, D. K. Reed and K. Bhatt for support with reagents, sample preparations and sorting; the Animal Resource Center for taking care of the zebrafish facility; and the whole Fiolka, Dean, Amatruda and Danuser laboratories for providing feedback and comments. Moreover, this research was supported in part by the computational resources provided by the BioHPC initiative at UT Southwestern Medical Center. Funding for this work is acknowledged from the Swiss National Science Foundation, grant no. 191347 to S.D.; National Institute of General Medical Sciences, grant no. R35 GM133522 to R.F.; National Cancer Institute, grant no. U54 CA268072 to G.D. and R.F; and grant no. K99CA270285 to D.S.
Author information
Authors and Affiliations
Contributions
S.D. and R.F. wrote the paper; S.D., B.C. and R.F. built the microscope; S.D. programmed and applied the microscope; S.D., H.M.F. and F.Y.Z. performed data analysis; S.D. and E.S. performed sample preparations; J.M.W. and R.A.B. prepared cancer spheroids; D.S. and G.D. contributed to study design; S.D., R.F. and G.D. secured funding for the project; and all authors revised and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Tzung Hsiai and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Rita Strack, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Schematic of the multiscale microscope.
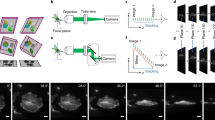
We introduce two designs for a multiscale microscope with dual-sided illumination (gray), a remote focusing unit (light gray) and detection with two resolutions (orange): a one design operating with cylindrical lenses and using a motorized slit, and b one design using Powell lenses, where no motorized slit is required (Supplementary Note 1). The microscope hardware comfortably accommodated several samples in one experiment, mounted in low melting agarose within fluorinated ethylene propylene (FEP) tubes. Abbreviations are as follows: 4 different lasers (LS1-LS4), dichroic beamsplitters (DC), 4x telescope (T1, f1 = 50 mm, f2 = 200 mm), pinhole (PI), halfwave plate (λ1/2), polarized beamsplitter cube (PBS), motorized flip mirror (FM), two cylindrical lenses (CT, f1 = 25 mm, 100 mm), vertical slit (VS), resonant galvanometer (RM), low-resolution cylindrical lens (Cl, f = 200 mm), tube lens (TLi, f = 100 mm), illumination objective (IL, NA 0.4), telescope (T2, 5x), motorized vertical slit (mVS), high-resolution cylindrical lens (Ch, f = 50 mm), tube lens (Trm, f = 200 mm), remote focusing objective (RF), a quarterwave plate (QWP), telescope (T3, f1 = 200 mm, f2 = 75 mm), 10° Powell lens (P), f = 30 mm lens (L1), f = 60 mm lens (L2), f = 400 mm lens (L3), voice coil with a mirror (VCM), LED illumination (LED), 20x NA1.0 detection objective (DL), filter wheel (FW), 100 mm tube lens (TL1), 500 mm tube lens (TL2).
Extended Data Fig. 2 Region tracking algorithm.
a Low-resolution maximum intensity projection of MDA-MB-231 breast cancer cells, labeled with F-tractin-GFP, in the zebrafish tail. Insets depict a high-resolution region of interest (blue, panel b) with its corresponding, enlarged (1.5x in both lateral dimensions) region used for better tracking. b 3D rendering of the high-resolution region of interest. c Schematic of the steps required for initialization of the region tracking algorithm. d X–Y, Y–Z, X–Z low-resolution maximum intensity projections saved in the image library at initialization, and each subsequent time point / update step. e Schematic of simultaneous data processing and acquisition (data streaming), enabled by a custom buffer architecture using shared memory arrays and tools from the concurrency tool library by Thayer, York et al. f Schematic of the pipeline for the update step to calculate on-the-fly the lateral and axial shifts for region tracking. Scale bar lengths are as follows: a 150 μm, b,d 50 μm.
Extended Data Fig. 3 Region tracking using transmission images.
a Selected maximum intensity projections of transmission images of the larval zebrafish tail over 11 hours of observation, starting at 2.5 days post-fertilization. The blue dashed line indicates the tip of the vasculature at the end of the tail, growing around 100 μm over the 11 hours window of observation (blue arrows). b Despite this growth, self-driving microscopy kept the vascular region of interest in focus over the observation window. Maximum intensity projections of the corresponding high-resolution region showcase details of the outgrowth of a single vessel (white arrowhead) with subsequent anastomosis (white arrows) with a neighboring vessel, including positioning of endothelial nuclei (asterisk). The vasculature was labeled with Tg(kdrl:EGFP). c Maximum intensity projection of the entire field of view of the low-resolution acquisition at start of the time-lapse imaging, with the inset highlighting the region used for tracking. d Representative X–Y, Y–Z, X–Z maximum intensity projections used in the region tracking algorithm. Scale bar lengths are as follows: a, c, d 150 μm; b 50 μm.
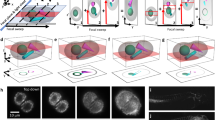
Extended Data Fig. 4 Phagocytosis of cancer cell by macrophages.
a Schematic of our in vivo zebrafish xenograft assay to study immune cell–cancer cell interactions in situ. Firstly, human cancer cells were cultured and modified as desired, for example, by expressing a fluorescent marker to label the cells. Secondly, cells were harvested and injected near the common cardinal vein (CCV) into the yolk of zebrafish larvae (violet arrow). Then, xenografts were imaged on our self-driving, multiresolution microscope, and subsequent analysis allowed visualization and quantification of cell spreading, cell–cell interactions, and cell morphological changes. b Low-resolution mSPIM images captured the distribution of macrophages (Tg(mpeg1:EGFP), top: magenta, bottom: grayscale image) in the entire zebrafish larvae after xenografting U-2 OS osteosarcoma cells (pVimentin-PsmOrange label, top: white, bottom: bright white). For tissue context, zebrafish also expressed the vascular marker Tg(kdrl:Hsa.HRAS-mCherry) (top: cyan). The image highlights how macrophages clustered around sites with cancer cells (Supplementary Movie 7). c In contrast, zebrafish without xenografts (control) displayed a uniform distribution of macrophages (Tg(mpeg1:EGFP), top: magenta, bottom: grayscale image) across the entire zebrafish embryo (top: vascular marker Tg(kdrl:Has.HRAS-mCherry in cyan). d The self-driving feature of the microscope enabled high-resolution imaging of selected cancer colonies in the zebrafish tail over many hours by keeping it in focus. Frequently, we observed how zebrafish macrophages (Tg(mpeg1:EGFP), top: magenta, bottom: gray) attached to the U-2 OS cancer cells (pVimentin-PsmOrange label, top and bottom: green), and phagocytosed them (Supplementary Movie 7, 8). Scale bar lengths are as follows: b,c 500 μm; d 50 μm.
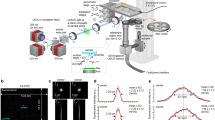
Extended Data Fig. 5 Workflow to obtain a high-resolution segmentation of zebrafish macrophages.
a Maximum intensity projection of a 3D volume in the caudal hematopoietic tissue visualizing zebrafish macrophages, labeled with Tg(mpeg1:EGFP)62, after xenografting human U-2 OS cells into the zebrafish larvae. As the cancer cells (yellow arrowhead) were labeled with a psmOrange fluorophore, there was bleed-through into the macrophage channel (GFP). Moreover, macrophages tended to cluster around cancer cells and at locations where cancer cells resided before they were phagocytosed by macrophages (yellow arrow). b 3D rendering with Fiji’s 3D viewer to display the final segmentation by the proposed workflow. c Schematic of the segmentation workflow. d We initially segmented the data with multi-Otsu thresholding and connected component labeling. This segmented individual macrophages well, for example in the control experiment (without xenografts) or at the beginning of the xenograft experiment. However, it could not separate touching macrophages (white arrowheads). e To clean up the segmentation, we first removed the cancer cell signal (white arrowhead) from the segmentation. f To separate clustering macrophages, we constructed 3D consensus segmentation from Cellpose-2D46 segmentations computed in orthogonal x–y, x–z, y–z views of the original raw data. While this segmentation separated clustering macrophages, it was observed to split individual macrophages into smaller fragments (white arrows) when the macrophages show extensive cytoplasmic extensions of several tens to hundreds of microns. g We merged the connected component segmentation and the Cellpose-based segmentation by replacing connected volumes that exceeded a defined threshold with the Cellpose-based segmentation. Moreover, we merged connected volumes that were below a certain threshold with neighboring volumes. Subsequent manual curation further cleaned up the segmentation of macrophages to obtain the finalized segmentation b. Scale bar lengths are as follows: a 30 μm.
Supplementary information
Supplementary Information (download PDF )
Supplementary Video captions 1–14, Notes 1–8, Tables 1–3, Figs. 1–21 and References.
Supplementary Video 1 (download AVI )
Multiscale, time-lapse imaging of a mosaic-labeled SUM159 breast cancer cell spheroid embedded into a collagen matrix with the low-resolution modality (top) and the high-resolution modality (bottom). The spheroids consisted of a 1:1 mixture of cells expressing the actin marker Lifeact-GFP (cyan) and Lifeact-mCherry (magenta). Boxed region on top indicates the location of the high-resolution region on the bottom. The yellow arrowhead points at a cell division at the invasive front. Maximum intensity projections of the time points are shown. Scale bars, 150 μm (top), 20 μm (bottom).
Supplementary Video 2 (download AVI )
3D rendering of multiscale, time-lapse imaging data of a mosaic-labeled SUM159 breast cancer cell spheroid embedded into a collagen matrix with the low-resolution modality (left) and the high-resolution modality (right). The spheroids consisted of a 1:1 mixture of cells expressing the actin marker Lifeact-GFP (cyan) and Lifeact-mCherry (magenta). Boxed region on left indicates the location of the high-resolution region on the right. Scale bars, 150 μm (left), 40 μm (right).
Supplementary Video 3 (download AVI )
Multiscale, time-lapse imaging of zebrafish gastrulation with cells expressing the histone marker Tg(h2afva:h2afva-GFP), starting at around 6 hours post-fertilization. Low-resolution imaging (top) captured the entire zebrafish embryo (color scale, depth of data in 3D volume from 0–450 μm), while the high-resolution imaging (bottom, maximum intensity projection) enabled near-simultaneous imaging of cell division. Scale bars, 250 μm (top), 25 μm (bottom).
Supplementary Video 4 (download AVI )
3D rendering of multiscale, time-lapse imaging data of zebrafish gastrulation with cells expressing the histone marker Tg(h2afva:h2afva-GFP), starting at around 6 h post-fertilization. Low-resolution imaging (left) captured the entire zebrafish embryo, while the high-resolution imaging (right) enabled near-simultaneous imaging of cell division. Scale bars, 250 μm (left), 25 μm (right).
Supplementary Video 5 (download AVI )
Multiscale, time-lapse imaging of human breast cancer cells MDA-MB-231 expressing F-tractin-EGFP (top, magenta; bottom, gray) in a larval zebrafish xenograft model, expressing the vascular marker Tg(kdrl:Hsa.HRAS-mCherry) for tissue context (top, cyan). Low-resolution imaging (top, maximum intensity projections) captured the whole organism, while high-resolution imaging (bottom, maximum intensity projections) revealed subcellular dynamics including an intricate network of protrusions. The cancer cells were xenografted into zebrafish larvae at 2.25 days post-fertilization. Scale bars, 250 μm (top), 25 μm (bottom).
Supplementary Video 6 (download AVI )
3D rendering of multiscale, time-lapse imaging data of human breast cancer cells MDA-MB-231 expressing F-tractin-EGFP (top, magenta; bottom, gray) in a larval zebrafish xenograft model, expressing the vascular marker Tg(kdrl:Hsa.HRAS-mCherry) for tissue context (top, cyan). Low-resolution imaging (top, 3D rendering) captured the whole organism, while high-resolution imaging (bottom, 3D rendering) revealed subcellular dynamics including an intricate network of protrusions. The cancer cells were xenografted into zebrafish larvae at 2.25 days post-fertilization. Scale bars, 500 μm (top), 25 μm (bottom).
Supplementary Video 7 (download AVI )
Multiscale, time-lapse imaging of U-2 OS osteosarcoma cancer cells (pVimentin-PsmOrange label; top, white; bottom, green) and their interactions with macrophages (Tg(mpeg1:EGFP), magenta) in a larval zebrafish xenograft model. The zebrafish expressed the vascular label Tg(kdrl:Hsa.HRAS-mCherry) (cyan) for tissue context. Maximum intensity projection of the low-resolution data is shown on top, while a 3D rendering of the high-resolution is shown on the bottom. Boxed region on top indicates the location of the high-resolution region on the bottom. Scale bars, 500 μm (top), 50 μm (bottom).
Supplementary Video 8 (download AVI )
High-resolution time-lapse video of U-2 OS osteosarcoma cancer cells (pVimentin-PsmOrange label; top and bottom, green) xenografted into larval zebrafish with labeled macrophages (Tg(mpeg1:EGFP); top, magenta; bottom, gray) and vasculature (Tg(kdrl:Hsa.HRAS-mCherry); top, cyan), displayed as maximum intensity projections over time. Same high-resolution data as in Supplementary Video 7. Scale bar, 30 μm.
Supplementary Video 9 (download AVI )
Low-resolution, time-lapse imaging of macrophages (Tg(mpeg1:EGFP), magenta) in larval zebrafish expressing the vascular marker Tg(kdrl:Hsa.HRAS-mCherry) (cyan) without any xenografted cancer cells (control). Scale bar, 500 μm.
Supplementary Video 10 (download AVI )
High-resolution time-lapse video of A375 melanoma cancer cells (green, pVimentin-PsmOrange label) and their interactions with macrophages (Tg(mpeg1:EGFP), gray), displayed as 3D rendering over time. Scale bar, 50 μm.
Supplementary Video 11 (download AVI )
3D rendering of the segmented macrophages (color scale, violet to green from head to tail) from the low-resolution imaging data of the xenograft experiment with injected U-2 OS osteosarcoma cancer cells (red) (Supplementary Video 7). For tissue context, the vasculature (label, Tg(kdrl:Hsa.HRAS-mCherry)) was rendered in gray. Scale bar, 500 μm.
Supplementary Video 12 (download AVI )
3D rendering of the segmented macrophages (color scale, violet to red from head to tail) from the low-resolution imaging data over time in the control experiment (that is no injected cancer cells, Supplementary Video 9). For tissue context, the vasculature (label, Tg(kdrl:Hsa.HRAS-mCherry)) was rendered in gray. Scale bar, 500 μm.
Supplementary Video 13 (download AVI )
Left: 3D rendering of high-resolution imaging data with segmented macrophages (in color), U-2 OS osteosarcoma cancer cell (in red), and vasculature (in gray). Right: Analysis of macrophage shapes (n = 825) using global morphological feature analysis, displayed in the principal-component space. Each gray point represents a single macrophage. Bagplots were used to visualize all macrophage shapes within an hour of observation in the PCA space (Tukey median (cross); inner polygon (darker color) that contains the 50% observations with largest Tukey depth; the outer polygon (lighter color) with all data excluding outliers). The color of the bagplots represents the classification of macrophage function from Fig. 3g. The red line indicates the change of the Tukey median over time. Scale bar, 50 μm
Supplementary Video 14 (download AVI )
Representative 3D renderings (not to scale) of individual macrophages from xenograft (top) and control (bottom). Color indicates the mean cellular curvature.
Supplementary Source Data (download ZIP )
Statistical Source Data for Supplementary Figs.
Source data
Source Data Fig. 1 (download XLSX )
Statistical Source Data.
Source Data Fig. 3 (download XLSX )
Statistical Source Data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Daetwyler, S., Mazloom-Farsibaf, H., Zhou, F.Y. et al. Imaging of cellular dynamics from a whole organism to subcellular scale with self-driving, multiscale microscopy. Nat Methods 22, 569–578 (2025). https://doi.org/10.1038/s41592-025-02598-2
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-025-02598-2
This article is cited by
-
Second-level high-speed 3D isotropic imaging of whole mouse brain using deep-learning spinning-disk light-sheet microscopy
PhotoniX (2025)
-
When imaging technology meets single-cell omics: new paradigm in developmental biology
Cell Regeneration (2025)
-
Imaging 3D cell cultures with optical microscopy
Nature Methods (2025)
-
Universal consensus 3D segmentation of cells from 2D segmented stacks
Nature Methods (2025)
-
Bioengineering structural anisotropy in living tissues
Nature Reviews Bioengineering (2025)