Abstract
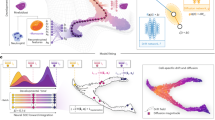
Single-cell sequencing has revolutionized our understanding of cellular heterogeneity and responses to environmental stimuli. However, mapping transcriptomic changes across diverse cell types in response to various stimuli and elucidating underlying disease mechanisms remains challenging. Here we present Squidiff, a diffusion model-based generative framework that predicts transcriptomic changes across diverse cell types in response to environmental changes. We demonstrate the robustness of Squidiff across cell differentiation, gene perturbation and drug response prediction. Through continuous denoising and semantic feature integration, Squidiff learns transient cell states and predicts high-resolution transcriptomic landscapes over time and conditions. Furthermore, we applied Squidiff to model blood vessel organoid development and cellular responses to neutron irradiation and growth factors. Our results demonstrate that Squidiff enables in silico screening of molecular landscapes and cellular state transitions, facilitating rapid hypothesis generation and providing valuable insights into the regulatory principles of cell fate decisions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The public dataset of scRNA-seq of iPSCs differentiating toward endoderm27 was downloaded from Zenodo (https://zenodo.org/records/3625024#.Xil-0y2cZ0s)66. The dataset for nonadditive gene perturbation34 was downloaded from GSE133344. The dataset for drug treatment of melanoma cells8 was downloaded from https://www.research-collection.ethz.ch/handle/20.500.11850/609681. The public dataset of drug screening in glioblastoma35 was downloaded from accession GSE148842. The sci-Plex3 dataset37 was downloaded from GSM4150378. The raw single-cell sequencing data of BVOs are available on figshare (https://doi.org/10.6084/m9.figshare.27948633)67. Source data are provided with this paper.
Code availability
The Squidiff package and code to reproduce the results in this study are available on the GitHub repositories: https://github.com/siyuh/Squidiff and https://github.com/siyuh/Squidiff_reproducibility. The code is also deposited at Zenodo (https://doi.org/10.5281/zenodo.15061773)65.
References
Schneider, G., Schmidt-Supprian, M., Rad, R. & Saur, D. Tissue-specific tumorigenesis: context matters. Nat. Rev. Cancer 17, 239–253 (2017).
Potente, M. & Mäkinen, T. Vascular heterogeneity and specialization in development and disease. Nat. Rev. Mol. Cell Biol. 18, 477–494 (2017).
Rafelski, S. M. & Theriot, J. A. Establishing a conceptual framework for holistic cell states and state transitions. Cell 187, 2633–2651 (2024).
Gullapalli, R. R., Desai, K. V., Santana-Santos, L., Kant, J. A. & Becich, M. J. Next generation sequencing in clinical medicine: challenges and lessons for pathology and biomedical informatics. J. Pathol. Inform. 3, 40 (2012).
Atanasov, A. G., Zotchev, S. B., Dirsch, V. M. & Supuran, C. T. Natural products in drug discovery: advances and opportunities. Nat. Rev. Drug Discov. 20, 200–216 (2021).
Lotfollahi, M., Wolf, F. A. & Theis, F. J. scGen predicts single-cell perturbation responses. Nat. Methods 16, 715–721 (2019).
Kana, O. et al. Generative modeling of single-cell gene expression for dose-dependent chemical perturbations. Patterns 4, 100817 (2023).
Bunne, C. et al. Learning single-cell perturbation responses using neural optimal transport. Nat. Methods 20, 1759–1768 (2023).
Roohani, Y., Huang, K. & Leskovec, J. Predicting transcriptional outcomes of novel multigene perturbations with GEARS. Nat. Biotechnol. 42, 927–935 (2023).
Chen, Y. & Zou, J. Simple and effective embedding model for single-cell biology built from ChatGPT. Nat. Biomed. Eng. 9, 483–493 (2025).
Kingma, D. P. & Welling, M. Auto-encoding variational Bayes. In 2nd International Conference on Learning Representations (ICLR 2014) (eds Bengio, Y. & LeCun, Y.) (OpenReview, 2014).
Yang, L. et al. Diffusion models: a comprehensive survey of methods and applications. ACM Comput. Surv. 56, 1–39 (2023).
Ho, J., Jain, A. & Abbeel, P. Denoising diffusion probabilistic models. In Advances in Neural Information Processing Systems 33 (NeurIPS 2020) (eds Larochelle, H. et al.) 6840–6851 (Curran Associates, 2020).
Guo, Z. et al. Diffusion models in bioinformatics and computational biology. Nat. Rev. Bioeng. 2, 136–154 (2023).
Rombach, R., Blattmann, A., Lorenz, D., Esser, P. & Ommer, B. High-resolution image synthesis with latent diffusion models. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition 10684–10695 (IEEE, 2022).
Pandey, K., Mukherjee, A., Rai, P. & Kumar, A. VAEs meet diffusion models: efficient and high-fidelity generation. In NeurIPS 2021 Workshop on Deep Generative Models and Downstream Applications (OpenReview, 2021); https://openreview.net/pdf?id=-J8dM4ed_92
Sadria, M. & Layton, A. scVAEDer: integrating deep diffusion models and variational autoencoders for single-cell transcriptomics analysis. Genome Biol. 26, 64 (2025).
Luo, E., Hao, M., Wei, L. & Zhang, X. scDiffusion: conditional generation of high-quality single-cell data using diffusion model. Bioinformatics 40, btae518 (2024).
Tang, W. et al. A general single-cell analysis framework via conditional diffusion generative models. Preprint at bioRxiv https://doi.org/10.1101/2023.10.13.562243 (2023).
Preechakul, K., Chatthee, N., Wizadwongsa, S. & Suwajanakorn, S. Diffusion autoencoders: toward a meaningful and decodable representation. In 2022 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) 10619–10629 (IEEE, 2022).
Bunne, C. et al. How to build the virtual cell with artificial intelligence: priorities and opportunities. Cell 187, 7045–7063 (2024).
Song, J., Meng, C. & Ermon, S. Denoising diffusion implicit models. In 9th International Conference on Learning Representations (ICLR 2021) (OpenReview, 2021); https://openreview.net/pdf?id=St1giarCHLP
Yahaya, B. H. Organoid Technology for Disease Modelling and Personalized Treatment (Springer Nature, 2022).
Nikolova, M. T. et al. Fate and state transitions during human blood vessel organoid development. Cell 188, 3329–3348.e31 (2025).
Ho, J., Jain, A. & Abbeel, P. Denoising diffusion probabilistic models. Adv. Neural Inf. Process. Syst. 33, 6840–6851 (2020).
Zappia, L., Phipson, B. & Oshlack, A. Splatter: simulation of single-cell RNA sequencing data. Genome Biol. 18, 174 (2017).
Cuomo, A. S. E., et al. Single-cell RNA-sequencing of differentiating iPS cells reveals dynamic genetic effects on gene expression. Nat. Commun. 11, 810 (2020).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Schrode, N., Saiz, N., Di Talia, S. & Hadjantonakis, A.-K. GATA6 levels modulate primitive endoderm cell fate choice and timing in the mouse blastocyst. Dev. Cell 29, 454–467 (2014).
Wilson, V. & Beddington, R. Expression of T protein in the primitive streak is necessary and sufficient for posterior mesoderm movement and somite differentiation. Dev. Biol. 192, 45–58 (1997).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Cannoodt, R. et al. SCORPIUS improves trajectory inference and identifies novel modules in dendritic cell development. Preprint at bioRxiv https://doi.org/10.1101/079509 (2016).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Norman, T. M. et al. Exploring genetic interaction manifolds constructed from rich single-cell phenotypes. Science 365, 786–793 (2019).
Zhao, W., et al. Deconvolution of cell type-specific drug responses in human tumor tissue with single-cell RNA-seq. Genome Med. 13, 82 (2021).
Qi, X., et al. Predicting transcriptional responses to novel chemical perturbations using deep generative model for drug discovery. Nat. Commun. 15, 9256 (2024).
Srivatsan, S. R., et al. Massively multiplex chemical transcriptomics at single-cell resolution. Science 367, 45–51 (2020).
Hofer, M. & Lutolf, M. P. Engineering organoids. Nat. Rev. Mater. 6, 402–420 (2021).
Yang, S. et al. Organoids: the current status and biomedical applications. MedComm 4, e274 (2023).
Huang, Y. et al. Deciphering the impact of aging on splenic endothelial cell heterogeneity and immunosenescence through single-cell RNA sequencing analysis. Immun. Ageing 21, 48 (2024).
Mettler, F. A. Jr & Voelz, G. L. Major radiation exposure — what to expect and how to respond. N. Engl. J. Med. 346, 1554–1561 (2002).
Kameni, L. E. et al. A review of radiation-induced vascular injury and clinical impact. Ann. Plast. Surg. 92, 181–185 (2024).
Durante, M. New challenges in high-energy particle radiobiology. Br. J. Radiol. 87, 20130626 (2014).
Tavakol, D. N. et al. Modeling and countering the effects of cosmic radiation using bioengineered human tissues. Biomaterials 301, 122267 (2023).
Chancellor, J. C., Scott, G. B. I. & Sutton, J. P. Space radiation: the number one risk to astronaut health beyond low earth orbit. Life 4, 491–510 (2014).
Wijerathne, H. et al. Mechanisms of radiation-induced endothelium damage: emerging models and technologies. Radiother. Oncol. 158, 21–32 (2021).
Oliner, J. D., Saiki, A. Y. & Caenepeel, S. The role of MDM2 amplification and overexpression in tumorigenesis. Cold Spring Harb. Perspect. Med. 6, a026336 (2016).
Ruef, C. & Coleman, D. L. Granulocyte–macrophage colony-stimulating factor: pleiotropic cytokine with potential clinical usefulness. Rev. Infect. Dis. 12, 41–62 (1990).
Ping, S., Qiu, X., Gonzalez-Toledo, M. E., Liu, X. & Zhao, L.-R. Stem cell factor in combination with granulocyte colony-stimulating factor reduces cerebral capillary thrombosis in a mouse model of CADASIL. Cell Transplant. 27, 637–647 (2018).
Furmento, V. A., Marino, J., Blank, V. C. & Roguin, L. P. The granulocyte colony-stimulating factor (G-CSF) upregulates metalloproteinase-2 and VEGF through PI3K/Akt and Erk1/2 activation in human trophoblast Swan 71 cells. Placenta 35, 937–946 (2014).
Boneberg, E. M. & Hartung, T. Molecular aspects of anti-inflammatory action of G-CSF. Inflamm. Res. 51, 119–128 (2002).
Magné, N. et al. NF-κB modulation and ionizing radiation: mechanisms and future directions for cancer treatment. Cancer Lett. 231, 158–168 (2006).
Hughson, R. L., Helm, A. & Durante, M. Heart in space: effect of the extraterrestrial environment on the cardiovascular system. Nat. Rev. Cardiol. 15, 167–180 (2018).
Delp, M. D., Charvat, J. M., Limoli, C. L., Globus, R. K. & Ghosh, P. Apollo lunar astronauts show higher cardiovascular disease mortality: possible deep space radiation effects on the vascular endothelium. Sci. Rep. 6, 29901 (2016).
Barcellos-Hoff, M. H., et al. Concepts and challenges in cancer risk prediction for the space radiation environment. Life Sci. Space Res. 6, 92–103 (2015).
Ho, J. et al. Video diffusion models. Adv. Neural Inf. Process. Syst. 35, 8633–8646 (2022).
Wu, K. E. et al. Protein structure generation via folding diffusion. Nat. Commun. 15, 1059 (2024).
Virshup, I. et al. The scverse project provides a computational ecosystem for single-cell omics data analysis. Nat. Biotechnol. 41, 604–606 (2023).
Werschler, N. & Penninger, J. Generation of human blood vessel organoids from pluripotent stem cells. J. Vis. Exp. https://doi.org/10.3791/64715 (2023).
Tomer, R. et al. SPED light sheet microscopy: fast mapping of biological system structure and function. Cell 163, 1796–1806 (2015).
Xu, Y. et al. Accelerator-based biological irradiation facility simulating neutron exposure from an improvised nuclear device. Radiat. Res. 184, 404–410 (2015).
Xu, Y. Broad energy range neutron spectroscopy using a liquid scintillator and a proportional counter: application to a neutron spectrum similar to that from an improvised nuclear device. Nucl. Instrum. Methods Phys. Res. A 794, 234–239 (2015).
Tavakol, D. N. et al. Modeling the effects of protracted cosmic radiation in a human organ-on-chip platform. Adv. Sci. 11, 2401415 (2024).
Holden, S. et al. Effects of acute and chronic exposure to a mixed field of neutrons and photons and single or fractionated simulated galactic cosmic ray exposure on behavioral and cognitive performance in mice. Radiat. Res. 196, 31–39 (2021).
He, S. et al. Squidiff. Zenodo https://doi.org/10.5281/zenodo.15061773 (2025).
Cuomo, A. S. E. Single-cell RNA-sequencing of iPS cells differentiating towards definitive endoderm. Zenodo https://zenodo.org/records/3625024#.Xil-0y2cZ0s (2020).
He, S. et al. Squidiff: predicting cellular development and responses to perturbations using a diffusion model. figshare https://doi.org/10.6084/m9.figshare.27948633 (2025).
Acknowledgements
We thank S. Quake and K. Preechakul for insightful discussions on virtual cells and the diffusion model; and S. Wang, J. H. Lee and R.B.-L. Berris for their assistance with scRNA-seq analysis and cell culture. We also appreciate the staff at Columbia’s Center for Radiological Research for their support in operating ionizing radiation equipment. This study used resources from the Herbert Irving Comprehensive Cancer Center Confocal, Specialized Microscopy Shared Resource and the Genomics and High Throughput Screening Shared Resource, partially funded by NIH/NCI Cancer Center Support Grant P30CA013696. The Columbia IND Neutron Facility was developed under NIAID grant U19 AI067773. We gratefully acknowledge funding support from the Translational Research Institute for Space Health (TRISH/NASA) (RAD0104 and NNX16A069A to G.V.-N. and K.W.L.).
Author information
Authors and Affiliations
Contributions
K.W.L., E.A. and J.Z. conceived the study and provided overall supervision of the study. S.H., Y.Z. and D.N.T. designed the BVO study and performed experiments. S.H. and H.Y. designed and developed the model. S.H., Y.Z., D.N.T., H.Y., Z.Z., C.X. and S.C. analyzed and interpreted data. G.G. performed irradiation experiments. R.T. and G.V.-N. provided additional supervision. S.H., Y.Z., D.N.T., H.Y., J.Z., E.A. and K.W.L. wrote the paper. All authors reviewed, contributed to and approved the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Sudin Bhattacharya, Qing Nie and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reportsare available. Primary Handling Editor: Madhura Mukhopadhyay, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information (download PDF )
Supplementary Table 1 and Figs. 1–10.
Supplementary Video 1 (download AVI )
3D rending of one CD31-stained blood vessel organoid.
Source data
Source Data Fig. 5 (download ZIP )
Multichannel confocal imaging for BVOs.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, S., Zhu, Y., Tavakol, D.N. et al. Squidiff: predicting cellular development and responses to perturbations using a diffusion model. Nat Methods 23, 65–77 (2026). https://doi.org/10.1038/s41592-025-02877-y
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-025-02877-y
This article is cited by
-
Interpretation, extrapolation and perturbation of single cells
Nature Reviews Genetics (2026)