Abstract
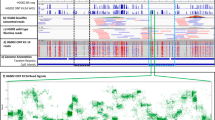
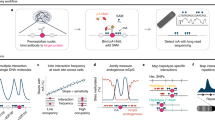
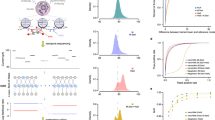
We recently developed directed methylation with long-read sequencing (DiMeLo-seq) to map protein–DNA interactions genome wide. DiMeLo-seq is capable of mapping multiple interaction sites on single DNA molecules, profiling protein binding in the context of endogenous DNA methylation, identifying haplotype-specific protein–DNA interactions and mapping protein–DNA interactions in repetitive regions of the genome that are difficult to study with short-read methods. With DiMeLo-seq, adenines in the vicinity of a protein of interest are methylated in situ by tethering the Hia5 methyltransferase to an antibody using protein A. Protein–DNA interactions are then detected by direct readout of adenine methylation with long-read, single-molecule DNA sequencing platforms such as Nanopore sequencing. Here we present a detailed protocol and practical guidance for performing DiMeLo-seq. This protocol can be run on nuclei from fresh, lightly fixed or frozen cells. The protocol requires 1–2 d for performing in situ targeted methylation, 1–5 d for library preparation depending on desired fragment length and 1–3 d for Nanopore sequencing depending on desired sequencing depth. The protocol requires basic molecular biology skills and equipment, as well as access to a Nanopore sequencer. We also provide a Python package, dimelo, for analysis of DiMeLo-seq data.
Key points
-
DiMeLo-seq uses long-read, single-molecule sequencing to map protein–DNA interactions genome wide, in nuclei from fresh, fixed or frozen cells, and from primary tissues or intact organisms.
-
Compared with short-read methods, this enables mapping of multiple interaction sites on single DNA molecules, profiling protein binding in the context of endogenous DNA methylation, identifying haplotype-specific protein–DNA interactions and mapping protein–DNA interactions in repetitive regions of the genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$32.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to the full article PDF.
USD 39.95
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
Data generated for this protocol: raw sequencing data are available in the Sequence Read Archive (SRA) under BioProject accession PRJNA855257 and processed data are available on Gene Expression Omnibus (GEO) under accession GSE208125. All raw fast5 sequencing data from the accompanying Altemose et al. manuscript are available in the SRA under BioProject accession PRJNA752170. External data sources used in this protocol: H3K27ac ChIP-seq data in GM12878 available from ENCODE Project Consortium under accession ENCFF218QBO (https://www.encodeproject.org/files/ENCFF218QBO/). H3K27me3 ChIP-seq data in GM12878 available from ENCODE Project Consortium under accession ENCFF119CAV (https://www.encodeproject.org/files/ENCFF119CAV/). H3K4me3 ChIP-seq data in GM12878 available from ENCODE Project Consortium under accession ENCFF228TWF (https://www.encodeproject.org/files/ENCFF228TWF/). H3K27ac CUT&Tag data in GM12878 available on Gene Expression Omnibus (GEO) under accession GSM5530639. H3K27me3 CUT&Tag data in GM12878 available on GEO under accession GSM5530673. ATAC-seq data in GM12878 available from ENCODE Project Consortium under accession ENCFF603BJO (https://www.encodeproject.org/files/ENCFF603BJO/). TSS and gene annotations from NCBI RefSeq downloaded from UCSC Genome Browser (https://genome.ucsc.edu/cgi-bin/hgTrackUi?g=refSeqComposite&db=hg38). RNA-seq data in GM12878 available from ENCODE Project Consortium under accession ENCFF978HIY (https://www.encodeproject.org/files/ENCFF978HIY/). D. melanogaster H3K9me3 ChIP-seq data available on GEO under accession GSE140539. File GSE140539_H3K9me3_sorted_deepnorm_log2_smooth.bw was used.
Code availability
The dimelo Python package for analysis of DiMeLo-seq data is available on Github: https://github.com/streetslab/dimelo.
References
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat. Methods 4, 651–657 (2007).
Johnson, D. S., Mortazavi, A., Myers, R. M. & Wold, B. Genome-wide mapping of in vivo protein–DNA interactions. Science 316, 1497–1502 (2007).
Barski, A. et al. High-resolution profiling of histone methylations in the human genome. Cell 129, 823–837 (2007).
Kaya-Okur, H. S. et al. CUT&Tag for efficient epigenomic profiling of small samples and single cells. Nat. Commun. 10, 1930 (2019).
Skene, P. J., Henikoff, J. G. & Henikoff, S. Targeted in situ genome-wide profiling with high efficiency for low cell numbers. Nat. Protoc. 13, 1006–1019 (2018).
van Steensel, B. & Henikoff, S. Identification of in vivo DNA targets of chromatin proteins using tethered dam methyltransferase. Nat. Biotechnol. 18, 424–428 (2000).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Altemose, N. et al. DiMeLo-seq: a long-read, single-molecule method for mapping protein–DNA interactions genome wide. Nat. Methods 19, 711–723 (2022).
van Schaik, T., Vos, M., Peric-Hupkes, D., Hn Celie, P. & van Steensel, B. Cell cycle dynamics of lamina-associated DNA. EMBO Rep. 21, e50636 (2020).
Stergachis, A. B., Debo, B. M., Haugen, E., Churchman, L. S. & Stamatoyannopoulos, J. A. Single-molecule regulatory architectures captured by chromatin fiber sequencing. Science 368, 1449–1454 (2020).
Shipony, Z. et al. Long-range single-molecule mapping of chromatin accessibility in eukaryotes. Nat. Methods 17, 319–327 (2020).
Abdulhay, N. J. et al. Massively multiplex single-molecule oligonucleosome footprinting. eLife 9, e59404 (2020).
Lee, I. et al. Simultaneous profiling of chromatin accessibility and methylation on human cell lines with nanopore sequencing. Nat. Methods 17, 1191–1199 (2020).
Wang, Y. et al. Single-molecule long-read sequencing reveals the chromatin basis of gene expression. Genome Res. 29, 1329–1342 (2019).
Weng, Z. et al. BIND&MODIFY: a long-range method for single-molecule mapping of chromatin modifications in eukaryotes. Genome Biol. 24, 61 (2023).
Yue, X. et al. Simultaneous profiling of histone modifications and DNA methylation via nanopore sequencing. Nat. Commun. 13, 1–14 (2022).
Statham, A. L. et al. Bisulfite sequencing of chromatin immunoprecipitated DNA (BisChIP-seq) directly informs methylation status of histone-modified DNA. Genome Res. 22, 1120–1127 (2012).
Brinkman, A. B. et al. Sequential ChIP-bisulfite sequencing enables direct genome-scale investigation of chromatin and DNA methylation cross-talk. Genome Res. 22, 1128–1138 (2012).
Gamba, R. et al. Enrichment of centromeric DNA from human cells. PLoS Genet. 18, e1010306 (2022).
Payne, A. et al. Readfish enables targeted nanopore sequencing of gigabase-sized genomes. Nat. Biotechnol. 39, 442–450 (2021).
Kovaka, S., Fan, Y., Ni, B., Timp, W. & Schatz, M. C. Targeted nanopore sequencing by real-time mapping of raw electrical signal with UNCALLED. Nat. Biotechnol. 39, 431–441 (2021).
Hoffman, E. A., Frey, B. L., Smith, L. M. & Auble, D. T. Formaldehyde crosslinking: a tool for the study of chromatin complexes. J. Biol. Chem. 290, 26404–26411 (2015).
Vogel, M. J., Peric-Hupkes, D. & van Steensel, B. Detection of in vivo protein-DNA interactions using DamID in mammalian cells. Nat. Protoc. 2, 1467–1478 (2007).
Altemose, N. et al. DiMeLo-Seq: directed methylation with long-read sequencing v2. Protocols.io https://www.protocols.io/view/dimelo-seq-directed-methylation-with-long-read-seq-b2u8qezw (2021).
De Coster, W., Stovner, E. B. & Strazisar, M. Methplotlib: analysis of modified nucleotides from nanopore sequencing. Bioinformatics 36, 3236–3238 (2020).
Jain, M. et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 36, 338–345 (2018).
Brothers, M. & Rine, J. Distinguishing between recruitment and spread of silent chromatin structures in Saccharomyces cerevisiae. eLife 11, e75653 (2022).
Srinivasan, M., Sedmak, D. & Jewell, S. Effect of fixatives and tissue processing on the content and integrity of nucleic acids. Am. J. Pathol. 161, 1961–1971 (2002).
Altemose, N. et al. PA-Hia5 protein expression and purification v1. Protocols.io https://www.protocols.io/view/pa-hia5-protein-expression-and-purification-bv82n9ye (2021).
Altemose, N. et al. AlphaHOR-RES: a method for enriching centromeric DNA v1. Protocols.io https://www.protocols.io/view/alphahor-res-a-method-for-enriching-centromeric-dn-bv9vn966 (2021).
Kim, B. Y. et al. Highly contiguous assemblies of 101 drosophilid genomes. eLife 10, e66405 (2021).
Kim, B. Y., Miller, D. E. & Wang, J. DNA extraction and nanopore library prep from 15-30 whole flies v1. Protocols.io https://www.protocols.io/view/dna-extraction-and-nanopore-library-prep-from-15-3-bdfqi3mw (2021).
Luo, Y. et al. New developments on the Encyclopedia of DNA Elements (ENCODE) data portal. Nucleic Acids Res. 48, D882–D889 (2020).
Zhao, L. et al. FACT-seq: profiling histone modifications in formalin-fixed paraffin-embedded samples with low cell numbers. Nucleic Acids Res. 49, e125 (2021).
Karlić, R., Chung, H.-R., Lasserre, J., Vlahovicek, K. & Vingron, M. Histone modification levels are predictive for gene expression. Proc. Natl Acad. Sci. USA 107, 2926–2931 (2010).
Cedar, H. & Bergman, Y. Linking DNA methylation and histone modification: patterns and paradigms. Nat. Rev. Genet. 10, 295–304 (2009).
Zenk, F. et al. HP1 drives de novo 3D genome reorganization in early Drosophila embryos. Nature 593, 289–293 (2021).
Acknowledgements
Research reported in this publication was supported by the Chan Zuckerberg Biohub, San Francisco, and by the National Human Genome Research Institute and the National Institute of General Medical Sciences of the National Institutes of Health under award number R01HG012383 to A.S., R01GM074728 to A.F.S. and R35GM139653 to G.K. A.M. is supported by a NSF GRFP award. L.D.B. is supported by Volkswagen Stiftung (98196). N.A. is an HHMI Hanna H. Gray Fellow. A.S. is a Chan Zuckerberg Biohub Investigator and a Pew Scholar in the Biomedical Sciences. The sequencing was carried out by the DNA Technologies and Expression Analysis Core at the UC Davis Genome Center, supported by NIH Shared Instrumentation Grant 1S10OD010786-01. This project has been made possible in part by grant number 2022-253563 from the Chan Zuckerberg Initiative DAF, an advised fund of Silicon Valley Community Foundation.
Author information
Authors and Affiliations
Contributions
A.M., N.A. and A.S. designed the study. A.M., N.A. and L.D.B. performed the experiments. A.M., R.M. and J.M. developed dimelo software package. A.M. and N.A. analyzed and interpreted the data. A.M., N.A. and R.M. made the figures. A.M. and J.M. wrote the manuscript, with input from N.A., R.M., L.D.B., K.S., G.K., A.F.S. and A.S. A.S. and N.A. supervised the study.
Corresponding author
Ethics declarations
Competing interests
N.A., A.M., K.S., A.F.S. and A.S. are co-inventors on a patent application related to this work. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
Altemose, N. et al. Nat. Methods 19, 711–723 (2022): https://doi.org/10.1038/s41592-022-01475-6
Supplementary information
Supplementary Information (download PDF )
Supplementary Methods and Table 1.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Maslan, A., Altemose, N., Marcus, J. et al. Mapping protein–DNA interactions with DiMeLo-seq. Nat Protoc 19, 3697–3720 (2024). https://doi.org/10.1038/s41596-024-01032-9
Received:
Accepted:
Published:
Version of record:
Issue date:
DOI: https://doi.org/10.1038/s41596-024-01032-9