Abstract
Despite advances in whole-brain imaging technologies, the lack of quantitative approaches to bridge rodent preclinical and human studies remains a critical challenge. Here we present TransBrain, a computational framework enabling bidirectional translation of brain-wide phenotypes between humans and mice. TransBrain improves human–mouse homology mapping accuracy through (1) a cortical and subcortical detached region-specific deep neural network trained on integrated multimodal human transcriptomics to improve cortical correspondence (89.5% improvement over the original transcriptome), which revealed 2 evolutionarily conserved gradients, and (2) a graph-based approach to construct a unified cross-species representational space incorporating anatomical hierarchies and structural connectivity. We demonstrate TransBrain’s utility through three cross-species applications: quantitative assessment of resting-state brain organizational features, inferring human cognitive functions from mouse optogenetic circuits and translating molecular insights from mouse models to individual-level mechanisms in autism. TransBrain enables quantitative cross-species comparison and mechanistic investigation of both normal and pathological brain functions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The AHBA is available at http://human.brain-map.org; the single-nucleus dataset for the human brain is available at https://storage.cloud.google.com/linnarsson-lab-human. Processed mouse gene expression data and 3D coordinates of spots can be accessed at https://www.molecularatlas.org, and mouse connectivity data from the Allen Mouse Brain Connectivity Atlas are available at https://connectivity.brain-map.org. Neuroimaging data for this research were procured from the publicly available HCP Young Adult Project, which can be found at https://db.humanconnectome.org. The awake mouse resting fMRI dataset can be downloaded from https://data.mendeley.com/datasets/np2fx99hn6/2. Processed optogenetic fMRI datasets for the insula and the DRN are available at https://data.ru.nl/collections/di/dcmn/DSC_4180000.18_380. Preprocessed ABIDE I data of 1,102 participants can be accessed at https://fcon_1000.projects.nitrc.org/indi/abide/. The mouse structural imaging dataset related to autism is available at https://www.braincode.ca/content/public-data-releases#dr001. Processed transcriptomic datasets combining AHBA and snRNA-seq data are available on Zenodo, with details at https://github.com/ibpshangzheng/transbrain/blob/main/pipeline/datasets/README.md. Other atlases and imaging datasets used in this study were all obtained from previous publications, as detailed in Methods. Source data are provided with this paper.
Code availability
The original graph representation learning framework is publicly available on GitHub (https://github.com/aditya-grover/node2vec) under the MIT License. The TransBrain toolbox, including installation instructions, step-by-step tutorials, analysis processes for main results, application cases and visualizations, is publicly available at https://github.com/ibpshangzheng/transbrain under the Apache License version 2.0, with comprehensive documentation at https://transbrain.readthedocs.io/en/latest/ and an interactive platform at https://transbrain.streamlit.app/ for users without programming experience. All code is also available in a Code Ocean capsule66, accessible at https://doi.org/10.24433/CO.0833860.v1, under the Apache License version 2.0.
References
Finn, E. S., Poldrack, R. A. & Shine, J. M. Functional neuroimaging as a catalyst for integrated neuroscience. Nature 623, 263–273 (2023).
Gozzi, A. & Zerbi, V. Modeling brain dysconnectivity in rodents. Biol. Psychiatry 93, 419–429 (2023).
Buch, A. M. et al. Molecular and network-level mechanisms explaining individual differences in autism spectrum disorder. Nat. Neurosci. 26, 650–663 (2023).
Tozzi, L. et al. Personalized brain circuit scores identify clinically distinct biotypes in depression and anxiety. Nat. Med. 30, 2076–2087 (2024).
Ellegood, J. et al. Clustering autism: using neuroanatomical differences in 26 mouse models to gain insight into the heterogeneity. Mol. Psychiatry 20, 118–125 (2015).
Zerbi, V. et al. Brain mapping across 16 autism mouse models reveals a spectrum of functional connectivity subtypes. Mol. Psychiatry 26, 7610–7620 (2021).
Liu, J. et al. Astrocyte dysfunction drives abnormal resting-state functional connectivity in depression. Sci. Adv. 8, eabo2098 (2022).
Balsters, J. H., Zerbi, V., Sallet, J., Wenderoth, N. & Mars, R. B. Primate homologs of mouse cortico-striatal circuits. eLife 9, e53680 (2020).
Oldham, S. & Ball, G. A phylogenetically-conserved axis of thalamocortical connectivity in the human brain. Nat. Commun. 14, 6032 (2023).
Gutierrez-Barragan, D., Ramirez, J. S. B., Panzeri, S., Xu, T. & Gozzi, A. Evolutionarily conserved fMRI network dynamics in the mouse, macaque, and human brain. Nat. Commun. 15, 8518 (2024).
Pagani, M. et al. mTOR-related synaptic pathology causes autism spectrum disorder-associated functional hyperconnectivity. Nat. Commun. 12, 6084 (2021).
Guma, E. et al. Comparative neuroimaging of sex differences in human and mouse brain anatomy. eLife 13, RP92200 (2024).
Beauchamp, A. et al. Whole-brain comparison of rodent and human brains using spatial transcriptomics. eLife 11, e79418 (2022).
Mandino, F. et al. A triple-network organization for the mouse brain. Mol. Psychiatry 27, 865–872 (2022).
Lein, E. S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Zhang, B., Zhang, S. & Zhang, S. Whole brain alignment of spatial transcriptomics between humans and mice with BrainAlign. Nat. Commun. 15, 6302 (2024).
Hawrylycz, M. et al. Canonical genetic signatures of the adult human brain. Nat. Neurosci. 18, 1832–1844 (2015).
Siletti, K. et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023).
Mars, R. B. et al. Whole brain comparative anatomy using connectivity blueprints. eLife 7, e35237 (2018).
Xu, T. et al. Cross-species functional alignment reveals evolutionary hierarchy within the connectome. Neuroimage 223, 117346 (2020).
de Schotten, M. T. & Forkel, S. J. The emergent properties of the connected brain. Science 378, 505–510 (2022).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Oh, S. W. et al. A mesoscale connectome of the mouse brain. Nature 508, 207–214 (2014).
Carlén, M. What constitutes the prefrontal cortex? Science 358, 478–482 (2017).
Vann, S. D., Aggleton, J. P. & Maguire, E. A. What does the retrosplenial cortex do? Nat. Rev. Neurosci. 10, 792–802 (2009).
Dear, R. et al. Cortical gene expression architecture links healthy neurodevelopment to the imaging, transcriptomics and genetics of autism and schizophrenia. Nat. Neurosci. 27, 1075–1086 (2024).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Ortiz, C. et al. Molecular atlas of the adult mouse brain. Sci. Adv. 6, eabb3446 (2020).
Fulcher, B. D., Murray, J. D., Zerbi, V. & Wang, X.-J. Multimodal gradients across mouse cortex. Proc. Natl Acad. Sci. USA 116, 4689–4695 (2019).
Croxson, P. L., Forkel, S. J., Cerliani, L. & de Schotten, M. T. Structural variability across the primate brain: a cross-species comparison. Cereb. Cortex 28, 3829–3841 (2017).
Uylings, H. B. M., Groenewegen, H. J. & Kolb, B. Do rats have a prefrontal cortex? Behav. Brain Res. 146, 3–17 (2003).
Assaf, Y., Bouznach, A., Zomet, O., Marom, A. & Yovel, Y. Conservation of brain connectivity and wiring across the mammalian class. Nat. Neurosci. 23, 805–808 (2020).
Schaeffer, D. J. et al. Divergence of rodent and primate medial frontal cortex functional connectivity. Proc. Natl Acad. Sci. USA 117, 21681–21689 (2020).
Rosenthal, G. et al. Mapping higher-order relations between brain structure and function with embedded vector representations of connectomes. Nat. Commun. 9, 2178 (2018).
Bang, D., Lim, S., Lee, S. & Kim, S. Biomedical knowledge graph learning for drug repurposing by extending guilt-by-association to multiple layers. Nat. Commun. 14, 3570 (2023).
Margulies, D. S. et al. Situating the default-mode network along a principal gradient of macroscale cortical organization. Proc. Natl Acad. Sci. USA 113, 12574–12579 (2016).
Coletta, L. et al. Network structure of the mouse brain connectome with voxel resolution. Sci. Adv. 6, eabb7187 (2020).
Gutierrez-Barragan, D. et al. Unique spatiotemporal fMRI dynamics in the awake mouse brain. Curr. Biol. 32, 631–644 (2022).
Huang, Z., Zhang, J., Wu, J., Mashour, G. A. & Hudetz, A. G. Temporal circuit of macroscale dynamic brain activity supports human consciousness. Sci. Adv. 6, eaaz0087 (2020).
Yeo, B. T. T. et al. The organization of the human cerebral cortex estimated by intrinsic functional connectivity. J. Neurophysiol. 106, 1125–1165 (2011).
Pagani, M., Gutierrez-Barragan, D., de Guzman, A. E., Xu, T. & Gozzi, A. Mapping and comparing fMRI connectivity networks across species. Commun. Biol. 6, 1238 (2023).
Váša, F. & Mišić, B. Null models in network neuroscience. Nat. Rev. Neurosci. 23, 493–504 (2022).
Yarkoni, T., Poldrack, R. A., Nichols, T. E., Van Essen, D. C. & Wager, T. D. Large-scale automated synthesis of human functional neuroimaging data. Nat. Methods 8, 665–670 (2011).
Uddin, L. Q. Salience processing and insular cortical function and dysfunction. Nat. Rev. Neurosci. 16, 55–61 (2015).
Salvan, P. et al. Serotonin regulation of behavior via large-scale neuromodulation of serotonin receptor networks. Nat. Neurosci. 26, 53–63 (2023).
Rutherford, S. et al. The normative modeling framework for computational psychiatry. Nat. Protoc. 17, 1711–1734 (2022).
Köhler, S. et al. The Human Phenotype Ontology in 2021. Nucleic Acids Res. 49, D1207–D1217 (2021).
Liu, X., Shen, Q. & Zhang, S. Cross-species cell-type assignment from single-cell RNA-seq data by a heterogeneous graph neural network. Genome Res. 33, 96–111 (2023).
Shi, Y. et al. Mouse and human share conserved transcriptional programs for interneuron development. Science 374, eabj6641 (2021).
Maier-Hein, K. H. et al. The challenge of mapping the human connectome based on diffusion tractography. Nat. Commun. 8, 1349 (2017).
Fan, L. et al. The Human Brainnetome Atlas: a new brain atlas based on connectional architecture. Cereb. Cortex 26, 3508–3526 (2016).
Winterburn, J. L. et al. A novel in vivo atlas of human hippocampal subfields using high-resolution 3 T magnetic resonance imaging. Neuroimage 74, 254–265 (2013).
Ding, S.-L. et al. Allen Human Reference Atlas—3D, 2020. Allen Brain Map Community Forum https://community.brain-map.org/t/allen-human-reference-atlas-3d-2020-new/405 (2020).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Tzourio-Mazoyer, N. et al. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 15, 273–289 (2002).
Wang, Q. et al. The Allen Mouse Brain Common Coordinate Framework: a 3D reference atlas. Cell 181, 936–953 (2020).
Markello, R. D. et al. Standardizing workflows in imaging transcriptomics with the abagen toolbox. eLife 10, e72129 (2021).
Zhou, Y. et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat. Commun. 10, 1523 (2019).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80, 105–124 (2013).
Tournier, J.-D. et al. MRtrix3: a fast, flexible and open software framework for medical image processing and visualisation. Neuroimage 202, 116137 (2019).
Knox, J. E. et al. High-resolution data-driven model of the mouse connectome. Netw. Neurosci. 3, 217–236 (2018).
Alexander-Bloch, A. F. et al. On testing for spatial correspondence between maps of human brain structure and function. Neuroimage 178, 540–551 (2018).
Markello, R. D. et al. neuromaps: structural and functional interpretation of brain maps. Nat. Methods 19, 1472–1479 (2022).
Grandjean, J. et al. A brain-wide functional map of the serotonergic responses to acute stress and fluoxetine. Nat. Commun. 10, 350 (2019).
Esteban, O. et al. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16, 111–116 (2019).
Huang, S. TransBrain: a computational framework for translating brain-wide phenotypes between humans and mice. Code Ocean https://doi.org/10.24433/CO.0833860.v1 (2025).
Acknowledgements
This work was supported by the STI2030-Major Projects (2022ZD0211900 to A.L. and Y.S.), the National Natural Science Foundation of China (32192411 to X.W. and 32122037 to Q. Wu), the Beijing Nova Program (20250484761 to A.L.), the New Cornerstone Science Foundation (to X.W.), Changping Laboratory and the Fundamental Research Funds for the Central Universities (to X.W.). The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript. We are grateful to L. Fan for expert help in diffusion-based tractography and to Y. Wang for help with the figures.
Author information
Authors and Affiliations
Contributions
A.L. and X.W. led the project. A.L., X.W. and S.H. were responsible for the study concept and the design of the study. S.H., A.L. and T.Z. analyzed data. S.H. and A.L. created figures and wrote the paper. T.Z., C.D., Y.S. and Y.P. aided in writing paper. X.L., K.L., L.C., Q. Wang, Y.H., Y.G., F.X., X.T., J.X., C.Z., Q. Wu, Y.Z., L.L. and B.L. participated in discussions of the results and the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks James Shine and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Nina Vogt, in collaboration with the Nature Methods team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The overall framework to develop TransBrain.
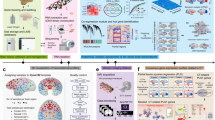
(a) TransBrain’s goal: To quantitatively map brain-wide phenotypes between humans and mice through (1) cross-species transcriptional alignment and (2) integration of anatomical hierarchies and connectivity patterns. (b) Overview of the TransBrain pipeline. We integrated two modalities of human brain transcriptional data as input. A detached deep neural network model was used to capture region-specific transcriptional embeddings for cortical and subcortical, respectively, which were then directly applied to spatial transcriptomics data of mice. Using species-specific transcriptional embeddings as a bridge, we constructed a heterograph that integrates structural connectivity and anatomical hierarchies. Random walk-based graph representation learning algorithm was performed on the heterograph to obtain latent embeddings. Finally, a dual regression approach was employed to compare macroscopic imaging phenotypes between species. (c) Applications include: cross-species comparison of resting-state networks, translation of mouse optogenetic circuits to human brain functions, and evaluation of mouse models for human brain disorders. Human brain image in a from Freepik.com; mouse brain image in a from Shutterstock; human icons in c from FlatIcon.com.
Extended Data Fig. 2 The pipeline to test the deep neural network model’s generalizability and accuracy.
(a) One of the six AHBA donors was excluded as the independent test set, while the remaining five were used for training. Each AHBA/single-nucleus donor pair underwent independent integration. For all combinations, integration was repeated 100 times, and each dataset underwent 10 rounds of model training and testing. During training, stratified 10% validation sets were defined based on pre-integration AHBA sample labels per brain region.
Extended Data Fig. 3 Leave-one-out independent validation of the deep neural network models.
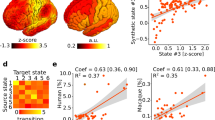
(a) and (b), Average confusion matrix of cortical and subcortical regions (cortical: detached model mean accuracy=32%, shared model mean accuracy=24%; subcortical: detached model mean accuracy=36%, shared model mean accuracy=29%). (c) True positives rate (%) visualization of average confusion matrix of cortical and subcortical regions. (d) Comparison of true positives rate (%) in the average confusion matrix (detached model vs shared model; cortical (n = 61 ROIs): t(60) = 3.77, P = 3.75×10−4; subcortical (n = 18 ROIs): t(17) = 2.20, P = 0.042; detached model vs random classifier; cortical (n = 61 ROIs): t(60) = 11.42, P = 1.09×10−16; subcortical (n = 18 ROIs): t(17) = 5.29, P = 5.99×10−5; two-sided paired-samples t-test). Box plots show the median (centre line), 25th and 75th percentiles (box bounds), and whiskers extending to 1.5 × IQR. *P < 0.05, ***P < 0.001.
Extended Data Fig. 4 Region-specific transcriptional profiles in the human subcortex.
(a) The autocorrelation matrix of region-specific transcriptional embeddings in the human subcortex was divided into four sections. (Amy: Amygdala, red box; Hippo: Hippocampus, green box; BG: Basal ganglia, blue box; Tha: Thalamus, purple box). (b) Comparison of the hierarchical value of region-specific transcriptional embeddings. The hierarchy value, defined as the ratio of the mean similarity of each ROI to other ROIs within the same hierarchy versus the mean similarity to ROIs outside that hierarchy in the autocorrelation matrix (detached model vs shared model; Hippo (n = 4 ROIs): t(3) = 11.69, P = 0.0013; BG (n = 8 ROIs): t(7) = 2.61, P = 0.035; Tha (n = 8 ROIs): t(7) = 2.96, P = 0.021; two-sided paired-samples t-test). Box plots show the median (centre line), 25th and 75th percentiles (box bounds), and whiskers extending to 1.5 × IQR. (c) Comparison of the specificity of region-specific transcriptional embeddings. Similarity vectors for each ROI were normalized to a range of [0,1], sorted in descending order, and average (n = 22 ROIs). By comparing the differences in scaled similarity at the same rank, we assessed the specificity of transcriptional embeddings (detached model vs shared model: t(21) = -8.81, P = 1.70×10−8; two-sided paired-samples t-test). Lines show the mean. *P < 0.05, **P < 0.01, ***P < 0.001.
Extended Data Fig. 5 Pathway and process enrichment of model-contributing genes.
(a) and (b) Bubble plot illustrating the results of gene enrichment analysis across biological processes. The X-axis represents enrichment percentage (%), and the Y-axis lists enriched GO terms. Bubble size corresponds to the number of genes associated with each term, and bubble color indicates the statistical significance (adjusted q-value), with darker colors representing lower Q-values. Only the top enriched terms are shown for clarity; the additional gene enrichment analysis results are available in the Supplementary Tables 3 and 4. (c) Representative enriched modules were selected for display. (cortex: 697 genes, subcortex: 331 genes, overlap: 63 genes). (d) Permutation test (1000 iterations) showing minimal overlap between cortical and subcortical region-specific genes (overlap rate=0.054, Pperm = 0.001, one-sided permutation test).
Extended Data Fig. 6 Cross-species cortical region-specific transcriptional similarity patterns.
(a) Visualization of region-specific transcriptional similarity patterns between specific mouse brain regions and the entire human brain. (PL, prelimbic cortex; SSp-bfd, primary somatosensory cortex, barrel field; VISp, primary visual cortex; M1, primary motor cortex; S1, primary somatosensory cortex; vmPFC, ventromedial prefrontal cortex; ACA, anterior cingulate cortex). (b) and (c) Region-specific transcriptional similarity matrices from the shared model and detached model for cortical regions.
Extended Data Fig. 7 Predicting mouse cortical gradients using linear model.
(a) Spatial distribution of principal components (z-score) across species predicted by a linear model. (b–d) Compared the similarity between linear model predicted mouse cortical transcriptional gradients and detached model transformed gradients (Gradient 1: r = 0.90, P = 2.25×10−14; Gradient 2: r = 0.53, P = 5.07×10−4; Gradient 3: r = 0.60, P = 5.11×10−5; df = 37, n = 39 mouse cortical ROIs; Two-sided Pearson correlations). Shaded bands represent 95% confidence intervals around the regression line.
Extended Data Fig. 8 Optimization of cross-species graph representation learning approach.
(a) Cross-species fully connected graph used for constructing graph embeddings. (b) Selection of exploratory factor p and locality factor q during the construction of graph embeddings. The parameters were shifted from local (p = 10−4, q = 10) to more global (p = 0.1, q = 0.1), with 100 gradient parameters set within this range. Graph embeddings were reconstructed from different parameters to obtain cross-species similarity matrices, and the rank of potential homologous brain regions was evaluated. The rank for identifying homologous regions reaching a minimum at p = 0.01, q = 0.1 (indicated by the orange scatter point in the figure, rank=5.72). (c) The averaged fully connected graph between humans and mice was reconstructed using graph embeddings. (d) Cross-species similarity matrix reconstructed from graph embeddings was compared to the similarity matrix derived from region-specific transcriptional embeddings after hierarchical constraint (with 0 elements in the matrices excluded; r = 0.53, df = 715, n = 717 human-mouse regional pairs, P = 7.97×10−56; Two-sided Pearson correlations). Shaded bands represent 95% confidence intervals around the regression line. (e) Rank of potential homologous regions (n = 38) derived from the two cross-species similarity matrices showed no significant difference (graph embeddings vs transcriptional embeddings; t(37) = 0.02, P = 0.985; two-sided paired-samples t-test). Box plots show the median (centre line), 25th and 75th percentiles (box bounds), and whiskers extending to 1.5 × IQR.
Extended Data Fig. 9 Evolutionary relationships in resting-state networks of humans and mice.
(a–c) Detailed visualization of evolutionary relationships in Fig. 4. Cross-species similarity is depicted by solid lines, where thicker and more reddish lines indicate stronger evolutionary matching between network organizational patterns of humans and mice. The map of the mouse is displayed on the coronal plane using anterior-posterior coordinates. Z-score normalization was applied to the data before visualization.
Supplementary information
Supplementary Information (download PDF )
Supplementary Notes 1–9 and Figs. 1–20.
Supplementary Tables 1–4 (download XLSX )
Ranking of potential homologous regions, hierarchical brain relationships and region-specific gene enrichments.
Source data
Source Data Fig. 1 (download XLSX )
Data used to generate Fig. 1.
Source Data Fig. 2 (download XLSX )
Data used to generate Fig. 2.
Source Data Fig. 3 (download XLSX )
Data used to generate Fig. 3.
Source Data Fig. 4 (download XLSX )
Data used to generate Fig. 4.
Source Data Fig. 5 (download XLSX )
Data used to generate Fig. 5.
Source Data Fig. 6 (download XLSX )
Data used to generate Fig. 6.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, S., Zhang, T., Dong, C. et al. TransBrain: a computational framework for translating brain-wide phenotypes between humans and mice. Nat Methods 23, 426–437 (2026). https://doi.org/10.1038/s41592-025-02961-3
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41592-025-02961-3