Abstract
AI image classification algorithms have shown promising results when applied to skin cancer detection. Most public skin cancer image datasets are comprised of dermoscopic photos and are limited by selection bias, lack of standardization, and lend themselves to development of algorithms that can only be used by skilled clinicians. The SLICE-3D (“Skin Lesion Image Crops Extracted from 3D TBP”) dataset described here addresses those concerns and contains images of over 400,000 distinct skin lesions from seven dermatologic centers from around the world. De-identified images were systematically extracted from sensitive 3D Total Body Photographs and are comparable in optical resolution to smartphone images. Algorithms trained on lower quality images could improve clinical workflows and detect skin cancers earlier if deployed in primary care or non-clinical settings, where photos are captured by non-expert physicians or patients. Such a tool could prompt individuals to visit a specialized dermatologist. This dataset circumvents many inherent limitations of prior datasets and may be used to build upon previous applications of skin imaging for cancer detection.
Similar content being viewed by others
Background & Summary
Algorithms that can distinguish benign from malignant skin lesions with sufficient accuracy could improve triage for skin cancer detection and greatly benefit populations without access to specialized dermatologic care1,2,3,4,5,6,7,8. Current algorithms typically use images captured with a handheld medical device called a dermatoscope9, which uses a magnifying lens and a lighting source to illuminate morphologic features not otherwise visible to the naked eye10,11. Dermatoscopes are used commonly in dermatology clinics as an aid to evaluate skin lesions and diagnose skin cancers such as melanoma (MM)10,12,13, basal cell carcinoma (BCC)14, and squamous cell carcinoma (SCC)15. Dermoscopy has been integrated into the clinical practice worldwide, and a large number of images have been routinely collected for skin cancer diagnosis and monitoring. As a result, most state-of-the-art open-source skin lesion datasets for training AI models consist of dermoscopy images16,17,18. Over the past several years, researchers have explored how clinicians may benefit from utilizing dermoscopy-based AI algorithms19,20,21,22. However, determining which individuals should see a clinician in the first place has great potential impact23. Triaging applications have a significant potential to benefit underserved populations24,25 and improve early skin cancer detection, the key factor in long-term patient outcomes26,27. Similarly, decreasing unnecessary referrals can decrease delays in treating patients in true need of care and reduce burden on health systems28,29. With this work, we have published the SLICE-3D (“Skin Lesion Image Crops Extracted from 3D TBP”) dataset of 400,000+ standardized, de-identified, and diagnostically labeled skin images relevant to use-cases outside of specialized clinics. The image quality resembles cropped smartphone photos, which are regularly submitted by patients to their clinicians for telehealth purposes.
Contributors of this dataset are members of the International Skin Imaging Collaboration (ISIC), an international academia and industry partnership designed to reduce skin cancer morbidity and mortality through the development and use of digital skin imaging applications. ISIC engages both the dermatology and computer vision communities and works to achieve its goals by developing standards and guidelines to improve the quality, privacy, and interoperability for digital skin imaging, by making available a large and expanding open-source archive of quality labeled skin images, and by holding machine learning Grand Challenges for the computer science community in association with leading computer vision conferences. Since 2016, ISIC has hosted 5 Grand Challenges that have all been focused on developing AI for diagnostic classification using dermoscopy images30,31,32,33. Reader studies have demonstrated that top-performing algorithms outperform the average dermatologist on single image diagnosis in preselected lesions and limited diagnostic classes34,35. One prospective study tested a winning algorithm prospectively in the hands of real clinicians and in front of real patients19. However, these applications are confined to settings that involve the presence of a dermatologist at the time of photo capture.
Algorithms trained on more readily accessible, lower quality image types could be used by non-experts, especially in areas lacking specialized dermatologic care, for purposes of triaging concerning lesions and improving early detection8. Such an application might also benefit dermatologists who practice teledermatology, which has become quite common since COVID-1923,36,37,38,39, during which patients submit smartphone images of their skin24. Therefore, there is a need for algorithms to be robust with respect to image quality40,41. The objective for this data descriptor is to publicize a large, novel dataset and to facilitate the development of open-source AI algorithms capable of rendering diagnostic decisions from reduced quality, clinical photos resembling the resolution of smartphone images.
The images in this dataset are extracted from 3D total body photographs (3D TBP). 3D TBP is an emerging standardized imaging modality in dermatologic practice that uses a series of DSLR cameras fixed in an apparatus to capture the complete visible cutaneous surface area in one macro-quality resolution tomographic image42,43,44. Canfield Scientific, an imaging system vendor and manufacturer of a 3D TBP product45, has a proprietary AI-based software that identifies individual lesions on a given 3D capture46. This allows for the image capture and identification of all lesions on a patient, which are exportable as individual cropped photos from each lesion’s most orthogonal DSLR sensor. The SLICE-3D dataset represents every lesion from a sample of 1,000+ patients seen between 2015 and 2024 across seven high-risk dermatologic practices and three continents. Conversely, dermatologists normally use digital dermoscopy to document the more atypical lesions such as those that undergo biopsy or short-term monitoring. Utilizing 3D TBP helps circumvent the lesion-selection bias inherent to routinely collected dermoscopy image datasets47, where the ordinary benign examples tend to be underrepresented. Consequences from utilizing algorithms that have not been sufficiently trained on common benign lesions are uncertain.
Benign moles on an individual tend to resemble each other in terms of color, shape, size, and pattern while outlier lesions are more likely to be melanoma is an observation known as the “ugly duckling sign”48,49,50. Patient-level context is a relevant clinical consideration in addition to analysis of individual lesions for features predictive of malignancy. However, most skin lesion classification algorithms are trained for independent image analysis51. ISIC previously published a patient-centric dataset containing multiple dermoscopy images from a set of patients18, but the lesion selection bias mentioned above remained a limitation. The dataset presented here is novel because it represents each individual’s lesion phenotype more completely. This might be helpful in developing algorithms that utilize the full patient-level context.
This dataset is comprised of standardized images captured using a consistent device model and field of view (FOV). While the image resolution and detail content resemble photos captured with smartphone cameras, patient-captured images vary greatly in lighting and FOV. Standardization helps to mitigate biases that arise from differences in capture conditions specific to the image source. This is a common trait in multisource dermoscopic datasets, which typically contain source-related variations in color, lighting, and vignetting33. When algorithms overfit in training to features unrelated to lesion morphology, they may generalize poorly to unseen populations. Standardized photos in training may prevent overfitting and help produce robust algorithms.
Methods
VECTRA WB360 imaging system
VECTRA WB360 is a 3D TBP image capture system and processing software developed by Canfield Scientific that is used for documenting skin disorders. The physical device captures the entire exposed body efficiently in a single capture using an imaging pod equipped with 92 fixed cameras (46 stereo-pairs of cameras) and xenon flashes capable of both crossed-polarized and non-polarized lighting. The fully integrated DermaGraphix software allows clinicians to tag (put on record) and monitor lesions within a secure database, which can be associated with pathology reports. Clinicians can manually set tagged lesions to one of several statuses, including “biopsied/excised”. The Lesion Visualizer52 (LV) research tool for DermaGraphix uses AI algorithms to automatically locate lesions throughout the 3D TBP capture and estimates a set of measures for each lesion including size, shape, color, “nevus confidence”43, and asymmetry, among others. Hence, lesions are identified in one of two ways; (1) manual lesion tagging performed by the human, often to attribute other clinical and dermoscopy photos to the lesion; (2) automated lesion detection performed by the LV. Both forms assign a 3D coordinate to the 3D TBP image, which then guides cropping in an automated fashion.
Lesion-image “tile” cropping
The ISIC2024 Tile Export Tool was developed by Canfield Scientific for the purpose of extracting standardized, de-identified, single-image photos from individual 3D TBP captures. The cropping method offers a solution to sharing TBP data for research while maintaining patient privacy and confidentiality53,54,55. The tool is capable of batch processing a list of patients. Then, for each 3D TBP capture, the tool first compares the coordinate proximities of each manual tag with each automatically detected lesion to merge pairs that represent the same lesion. Each lesion is thus represented once per 3D capture. For each lesion in the 3D TBP capture, the optimal source photo is selected as that which contains the most orthogonal view of the lesion in order to provide the least distortion in the crop. 15mm-by-15mm field-of-view cropped images centered on each lesion are exported for automatically detected lesions larger than 2.5 mm and for all manually tagged lesions regardless of size. These images are here referred to as “tiles.” Metadata files are also exported and used to associate these tiles to the 3D TBP from which each such image was derived, and include data derived from the LV software and manually tagged lesion statuses. Figure 1 shows a 2D-projection of a portion of a 3D TBP next to examples of tile images with their associated dermoscopy photos. No image preprocessing was performed on the tile images after being extracted with the ISIC2024 Tile Export Tool.
Examples of image types. The SLICE-3D dataset is comprised of tile images, which were extracted from 3D TBP images. Tile images display fewer morphologic features than dermoscopic images, which are captured in clinical settings to document and monitor specific lesions.
Data collection
Plans for this dataset were initiated by the ISIC Artificial Intelligence working group and were presented at the VECTRA WB360 user group meeting during the 2023 EADV Congress. Users from seven institutions ultimately received ethics approval from their respective institutional review board (IRB) or human research ethics committee (HREC) and contributed data to ISIC for the present dataset: (1) Memorial Sloan Kettering Cancer Center, New York, USA; (2) Hospital Clínic de Barcelona, Barcelona, Spain; (3) the University of Queensland, Brisbane, Australia; (4) Medical University of Vienna, Vienna, Austria; (5) University of Athens, Athens, Greece; (6) Melanoma Institute Australia, Sydney, Australia; and (7) the University Hospital of Basel, Basel, Switzerland.
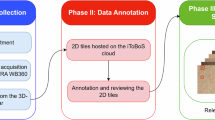
Each site identified patients who had been imaged with 3D TBP between 2015 and 2024. Patients who had previously undergone a skin lesion biopsy or who were known to have been diagnosed with skin cancer were prioritized, although neither were required for inclusion. Electronic medical records (EMR) were reviewed to identify all pathology reports for skin lesion biopsies performed on the included patients. The date of each biopsy, patient- and lesion- identification numbers (from manual tags in DermaGraphix), and coded histologic diagnosis labels were recorded. The data collection process is summarized in panel 1 of Fig. 2. Tile data was exported in bulk at each site using the ISIC2024 Tile Export Tool.
Schematic workflow of dataset curation.
Tile sub-selection
Tiles were then sub-selected according to an algorithm, which is summarized in panel 2 of Fig. 2. The term “strong-labels” refers to tiles associated with a pathology report for a primary biopsy performed within 3 months of the 3D TBP capture, while “weak-labels” refers to tiles assumed benign based on clinical evaluation, which were not associated with a reported biopsy. Tiles associated with a pathology report prior to or after 3 months of imaging were excluded. Also excluded were tiles that were not associated with a pathology report but whose status was set to “biopsied/excised”, as their diagnostic label could not be found.
Weak-label tiles were selected from each patient’s most recent 3D TBP capture containing examples of manual tags. The newest 3D TBP capture was chosen if the patient had no manual tags. When multiple qualifying strong labeled tiles were available for a given biopsied lesion (due to multiple 3D TBPs) one was selected at random.
Quality assurance
Protective health information (PHI) was scraped from metadata files and replaced with coded patient identifiers. All tile images underwent a manual review process in which examples of PHI were flagged and removed, in addition to occasionally included examples of non-lesions (e.g., knuckles, nipples, bellybuttons, tattoos) that were erroneously identified by the lesion detection software. Images underwent two duplicate detection measures. First, exact pixel information was compared across all possible image pairs to confirm that no exact duplicates were present. As a second measure, near duplicates to the strong-labeled images were identified and removed. All image embeddings were extracted using the MONET model published by Kim et al.56. Every strong-labeled image and its weak-labeled nearest neighbor were reviewed. If both images represented the same lesion, the weak-labeled example was excluded. Near-duplicates were removed in less than 5% of pairs and could occur in instances where the software failed to merge the manual tag with the automatically detected lesions based on 3D coordinate proximities due primarily to imprecise manual tag placement.
Obtaining ethics approvals for data sharing
Under the ISIC terms-and-conditions57, a contributor agrees to upload only images to which they have been duly authorized by their own institution to transit and license to data users in accordance with the ISIC terms and one of three Creative Commons licenses selected by the contributor: CC-0, CC-BY, or CC-BY NC. Accordingly, the data from each institution has been discretely licensed for use in this dataset in accordance with what is permitted by the respective institutional IRB and/or HREC.
Memorial Sloan Kettering Cancer Center, under IRB protocol 16–974, contributed anonymized data collected under clinical image capture and consent was waived due to the anonymized nature of the images. This data subset is provided under CC-BY.
Data from the Department of Dermatology, Hospital Clínic, University de Barcelona were compiled from patients who previously had given written consent for the use of anonymized images for clinical and scientific purposes. The research ethics committee of the Hospital Clínic de Barcelona approved (HCB/2023/0213) the licensing of their data for inclusion in the dataset. This data subset is provided under CC-BY NC.
Images from the University of Queensland Dermatology Research Centre were collected from a subset of participants from two studies conducted at the Princess Alexandria Hospital, Brisbane, Australia who provided written informed consent to share de-identified images through open access research databases. The two studies, ‘Mind Your Moles’ Study58 and Health Outcomes Programs Study45, were approved by Metro South Health HREC (HREC/16/QPAH/125 and HREC/17/QPAH/816) and University of Queensland HREC (2016000554 and 2018000074), respectively. The ‘Mind Your Moles’ study58 was additionally approved by Queensland University of Technology (Brisbane, Australia) (1600000515), and QIMR Berghofer Medical Research Institute (Herston, Australia) (P2271). This data subset is provided under CC-BY.
The images from the Department of Dermatology, Medical University of Vienna were acquired from total body photographs of patients who were photographed with one of the total body photography systems used at the Department of Dermatology (Vectra-3D, Fotofinder 2D, conventional photography) and who have given their written consent for the use of pseudonymised images for scientific purposes. This was approved by the ethics committee of the Medical University of Vienna, Austria (1996/2023). This data subset is provided under CC-BY NC.
Data from Andreas Sygros Hospital for Cutaneous and Venereal Diseases, University of Athens were compiled from patients who previously had given written consent for the use of anonymized images for clinical and scientific purposes. The research ethics committee of Andreas Sygros Hospital approved the licensing of this data to Memorial Sloan Kettering Cancer Center for inclusion in the dataset through the data transfer agreement MSK.DTA.0000.0582. This data subset is provided under CC-BY.
Melanoma Institute Australia received approval (X20-0241 and 2020/ETH01411: Melanoma Image Annotation and Analysis Collaboration) from the Sydney Local Health District HREC to share data accrued for registered trial ACTRN12619001706167 (ACEMID) approved by the Metro South HREC under protocol HREC/2019/QMS/57206. All participants provided written consent for ACEMID, and separate consent was provided for use of images and other clinical data in research projects. This data subset is provided under CC-0.
Data from the University Hospital Basel were acquired during a clinical trial approved by the local ethics committee in Switzerland (2020-02482) and registered with ClinicalTrials.gov (NCT04605822). All patients who contributed to this dataset provided optional written informed consent for both publication and transferring of their data and images to other databases for analysis in Switzerland and abroad. This data subset is provided under CC-BY NC.
Data Records
Data accessibility
Note that the project contains two versions of the dataset: SLICE-3D and SLICE-3D Permissive. Both versions are stored on the ISIC Archive. SLICE-3D, comprising all data from the seven institutions, is permanently available at https://doi.org/10.34970/2024-slice-3d59. Note that SLICE-3D is released under a Creative Commons non-commercial attribution (CC-BY NC) license due to the agreed licensing terms and conditions of the providing institutions. Four of seven institutions provided data under either an attribution license (Memorial Sloan Kettering Cancer Center, the University of Queensland, and the University of Athens; CC-BY) or no rights reserved (Melanoma Institute Australia; CC-0), which are not limited to non-commercial use. This subset, called SLICE-3D Permissive, is distinctly accessible at https://doi.org/10.34970/2024-slice-3d-permissive60 in perpetuity and is released under the CC-BY license. Future changes to the ISIC Archive will not affect the versions of the dataset, as described here, accessible at the listed DOIs.
Record description
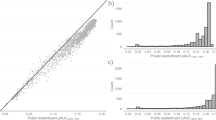
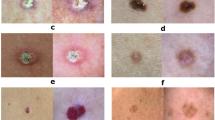
This dataset consists of diagnostically labeled images (tiles). Each example consists of a.jpg file and associated metadata, including a diagnosis label and relevant attributes, patient sex and age at time of imaging, anatomic location of the lesion, the lighting modality of the 3D TBP capture, data from the Lesion Visualizer (e.g., estimate of lesion diameter), and a coded patient identification number for attributing separate tiles to the same patient. Tiles that were associated with a manual tag also contain a unique lesion-identification number. More information on each metadata field is listed in Table 1. Distributions for the overall dataset are presented in Table 2. Image files depict individual skin lesions with a 15mm-by-15mm field-of-view and average image size of 133px-by-133px. Examples of tiles of diagnostic classes are shown in Fig. 3.
Example tiles of varying diagnostic classes.
The diagnostic labels conform to a consensus taxonomy for skin neoplasms developed in a Delphi process involving dermatologic experts61. While strong-labeled images contain descriptive precise lower-level terms, weak-labeled images are labeled “Benign”, a branch in the first layer of the model.
Technical Validation
Lesions were identified on 3D TBP images in an automatic fashion and exported as individual lesion crops. The existence of a primary lesion in each image was confirmed by visual inspection.
The entire malignant diagnostic class consists of strong-labels because their ground truth was confirmed through retrospective review of pathology reports and histopathology is considered the gold standard for diagnosing skin lesions62,63. The malignant class included lesions diagnosed as either melanoma, basal cell carcinoma, or squamous cell carcinoma within 3 months of 3D TBP capture. Melanoma in situ and invasive melanoma were both coded as melanoma.
The benign diagnostic class contains both strong-labels and weak-labels because, while some were attributable to a pathology report dated within 3 months of 3D TBP capture, most never underwent a skin biopsy. Tiles that were never attributed to a pathology report and were not tagged by the clinician as biopsied/excised were assumed benign NOS. All patients in this dataset were treated per standard of care and the decision to biopsy any given lesion was based on clinician discretion. The vast majority of skin lesions are non-cancerous64. The standard of care when a clinician is presented with a concerning lesion is to perform a biopsy62. The weak-label tiles in this dataset depict lesions that were viewed by a dermatologist during a full body skin exam and were not determined to be concerning.
Usage Notes
The field of view of each tile is constant (15 × 15 mm), but the pixel resolution is variable due to varying angles and distances between the patient’s skin and the DSLR sensors in the apparatus. Users of the dataset should use relative pixel distance (as opposed to absolute pixel distance) when conducting image analysis. In addition, the metadata element clin_size_long_diam_mm approximates the maximum diameter of the lesion. However, this measurement is most robust on lesions with clearly outlined boundaries and less conclusive in presence of sun damaged skin. Like the other metadata fields derived from the LV feature, this measurement is an approximation made with an AI model, which serves as a benchmark.
Each tile is derived from either a cross-polarized light (XP) or a non-polarized white light 3D TBP. Lighting modality is the predominant technical factor influencing variability in hue, tint, tone, and shade between the otherwise standardized images contained in this dataset. Thus, the lighting modality evident in the metadata element tbp_tile_type should be acknowledged.
There are no exact duplicate tiles in the dataset, and each corresponds to a unique lesion. When lesions are closely adjacent, however, they may be visible toward the boundary of one another’s tile. The metadata of each tile corresponds to the point-of-interest lesion present nearest to the center of the frame.
Numerous tiles can be associated to the same patient with the metadata element patient_id. Interaction of patient_id and body location fields such as anatom_site_general, location, or location_simple may be used to fine cluster the dataset for similar contextual analyses observed by clinicians51, such as the ugly duckling sign. Lesions in these clusters tend to exhibit similarities due to patient phenotypes and like sun exposure65.
While effort was made to recruit populations at high risk for skin cancer around the world, it was not possible to document the skin tone of the individuals in this dataset due to the retrospective nature of the collection. Dataset users should be aware of this limitation.
The SLICE-3D dataset was used as the training dataset for the ISIC 2024 Grand Challenge. Additional data from each of the seven contributors (1–7) was collected for a separate test set, not described here, for evaluating results of that competition. Two other sites provided data that was allocated entirely to the test set: (8) Alfred Hospital, Melbourne, Australia; and (9) FNQH Cairns, Westcourt, Australia. While the methods of curating the SLICE-3D dataset and the ISIC 2024 test set were consistent, the test set is beyond the scope of this data descriptor.
Code availability
Custom generated code used by all sites for collating the raw data from the ISIC2024 Tile Export Tool, which is described in the Methods section, is available at https://github.com/ISIC-Research/2024-challenge-dataset. All code is written in Python 3.10 and utilizes commonly used open-source packages, all of which are available on pypi.org.
References
James, O. T. et al. Artificial intelligence and machine learning algorithms for early detection of skin cancer in community and primary care settings: a systematic review. Lancet Digit. Health 4, e466–e476 (2022).
Beltrami, E. J. et al. Artificial intelligence in the detection of skin cancer. J. Am. Acad. Dermatol. 87, 1336–1342 (2022).
Vaidya, T. et al. Socioeconomic and geographic barriers to dermatology care in urban and rural US populations. J. Am. Acad. Dermatol. 78, 406–408 (2017).
Glazer, A. M. & Rigel, D. S. Analysis of trends in geographic distribution of US dermatology workforce density. JAMA Dermatol. 153, 472 (2017).
Leavitt, E. R. et al. Teledermatology as a tool to improve access to care for medically underserved populations: A retrospective descriptive study. J. Am. Acad. Dermatol. 75, 1259–1261 (2016).
Fludiona, N. et al. Impact of dermatology eConsults on access to care and skin cancer screening in underserved populations: A model for teledermatology services in community health centers. J. Am. Acad. Dermatol. 78, 293–302 (2018).
Peracca, S. B. et al. Implementing teledermatology for rural veterans: an evaluation using the RE-AIM framework. Telemed J E Health 27, 218–226 (2021).
Udrea, A. et al. Accuracy of a smartphone application for triage of skin lesions based on machine learning algorithms. J Eur Acad Dermatol Venereol 34, 648–655 (2020).
Zafar, M. et al. Skin lesion analysis and cancer detection based on machine/deep learning techniques: a comprehensive survey. Life. 13, 146 (2023).
Marghoob, A. A., Usatine, R. P. & Jaimes, N. Dermoscopy for the family physician. Am Fam Physician 88, 441–450 (2013).
Marghoob, A. A. et al. Instruments and new technologies for the in vivo diagnosis of melanoma. J Am Acad Dermatol 49, 777–799 (2003).
Menzies, S. W., Ingvar, C., Crotty, K. A. & McCarthy, W. H. Frequency and morphologic characteristics of invasive melanomas lacking specific surface microscopic features. Arch Dermatol 132, 1178–1182 (1996).
Bafounta, M. L., Beauchet, A., Aegerter, P. & Saiag, P. Is dermoscopy (epiluminescence microscopy) useful for the diagnosis of melanoma? Results of a meta-analysis using techniques adapted to the evaluation of diagnostic tests. Arch Dermatol 137, 1343–1350 (2001).
Menzies, S. W. et al. Surface microscopy of pigmented basal cell carcinoma. Arch Dermatol 136, 1012–1016 (2000).
Wolner, Z. J. et al. Enhancing skin cancer diagnosis with dermoscopy. Dermatol Clin 35, 417–437 (2017).
Tschandl, P., Rosendahl, C. & Kittler, H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci Data 5, 180161 (2018).
Ricci Lara, M. A. et al. A dataset of skin lesion images collected in Argentina for the evaluation of AI tools in this population. Sci Data 10, 712 (2023).
Rotemberg, V., et al. A patient-centric dataset of images and metadata for identifying melanomas using clinical context. Sci Data 8 2021.
Marchetti, M.A. et al. Prospective validation of dermoscopy-based open-source artificial intelligence for melanoma diagnosis (PROVE-AI study). Npj Digit Med 6 (2023).
Tschandl, P. et al. Human–computer collaboration for skin cancer recognition. Nat Med 26, 1229–1234 (2020).
Cerminara, S.E. et al. Diagnostic performance of augmented intelligence with 2D and 3D total body photography and convolutional neural networks in a high-risk population for melanoma under real-world conditions: A new era of skin cancer screening? Eur J Cancer 190 (2023).
Barata, C. et al. A reinforcement learning model for AI-based decision support in skin cancer. Nat Med 29, 1941–1946 (2023).
Gabel, C. K. et al. Use of teledermatology by dermatology hospitalists is effective in the diagnosis and management of inpatient disease. J. Am. Acad. Dermatol. 84, 1547–1553 (2021).
Pearlman, R. L., Brodell, R. T. & Byrd, A. C. Enhancing access to rural dermatological care; the time to start is now. JAMA Dermatol. 158, 725–726 (2022).
Stitzenberg, K. B. et al. Distance to diagnosing provider as a measure of access for patients with melanoma. Arch Dermatol 143, 991–998 (2007).
Pennie, M. L. et al. Melanoma outcomes for Medicare patients: Association of stage and survival with detection by a dermatologist vs a nondermatologist. Arch Dermatol 143, 488–494 (2007).
Jerant, A. F., Johnson, J. T., Sheridan, C. D. & Caffrey, T. J. Early detection and treatment of skin cancer. Am Fam Physician 62, 357–385 (2000).
Cantisani, C. et al. Melanoma detection by non-specialists: an untapped potential for triage? Diagnostics. 12, 2821 (2022).
Yadav, G. et al. A cross sectional survey of population-wide wait times for patients seeking medical vs. cosmetic dermatologic care. PLOS ONE 11, e0162767 (2016).
Marchetti, M. A. et al. Results of the 2016 International Skin Imaging Collaboration International Symposium on Biomedical Imaging challenge: Comparison of the accuracy of computer algorithms to dermatologists for the diagnosis of melanoma from dermoscopic images. J Am Acad Dermatol 78, 270–277.e1 (2018).
Marchetti, M. A. et al. Computer algorithms show potential for improving dermatologists’ accuracy to diagnose cutaneous melanoma: Results of the International Skin Imaging Collaboration 2017. J Am Acad Dermatol 82, 622–627 (2020).
Codella, N. et al. Skin lesion analysis toward melanoma detection 2018: a challenge hosted by the International Skin Imaging Collaboration (ISIC). arXiv. https://doi.org/10.48550/arXiv.1902.03368 (2019).
Combalia, M. et al. Validation of artificial intelligence prediction models for skin cancer diagnosis using dermoscopy images: the 2019 International Skin Imaging Collaboration Grand Challenge. Lancet Digit Health 4, e330–e339 (2022).
Tschandl, P. et al. Comparison of the accuracy of human readers versus machine-learning algorithms for pigmented skin lesion classification: an open, web-based, international, diagnostic study. Lancet Oncol 20, 938–947 (2019).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature 542, 115–118 (2017).
Megna, M. et al. Teledermatology: a useful tool also after COVID-19 era? J Cosmet Dermatol 21, 2309–2310 (2022).
Trinidad, J., Kroshinsky, D., Kaffenberger, B. H. & Rojek, N. W. Telemedicine for inpatient dermatology consultations in response to the COVID-19 pandemic. J Am Acad Dermatol 83, e69–e71 (2020).
Kennedy, J. et al. Dermatologist perceptions of teledermatology implementation and future use after COVID-19: demographics, barriers, and insights. JAMA Dermatol 157, 595–597 (2021).
Gordon, A. et al. Use of asynchronous interprofessional e-consults in cancer patients during the COVID-19 pandemic: current and future role. Telemed J E Health 29, 304–309.
Young, A. T., Xiong, M., Pfau, J., Keiser, M. J. & Wei, M. L. Artificial intelligence in dermatology: a primer. J. Invest. Dermatol. 140, 1504–1512 (2020).
Sangers, T. E. et al. Position statement of the EADV Artificial Intelligence (AI) Task Force on AI-assisted smartphone apps and web-based services for skin disease. J Eur Acad Dermatol Venereol 38, 22–30 (2024).
Ji-Xu, A., Dinnes, J. & Matin, R. N. Total body photography for the diagnosis of cutaneous melanoma in adults: a systematic review and meta-analysis. Br. J. Dermatol. 185, 302–312 (2021).
Betz-Stablein, B. et al. Reproducible naevus counts using 3D total body photography and convolutional neural networks. Dermatology. 238, 4–11 (2021).
Young, A. T. et al. The role of technology in melanoma screening and diagnosis. Pigment Cell Melanoma Res. 34, 288–300 (2020).
Primiero, C. A. et al. Evaluation of the efficacy of 3D total-body photography with sequential digital dermoscopy in a high-risk melanoma cohort: protocol for a randomised controlled trial. BMJ Open 9, e032969 (2019).
Marchetti, M. A. et al. 3D Whole-body skin imaging for automated melanoma detection. J Eur Acad Dermatol Venereol 37, 945–950 (2023).
Wen, D. et al. Characteristics of publicly available skin cancer image datasets: a systematic review. Lancet Digit Health 4, e64–e74 (2022).
Scope, A. et al. The ‘ugly duckling’ sign: agreement between observers. Arch Dermatol 144, 58–64 (2008).
Gaudy-Marqueste, C. et al. Ugly duckling sign as a major factor of efficiency in melanoma detection. JAMA Dermatol 153, 279–284 (2017).
Grob, J. J. & Bonerandi, J. J. The “ugly duckling” sign: identification of the common characteristics of nevi in an individual as a basis for melanoma screening. Arch Dermatol 134, 103–104 (1998).
Primiero, C. A. et al. A narrative review: opportunities and challenges in artificial intelligence skin image analyses using total body photography. J Invest Dermatol https://doi.org/10.1016/j.jid.2023.11.007 (2024).
D’Alessandro, B. Methods and apparatus for identifying skin features of interest. (2021).
Horsham, C., Janda, M., Kerr, M., Soyer, H. P. & Caffery, L. J. Consumer perceptions on privacy and confidentiality in dermatology for 3D total-body imaging. Australas J Dermatol 64, 118–121 (2023).
Oh, Y. et al. Dermatology patient perspectives regarding consent, privacy, security, and identifiability of clinical photography at a tertiary cancer center: A qualitative study. JAAD Int 12, 121–123.
Ghafur, S., Van Dael, J., Leis, M., Darzi, A., Sheikh, A. Public perceptions on data sharing: key insights from the UK and the USA. Lancet Digit Health 2, e444–e446.
Kim, C. et al. Transparent medical image AI via an image-text foundation model grounded in medical literature. Nat. Med. 30, 1154–1165 (2024).
Koh, U. et al. ‘Mind your Moles’ study: protocol of a prospective cohort study of melanocytic naevi. BMJ Open 8, e025857 (2018).
Kurtansky, N. et al. SLICE-3D 2024 Challenge Dataset. International Skin Imaging Collaboration https://doi.org/10.34970/2024-slice-3d (2024).
Kurtansky, N. et al. SLICE-3D 2024 Permissive Challenge Dataset. International Skin Imaging Collaboration https://doi.org/10.34970/2024-slice-3d-permissive (2024).
Scope, A. et al. International Skin Imaging Collaboration-Designated Diagnoses (ISIC-DX): consensus terminology for lesion diagnostic labeling. J Eur Acad Dermatol Venereol https://doi.org/10.1111/jdv.20055 (2024).
Ferry, A. M., Sarrami, S. M., Hollier, P. C., Gerich, C. F. & Thornton, J. F. Treatment of non-melanoma skin cancers in the absence of mohs micrographic surgery. Plast Reconstr Surg Glob Open 8, e3300 (2022).
Ciuciulete, A. R., Stepan, A. E., Andreiana, B. C. & Simionescu, C. E. Non-melanoma skin cancer: statistical associations between clinical parameters. Curr Health Sci J 48, 110–115 (2022).
Tsao, H., Bevona, C., Goggins, W. & Quinn, T. The transformation rate of moles (melanocytic nevi) into cutaneous melanoma: a population-based estimate. Arch Dermatol 139, 282–288 (2003).
Ghiasvand, R. et al. Association of phenotypic characteristics and UV radiation exposure with risk of melanoma on different body sites. JAMA Dermatol 155, 39–49 (2019).
Acknowledgements
Doug Canfield and Lisong Sun of Canfield Scientific, Inc. have provided resources and expertise during development of the ISIC2024 Tile Export Tool. Brian Helba and Dan LaManna of Kitware are the main developers and maintainers of the ISIC Archive, which hosts the data and serves as the sharing platform for each contributor. Rachel Stephenson and Leonid Zaytsev performed data extraction and subselection for University of Sydney and Melanoma Institute Australia. Dr. Noel Codella participated in the ISIC Challenge planning committee meetings that informed plans for the dataset. This work was funded in part by NIH/NCI U24-CA285296, NIH/NCI U24-CA264369, MSK Cancer Center Support Grant/Core Grant (P30 CA008748), and the European Union through project iToBoS (SC1-BHC-06-2020-965221). The 2024 ISIC Grand Challenge is supported in part by Kaggle. Lastly, we are gracious for the clinical staff at each contributing center for their effort in photographing patients with the WB360 and for the patients who contributed images to the dataset.
Author information
Authors and Affiliations
Contributions
N. Kurtansky designed and collated the dataset, wrote the manuscript, and performed quality review. B. D’Alessandro developed the LV software that detects lesions from 3D TBP captures and the ISIC2024 Tile Export Tool. K. Kose and J. Weber assisted in drafting the manuscript and developed the programmatic assessments for duplicate image detection. M. Gillis performed quality review steps. V. Rotemberg and A. Halpern collected data and provided executive leadership on the project. H. Kittler, K. Liopyris, J. Malvehy, V. Mar, L.V. Maul, A. Navarini, V. Rajeswaran, and H.P. Soyer collected and contributed data. B. Betz-Stablein, P. Gottfrois, A. Mothershaw, C. Mueller, T. Rajeswaran, M. Sashindranath, L. Serra-García, and G. Theocharis performed data extraction and collation. S. Cerminara, M.A. Girundi, E. Goessinger, T. Mathew, and A. Mueller recruited and imaged patients. A. Vos imaged patients and collated data. P. Guitera was involved in conceptualizing triage using 3D TBP. V. Jakrot and L. Martin provided administrative support. R. Garcia and A. Saha provided technical support. All authors helped edit the final manuscript.
Corresponding author
Ethics declarations
Competing interests
B. D’Alessandro is an employee of Canfield Scientific, Inc. B. Betz-Stablein is anticipating employment with Canfield Scientific, Inc. H. Halpern receives consultation fees from Canfield Scientific, Inc. H. Kittler has received speaker honoraria from Fotofinder, is an advisor of Fotofinder and AI Medical Technology, and has received license fees from Heine, Casio, MetaOptima, and Barco. H. Kittler has received equipment from Heine, Casio, and DermaMedical. J. Malvehy is co-founder of Athena Tech, scientific advisor of Dermavision, and is chairman of the Task Force of Artificial Intelligence of the EADV. V. Mar has received speaker fees from Novartis, Bristol Myers Squibb, Merck and Janssen, and has participated in Advisory Boards for MSD, L’Oreal, and SkylineDx. L. Martin is funded by the Warwick L Morison Professorship in dermatology. A. Navarini and L.V. Maul received a grant from Canfield Scientific, Inc. for physician salary in a separate study that had no influence on this manuscript. H.P. Soyer is a shareholder of MoleMap N.Z. Limited and e-derm consult GmbH, and undertakes regular teledermatological reporting for both companies. H.P. Soyer is a Medical Consultant for Canfield Scientific, Inc. and a Medical Advisor for First Derm. H.P. Soyer is involved in several committees of the Australiasian College of Dermatology. V. Rotemberg is a consultant for Inhabit Brands, Inc., and receives in kind support from Kaggle and AWS.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kurtansky, N.R., D’Alessandro, B.M., Gillis, M.C. et al. The SLICE-3D dataset: 400,000 skin lesion image crops extracted from 3D TBP for skin cancer detection. Sci Data 11, 884 (2024). https://doi.org/10.1038/s41597-024-03743-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-03743-w
This article is cited by
-
A methodology for developing dermatological datasets: lessons from retrospective data collection for AI-based applications
BMC Medical Research Methodology (2025)
-
A longitudinal dataset of tile and corresponding dermoscopic images with metadata for identifying skin cancers
Scientific Data (2025)
-
Explainable multimodal AI for skin lesion risk prediction via 3D imaging and clinical data
Scientific Reports (2025)
-
New Release of User-Captured Images from the Oregon Health & Science University Melanoma MoleMapper Project
Scientific Data (2025)
-
Improving dataset transparency in dermatologic Artificial Intelligence using a dataset nutrition label
npj Digital Medicine (2025)