Abstract
Transient receptor potential channel subfamily M member 3 (TRPM3) is a Ca2+-permeable cation channel activated by the neurosteroid pregnenolone sulfate (PregS) or heat, serving as a nociceptor in the peripheral sensory system. Recent discoveries of autosomal dominant neurodevelopmental disorders caused by gain-of-function mutations in TRPM3 highlight its role in the central nervous system. Notably, the TRPM3 inhibitor primidone, an anticonvulsant, has proven effective in treating patients with TRPM3-linked neurological disorders and in mouse models of thermal nociception. However, our understanding of neurosteroids, inhibitors and disease mutations on TRPM3 is limited. Here we present cryogenic electron microscopy structures of the mouse TRPM3 in complex with cholesteryl hemisuccinate, primidone and PregS with the synthetic agonist CIM 0216. Our studies identify the binding sites for the neurosteroid, synthetic agonist and inhibitor and offer insights into their effects and disease mutations on TRPM3 gating, aiding future drug development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
For the CHS-bound TRPM3, the primidone-bound TRPM3 and the PregS/CIM-bound TRPM3 structures, the coordinates have been deposited in the Protein Data Bank (PDB) under IDs PDB 9B29, PDB 9B28 and PDB 9B2A; the cryo-EM density maps have been deposited in the Electron Microscopy Data Bank (EMDB) under IDs EMD-44101, EMD-44100 and EMD-44102. The molecular dynamics simulation data used in this study are available from the Zenodo OpenAIRE database (https://zenodo.org/doi/10.5281/zenodo.13369813) ref. 79. Source data are provided with this paper.
Code availability
For molecular dynamics simulations, the following software packages were used for performing simulations and analysis: CHARMM-GUI (free to academic users), OpenMM (v.7.7; open source), CHARMM (v.45b1; free to academic users) and VMD (v.1.9; open source).
References
Su, S., Yudin, Y., Kim, N., Tao, Y. X. & Rohacs, T. TRPM3 channels play roles in heat hypersensitivity and spontaneous pain after nerve injury. J. Neurosci. 41, 2457–2474 (2021).
Aloi, V. D. et al. TRPM3 as a novel target to alleviate acute oxaliplatin-induced peripheral neuropathic pain. PAIN 164, 2060–2069 (2023).
Vriens, J., Nilius, B. & Voets, T. Peripheral thermosensation in mammals. Nat. Rev. Neurosci. 15, 573–589 (2014).
Vriens, J. et al. TRPM3 is a nociceptor channel involved in the detection of noxious heat. Neuron 70, 482–494 (2011).
Held, K. et al. Activation of TRPM3 by a potent synthetic ligand reveals a role in peptide release. Proc. Natl Acad. Sci. USA 112, E1363–E1372 (2015).
Vandewauw, I. et al. A TRP channel trio mediates acute noxious heat sensing. Nature 555, 662–666 (2018).
Wagner, T. F. J. et al. Transient receptor potential M3 channels are ionotropic steroid receptors in pancreatic β cells. Nat. Cell Biol. 10, 1421–1430 (2008).
Straub, I. et al. Flavanones that selectively inhibit TRPM3 attenuate thermal nociception in vivo. Mol. Pharmacol. 84, 736–750 (2013).
Krügel, U., Straub, I., Beckmann, H. & Schaefer, M. Primidone inhibits TRPM3 and attenuates thermal nociception in vivo. PAIN 158, 856–867 (2017).
Gavva, N. R. et al. The vanilloid receptor TRPV1 is tonically activated in vivo and involved in body temperature regulation. J. Neurosci. 27, 3366–3374 (2007).
Gavva, N. R. et al. Pharmacological blockade of the vanilloid receptor TRPV1 elicits marked hyperthermia in humans. PAIN 136, 202–210 (2008).
Burglen, L. et al. Gain-of-function variants in the ion channel gene TRPM3 underlie a spectrum of neurodevelopmental disorders. Elife 12, e81032 (2023).
Van Hoeymissen, E. et al. Gain of channel function and modified gating properties in TRPM3 mutants causing intellectual disability and epilepsy. Elife 9, e57190 (2020).
Sundaramurthi, J. C. et al. De novo TRPM3 missense variant associated with neurodevelopmental delay and manifestations of cerebral palsy. Cold Spring Harb. Mol. Case Stud. 9, a006293 (2023).
Dyment, D., et al. TRPM3-related neurodevelopmental disorder. In GeneReviews® [Internet] (eds. Adam, M.P. et al.) (Univ. of Washington, 2023).
Lines, M. A. et al. Phenotypic spectrum of the recurrent TRPM3 p.(Val837Met) substitution in seven individuals with global developmental delay and hypotonia. Am. J. Med. Genet. A 188, 1667–1675 (2022).
Kang, Q. et al. A Chinese patient with developmental and epileptic encephalopathies (DEE) carrying a TRPM3 gene mutation: a paediatric case report. BMC Pediatr. 21, 256 (2021).
Becker, L. L. et al. Primidone improves symptoms in TRPM3-linked developmental and epileptic encephalopathy with spike-and-wave activation in sleep. Epilepsia 64, e61–e68 (2023).
Harteneck, C. Pregnenolone sulfate: from steroid metabolite to TRP channel ligand. Molecules 18, 12012–12028 (2013).
Reddy, D. S. Neurosteroids: endogenous role in the human brain and therapeutic potentials. Prog. Brain Res. 186, 113–137 (2010).
Reddy, D. S. & Rogawski, M. A. Neurosteroids as endogenous regulators of seizure susceptibility and use in the treatment of epilepsy. Epilepsia 51, 84 (2010).
Belelli, D., Hogenkamp, D., Gee, K. W. & Lambert, J. J. Realising the therapeutic potential of neuroactive steroid modulators of the GABAA receptor. Neurobiol. Stress 12, 100207 (2020).
Wong, P., Sze, Y., Chang, C. C., Lee, J. & Zhang, X. Pregnenolone sulfate normalizes schizophrenia-like behaviors in dopamine transporter knockout mice through the AKT/GSK3β pathway. Transl. Psychiatry 5, e528 (2015).
Seljeset, S., Liebowitz, S., Bright, D. P. & Smart, T. G. Pre- and postsynaptic modulation of hippocampal inhibitory synaptic transmission by pregnenolone sulphate. Neuropharmacology 233, 109530 (2023).
Vriens, J. et al. Opening of an alternative ion permeation pathway in a nociceptor TRP channel. Nat. Chem. Biol. 10, 188–195 (2014).
Badheka, D. et al. Inhibition of transient receptor potential melastatin 3 ion channels by G-protein βγ subunits. Elife 6, e26147 (2017).
Dembla, S. et al. Anti-nociceptive action of peripheral mu-opioid receptors by G-beta-gamma protein-mediated inhibition of TRPM3 channels. Elife 6, e26280 (2017).
Quallo, T., Alkhatib, O., Gentry, C., Andersson, D. A. & Bevan, S. G protein βγ subunits inhibit TRPM3 ion channels in sensory neurons. Elife 6, e26138 (2017).
Zhao, C. & MacKinnon, R. Structural and functional analyses of a GPCR-inhibited ion channel TRPM3. Neuron 111, 81–91.e7 (2023).
Behrendt, M. et al. The structural basis for an on–off switch controlling Gβγ-mediated inhibition of TRPM3 channels. Proc. Natl Acad. Sci. USA 117, 29090–29100 (2020).
Oberwinkler, J., Lis, A., Giehl, K. M., Flockerzi, V. & Philipp, S. E. Alternative splicing switches the divalent cation selectivity of TRPM3 channels. J. Biol. Chem. 280, 22540–22548 (2005).
Held, K. et al. Pharmacological properties of TRPM3 isoforms are determined by the length of the pore loop. Br. J. Pharmacol. 179, 3560–3575 (2022).
Yin, Y. et al. Structure of the cold- and menthol-sensing ion channel TRPM8. Science 359, 237–241 (2018).
Winkler, P. A., Huang, Y., Sun, W., Du, J. & Lu, W. Electron cryo-microscopy structure of a human TRPM4 channel. Nature 552, 200–204 (2017).
Diver, M. M., Cheng, Y. & Julius, D. Structural insights into TRPM8 inhibition and desensitization. Science 365, 1434–1440 (2019).
Naylor, J. et al. Pregnenolone sulphate- and cholesterol-regulated TRPM3 channels coupled to vascular smooth muscle secretion and contraction. Circ. Res. 106, 1507–1515 (2010).
Hirschi, M. et al. Cryo-electron microscopy structure of the lysosomal calcium-permeable channel TRPML3. Nature 550, 411–414 (2017).
Autzen, H. E. et al. Structure of the human TRPM4 ion channel in a lipid nanodisc. Science 359, 228–232 (2018).
Kwon, D. H. et al. Heat-dependent opening of TRPV1 in the presence of capsaicin. Nat. Struct. Mol. Biol. 28, 554–563 (2021).
Majeed, Y. et al. Cis-isomerism and other chemical requirements of steroidal agonists and partial agonists acting at TRPM3 channels. Br. J. Pharmacol. 161, 430–441 (2010).
Yin, Y. et al. Visualizing structural transitions of ligand-dependent gating of the TRPM2 channel. Nat. Commun. 10, 3740 (2019).
Zubcevic, L., Le, S., Yang, H. & Lee, S. Y. Conformational plasticity in the selectivity filter of the TRPV2 ion channel. Nat. Struct. Mol. Biol. 25, 405–415 (2018).
Kwon, D. H., Zhang, F., Fedor, J. G., Suo, Y. & Lee, S. Y. Vanilloid-dependent TRPV1 opening trajectory from cryoEM ensemble analysis. Nat. Commun. 13, 2874 (2022).
Pumroy, R. A. et al. Molecular mechanism of TRPV2 channel modulation by cannabidiol. Elife 8, e48792 (2019).
Zubcevic, L., Borschel, W. F., Hsu, A. L., Borgnia, M. J. & Lee, S. Y. Regulatory switch at the cytoplasmic interface controls TRPV channel gating. Elife 8, e47746 (2019).
Zhao, S., Yudin, Y. & Rohacs, T. Disease-associated mutations in the human TRPM3 render the channel overactive via two distinct mechanisms. Elife 9, e47746 (2020).
Zhao, Y., McVeigh, B. M. & Moiseenkova-Bell, V. Y. Structural pharmacology of TRP channels. J. Mol. Biol. 433, 166914 (2021).
Song, K. et al. Structural basis for human TRPC5 channel inhibition by two distinct inhibitors. Elife 10, e63429 (2021).
Yin, Y. et al. Activation mechanism of the mouse cold-sensing TRPM8 channel by cooling agonist and PIP2. Science 378, eadd1268 (2022).
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360 (1996).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Scheres, S. H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Moriarty, N. W., Grosse-Kunstleve, R. W. & Adams, P. D. Electronic ligand builder and optimization workbench (eLBOW): a tool for ligand coordinate and restraint generation. Acta Crystallogr. D Biol. Crystallogr. 65, 1074–1080 (2009).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Delano, W.L. The PyMol Molecular Graphics System (DeLano Scientific, 2002).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Ren, Z. et al. Structural basis for inhibition and regulation of a chitin synthase from Candida albicans. Nat. Struct. Mol. Biol. 29, 653–664 (2022).
Joyce, L. R. et al. Identification of a novel cationic glycolipid in Streptococcus agalactiae that contributes to brain entry and meningitis. PLoS Biol. 20, e3001555 (2022).
Best, R. B. et al. Optimization of the additive CHARMM all-atom protein force field targeting improved sampling of the backbone φ, ψ and side-chain χ1 and χ2 dihedral angles. J. Chem. Theory Comput. 8, 3257–3273 (2012).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Durell, S. R., Brooks, B. R. & Ben-Naim, A. Solvent-induced forces between two hydrophilic groups. J. Phys. Chem. 98, 2198–2202 (1994).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Wu, E. L. et al. CHARMM-GUI membrane builder toward realistic biological membrane simulations. J. Comput. Chem. 35, 1997–2004 (2014).
Lee, J. et al. CHARMM-GUI input generator for NAMD, GROMACS, AMBER, OpenMM, and CHARMM/OpenMM simulations using the CHARMM36 additive force field. J. Chem. Theory Comput. 12, 405–413 (2016).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Jo, S., Lim, J. B., Klauda, J. B. & Im, W. CHARMM-GUI membrane builder for mixed bilayers and its application to yeast membranes. Biophys. J. 97, 50–58 (2009).
Gao, Y. et al. CHARMM-GUI supports hydrogen mass repartitioning and different protonation states of phosphates in lipopolysaccharides. J. Chem. Inf. Model. 61, 831–839 (2021).
Eastman, P. et al. OpenMM 7: rapid development of high performance algorithms for molecular dynamics. PLoS Comput. Biol. 13, e1005659 (2017).
Yin, Y. et al. MD simulations for “molecular basis of neurosteroid and anticonvulsant regulation of TRPM3”. Zenodo https://doi.org/10.5281/zenodo.13369813 (2024).
Acknowledgements
Cryo-EM data were collected at the Shared Materials Instrumentation Facility (SMIF) at Duke University and the National Cancer Institute (NCI)’s National Cryo-EM facility (NCEF) at the Frederick National Laboratory for Cancer Research. We thank A. Wier at NCEF and N. Bhattacharya at SMIF for assistance with microscope operation. We thank J. Fedor for critical paper reading and S. Yang for help with data collection. This work was supported by the National Institutes of Health (R35NS097241 to S.-Y.L. and R01GM138472 to W.I.), the National Institute of Health Intramural Research Program; National Institute of Environmental Health Sciences (ZIC ES103326 to M.J.B) and by the NCI’s NCEF under contract 75N91019D00024.
Author information
Authors and Affiliations
Contributions
Y.Y. conducted biochemical preparation, cryo-EM experiments, single-particle 3D reconstruction and model building under the guidance of S.-Y.L. C.-G.P. and F.Z. carried out all the electrophysiological recordings under the guidance of S.-Y.L. S.F. carried out all the molecular dynamics simulations under the guidance of W.I. Z.G., H.-J.L. and Y.Y. conducted mass spectrometry experiments. K.S. helped with part of the grid screening under the guidance of M.J.B. S.-Y.L. wrote the paper with input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Functional characterization of TRPM3 channel activation by various agonists.
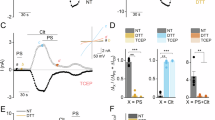
a, Representative currents at –60 mV (lower traces) and +60 mV (upper traces) from HEK293T cells expressing the mTRPM3α2 WT channels (left) and from GFP-only transfected cells (middle), in response to 200 μM PregS (blue), 200 μM Nif (orange) and 1 μM CIM (green) as indicated. Summary (right) of current amplitude in response to 200 μM PregS, 200 μM Nif and 1 μM CIM, respectively, from HEK293T cells expressing mTRPM3α2 WT (filled circles, n = 4 biological replicates; mean ± s.e.m.) and from cells transfected only with GFP (open circles, n = 3 biological replicates; mean ± s.e.m.) at +60 mV. P values from two-way ANOVA followed by Sidak post-hoc test are shown. b, Left: representative currents at –60 mV (lower traces) and +60 mV (upper traces) from HEK293T cells transfected only with GFP, in response to 200 μM PregS (blue) and 1 μM CIM (green) as indicated. Right: Summary of current amplitude in response to 200 μM PregS and 1 μM CIM from HEK293T cells transfected only with GFP at +60 mV (n = 3 biological replicates; mean ± s.e.m.). c, Left: representative traces at –60 mV (lower traces) and +60 mV (upper traces) from HEK293T cells transfected only with GFP, in response to 200 μM PregS (blue) and 200 μM Nif (orange) as indicated. Summary of current amplitude in response to 200 μM PregS and 200 μM Nif from HEK293T cells transfected only with GFP at +60 mV (n = 3 biological replicates; mean ± s.e.m.).
Extended Data Fig. 2 Cryo-EM data processing.
a–c, Cryo-EM data processing workflows, including representative 2D classification images, for the primidone-bound TRPM3 (a), the CHS-bound TRPM3 (b) and the PregS/CIM-bound TRPM3 (c) structures.
Extended Data Fig. 3 Cryo-EM data validation.
a, Representative micrographs with the physical pixel size indicated. b, Fourier shell correlation (FSC) curves of the final 3D reconstruction with different masking calculated in CryoSPARC. c, Local resolution estimation of the final 3D reconstruction. d, Orientation distribution diagram of the final 3D reconstruction. e, FSC curves between model and full- or half-maps calculated in PHENIX. Results are shown for the primidone-bound TRPM3 (left), the CHS-bound TRPM3 (middle) and the PregS/CIM-bound TRPM3 (right) structures.
Extended Data Fig. 4 Quality of cryo-EM maps.
a–c, Representative cryo-EM densities (meshes) for various structural elements in the primidone-bound TRPM3 (a, green), the CHS-bound TRPM3 (b, light blue) and the PregS/CIM-bound TRPM3 (c, magenta) structures. Residue ranges are specified in parentheses. d, EM densities (meshes) for primidone, CHS1 and CHS2 in the primidone-bound TRPM3 reconstruction (green), CHS1 and CHS2 in the CHS-bound TRPM3 reconstruction (light blue), CIM, PregS1 and PregS2 in the PregS/CIM-bound TRPM3 reconstruction (magenta).
Extended Data Fig. 5 Architecture of the TRPM3 channel and comparison with the published TRPM3 structure.
a, Structure model of the CHS-bound TRPM3 with a three-layer architecture. Each layer is labeled as transmembrane domain (TMD), top and bottom layers of the cytosolic domain (CD top and CD bottom). Protomers are colored blue, green, gold and red. b,c, Topology (b) and structure (c) of a single TRPM3 protomer. d, Comparisons of the TMDs between the published TRPM3 structure (PDB 8ED7) and the CHS-bound TRPM3 structure (left) and between the published TRPM3 structure and the PregS/CIM-bound TRPM3 structure (right). e, Side-by-side comparison of the published TRPM3 (PDB 8ED7), the CHS-bound TRPM3 and the PregS/CIM-bound TRPM3 structures.
Extended Data Fig. 6 Functional characterization of TRPM3.
a, Representative currents of the mutant mTRPM3α2 channels in HEK293T cells at –60 mV (lower traces) and +60 mV (upper traces) in response to 100 μM PregS (blue), 100 μM primidone (gold) and 50 μM ruthenium red (RR) (red) as indicated. b, Representative TEVC recording on the WT and mutant mTRPM3α2 channels in oocytes at –60 mV (lower traces) and +60 mV (upper traces) in response to 200 μM PregS (blue), 200 μM primidone (gold) and 50 μM RR (red) as indicated. c, Summary of current inhibition by 200 μM primidone in WT and mutant mTRPM3α2 channels at +60 mV (n = 8,7,6,8,8 and 8 biological replicates, respectively; mean ± s.e.m.). P values from one-way ANOVA followed by Dunnett’s post-hoc test for differences between WT and each mutant are shown. d, Representative currents of the mutant mTRPM3α2 channels in HEK293T cells at –60 mV (lower traces) and +60 mV (upper traces) in response to 200 μM Nif (orange) and 200 μM PregS (blue) as indicated. e, Representative currents of the mutant mTRPM3α2 channels in HEK293T cells at –60 mV (lower traces) and +60 mV (upper traces) in response to 200 μM PregS (blue), 200 μM Nif (orange) and 1 μM CIM (green) as indicated. Dotted lines in a,b,d,e indicate the zero-current level.
Extended Data Fig. 7 Identification of lipids and small molecules bound to TRPM3 by liquid chromatography–high-resolution mass spectrometry analysis.
a, Chemical structures and expected exact masses of pregnenolone sulfate (PregS), cholesteryl hemisuccinate (CHS) and CIM 0216 (CIM). b,c, PregS is detected in the +PregS sample (b) but not in the -PregS sample (c). d,e, PregS is detected in the +PregS sample (d) and CHS is detected in the -PregS sample (e). f,g, CHS is not detected in the +PregS sample (f) but is detected in the -PregS sample (g). h,i, CIM is not detected in the control sample (h) but is detected in the +CIM sample (i). Inset: mass spectrum showing the [M + H]+ and [M+Na]+ ions of CIM at m/z 348.176 and m/z 370.155, respectively. Normal LC–MS analysis of PregS was performed in the negative ion mode. Reverse LC–MS analysis of CIM was performed in the positive ion mode. EIC, extracted ion chromatogram.
Extended Data Fig. 8 Comparison of ligand binding at the outer pore in TRP channels.
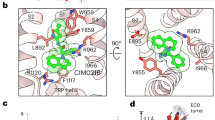
a, Cholesteryl hemisuccinate (CHS) (yellow, left) and pregnenolone sulfate (PregS) (purple, right) binding at site 1 and site 2 in TRPM3 channel identified in the current study. PregS at site 2 is shown as semi-transparent. b, CHS and phospholipid binding in the TRPML3 channel. c, Binding of the detergent glyco-diosgenin (GDN) in TRPM3 (left, yellow), CHS in TRPM4 (middle, yellow) and CHS in TRPM8 (right, yellow). d, Binding of the inhibitor HC-070 (left, green) and the putative diacylglycerol (DAG) (middle, yellow) in the TRPC5 channel. Binding of the agonist AM-0883 (right, green) in the TRPC6 channel. PDB IDs for the published structures in b–d are indicated. Ligands are shown as spheres, with agonists/inhibitors colored green, CHS/phospholipids/GDN/DAG colored yellow and PregS colored purple.
Extended Data Fig. 9 All-atom molecular dynamics simulation studies of TRPM3.
a, Conformational ensemble of PregS, CHS and cholesterol binding at site 1 and site 2, with twelve replicas shown. b,d, The starting configurations of CIM ligand in pose 1 (b) and pose 2 (d), with a schematic showing the orientation of the three ring systems. c,e, Conformational ensemble overlay of the CIM ligand in pose 1 (c) and pose 2 (e) after 1200-ns simulation. Twelve replicas (3 replicas x 4 subunits) are overlaid with two views shown. Pose 1 leads to more consistent ligand-binding configurations, as indicated by the well aligned oxazole and amide. f, Whole-molecule ligand r.m.s.d. of CIM in pose 1 and pose 2. g, Oxazole ring-based ligand r.m.s.d. of CIM in pose 1 and pose 2. The difference is salient between the two poses, as the ligand r.m.s.d. is more stable in pose 1, consistent with the structural overlay in (c).
Extended Data Fig. 10 Potential ion permeation pathways within VSLDs and disease mutation mapping on the TRPM3 structures.
a, Dot surface representations of the potential ion permeation pathways within the VSLDs in both the CHS-bound and the PregS/CIM-bound states. Regions inaccessible to water (pore radius <1.15 Å) are colored red. b–e, In silico mutagenesis (cyan) on both the CHS-bound and the PregS/CIM-bound TRPM3 structures. (b,c) Both L759 (L769 in human, pre-S1) and V992 (V1002 in human, S4b) interact with the TRP domain. However, the changes are subtle, making the increased activity of these mutations not entirely clear. (d) P1092 (P1102 in human) is located in S6 at the outer pore interface, comprising part of the CHS (or cholesterol) site 2 in the CHS-bound or the primidone-bound TRPM3 structures. In the PregS/CIM-bound TRPM3 structure, S6 rotates to position P1092 away from the PregS site 2 to form an interaction with E1057 on PHb. Therefore, P1092Q likely destabilizes the apo (closed) state by reducing cholesterol binding and stabilizes the PregS/CIM-bound conformation through interaction with PHb. CHS at site 2 of the CHS-bound structure is shown as yellow sticks (left panels). (e) N1116 (N1126 in human) is located toward the C-terminus of S6, adjacent to the neighboring S6. N1116D can interact electrostatically with K1122′ of the neighboring S6 only in the PregS/CIM-bound conformation. The salt bridge interaction between N1116D and K1122′ in the PregS/CIM-bound state is shown in the PregS/CIM N1116D panel.
Supplementary information
Supplementary Table 1 (download XLSX )
Primers used in this study.
Source data
Source Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Fig. 2 (download XLSX )
Statistical source data.
Source Data Fig. 3 (download XLSX )
Statistical source data.
Source Data Fig. 4 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 1 (download XLSX )
Statistical source data.
Source Data Extended Data Fig. 6 (download XLSX )
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, Y., Park, CG., Feng, S. et al. Molecular basis of neurosteroid and anticonvulsant regulation of TRPM3. Nat Struct Mol Biol 32, 828–840 (2025). https://doi.org/10.1038/s41594-024-01463-8
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41594-024-01463-8
This article is cited by
-
Structural basis for agonist and heat activation of nociceptor TRPM3
Nature Structural & Molecular Biology (2026)
-
A duo of redox-sensitive pore-loop cysteines controls the activity of the neural ion channel TRPM3
Communications Chemistry (2026)
-
A single allosteric site merges activation, modulation and inhibition in TRPM5
Nature Chemical Biology (2026)
-
Direct crosstalk between GPCRs and ion channels via G proteins
Experimental & Molecular Medicine (2025)
-
Visualizing insecticide control of insect TRP channel function and assembly
Nature Communications (2025)