Abstract
Advances in theoretical understanding are frequently unlocked by access to large, diverse experimental datasets. Our understanding of olfactory neuroscience and psychophysics remain years behind the other senses, in part because rich datasets linking olfactory stimuli with their corresponding percepts, behaviors, and neural pathways remain scarce. Here we present a concerted effort to unlock and unify dozens of stimulus-linked olfactory datasets across species and modalities under a unified framework called Pyrfume. We present examples of how researchers might use Pyrfume to conduct novel analyses uncovering new principles, introduce trainees to the field, or construct benchmarks for machine olfaction.
Similar content being viewed by others
Background & Summary
Olfaction is critical to the enjoyment of food, the avoidance of danger, emotional memory, and social interaction. Yet 150 years after Helmholtz and Young first developed a working theory of color vision1, and 100 years after Alexander Graham Bell asked, “Can you measure the difference between one kind of smell and another?”2 we do not yet have a theory that relates chemical features of molecules to neural responses or perception. The design and interpretation of future olfactory neuroscience experiments would greatly benefit from a better understanding of olfactory psychophysics. What is the relationship between the physical properties of the stimulus and the percept it evokes? How do the perceptual properties of monomolecular odorants blend when mixed? What is the size, shape, and structure of odor space? What neural features represent olfactory perceptual properties? There are many datasets and models probing these questions, and yet these are still all open questions in olfaction. By curating and aligning datasets, Pyrfume enables a wide range of specific hypotheses about olfactory perception to be tested against a variety of experimental data. By illustrating the strengths, weaknesses, and blind spots in past research, the results of these tests can motivate, guide, and constrain the next generation of experimental and theoretical investigations into the olfactory system.
The tremendous success of data-driven approaches for problems in visual coding and scene analysis have been propelled by the wide availability and accessibility of imaging data, as well as the communal adoption of key datasets for testing and benchmarking3,4. In the computer vision community, for example, the MNIST5 and ImageNet6 datasets are understood to be essential proving grounds for any newly proposed algorithm or coding principle. Additionally, new visual coding theories can be quickly tested and prototyped across a large number of heterogeneous datasets that expose them to different contexts and edge-cases, making models more robust through out-of-sample testing7. Large datasets have been curated to further build upon machine learning algorithms within the molecular space, as well8,9,10. Here, we describe a newly curated collection of >40 olfactory datasets and a new suite of data fetching, management, and curation tools that we believe can create a set of benchmarks for olfactory theories and models to help stimulate a new era of data-driven inquiry in the olfaction community.
In the remainder of this paper, we introduce Pyrfume, an integrated data archive which aims to accelerate inquiry in olfactory science. While the importance of data aggregation in olfaction has been recognized by others, and there have been other efforts on this front, Pyrfume is notable for its breadth and coverage, spanning > 40 odorant-linked datasets in mammalian olfaction including human psychophysics and perception, as well as animal psychophysics, behavior, brain imaging, physiology, and pharmacology. All together, it contains information about more than 20,000 identified odorants.
There are many archives, papers, and search engines with data that are useful to olfactory scientists, but it has been difficult to coordinate structured queries across these, and each alone has limitations pertaining to size, coverage, or accessibility. PubChem11, for example, has detailed chemical information for over 10 M molecules, but has very little olfactory data. The well-studied Dravnieks atlas12 data set has molecules whose perceptual qualities are described with a structured vocabulary amenable to machine learning, but contains data for only 138 unique molecules. The National Geographic Smell Survey13 has odor intensity ratings collected from an impressive 1.4 million people, but only for a total of six odorants. Even for investigators with the motivation and technical acumen to integrate olfactory data from different sources, there are still the additional challenges of scraping and cleaning datasets that are effectively siloed in separate repositories, and which employ idiosyncratic formats for organizing data.
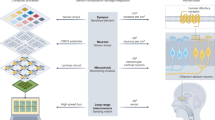
Pyrfume aims to overcome these limitations, and is premised on the simple ideas that: 1) most olfaction experiments are straightforward to index (on the unique identifier (ID) of an olfactory stimulus, e.g. molecules, substances, or mixtures at a given concentration), and 2) any olfactory experiment can be generically described as a machine-readable pairing of such stimulus IDs, the task performed with the corresponding stimuli, and the observed individuals and their behaviors. Linking these experimental components allows for a robust data-formatting standard which is flexible enough to accommodate a wide array of experimental designs and data types, and which conforms to principles of good database design. Note that behavior, as used here and throughout the paper, refers to any experimental measurement. These could be human perceptual ratings applied to a given chemical, glomerular responses observed in mouse physiology experiments, or measured sensitivities of receptors in pharmacology experiments, etc. (Fig. 1).
Overview of the Pyrfume ecosystem. Under the Pyrfume standard, data are always linked to the odor stimuli comprising a given experiment. The features, or experimental measurements, will depend on the particular experiment, but could include data such as amplitudes of glomerular calcium transients, vectorized perceptual descriptors, receptor sensitivities, etc. Any given data archive on Pyrfume (>40 to date) can be easily fetched by using the Pyrfume python and R packages or downloaded directly from GitHub as a zipfile. The orange rows of the odorant x feature matrices indicate odorant molecules, identified with CIDs, common across the experiments. The ability to easily extract data for a given molecule across experimental modalities and model systems is a unique strength of Pyrfume. See Supplementary Figures S1, S2 for examples of this.
Methods
Data collection
There are many archives, papers, and search engines with data that are useful to olfactory scientists, but it has been difficult to coordinate structured queries across these, and each alone has limitations pertaining to size, coverage, or accessibility. In an effort to overcome these limitations, the Pyrfume repository was created to integrate various sources of olfactory data, and format them around a subject-object distinction. Data were derived from online databases and several academic datasets. Table 1 provides a subset of sampled data currently available through Pyrfume.
Reformatting and organization of collected data
Each data source curated in Pyrfume is standardized to conform to a subject/object design framework, separating the odor objects from behavior of the subject(s) under study. At the object level, the most essential file, called stimuli.csv is indexed on a stimulus ID and maps this ID to the chemical/molecular details of the odorants used. A stimulus could represent a single molecule, substance, or mixture, the applied concentration(s), and potentially other experimental conditions. In (typical) cases where at least one stimulus is a single molecule with known structure, the Pyrfume archive will also contain a file, molecules.csv, that lists all molecules used in that dataset, with columns providing PubChem Compound IDs (CIDs), SMILES14, common names, and IUPAC names. This file is useful for indexing the usage of each kind of molecule across datasets, and also for computing physicochemical features for each such molecule (software packages such as RDKit15 and Mordred16 compute these directly from SMILES). When stimuli correspond to mixtures of unknown provenance, e.g. “cloves”, a unique stimulus ID is generated but, in such cases, it may not be possible to link it back to specific compounds in molecules.csv. In cases where a dataset includes calculated or experimentally measured physicochemical properties of molecules, these may be included in physics.csv, typically indexed on CID. Collectively these describe the odorant “object”. The subject side of the data is principally described in behavior.csv, a long-format dataframe, indexed on stimulus ID. This standard, widely referred to as ‘Tidy’, or sometimes ‘Third normal form’17, can easily accommodate more complex, multi-level designs, and reduces both redundancy in data representation, as well as the unnecessary proliferation of files and tables.
Along with these files, each data archive includes a simple, standardized, machine-readable file named manifest.toml, which describes the contents of the archive. The manifest outlines relevant citations and credits, lists both raw and processed data files, and provides essential context or metadata for interpreting the data. It also includes a list of the code used to process the raw data. Additionally, every archive contains a Python script, main.py, that documents the data processing workflow. This script allows Pyrfume users to view how raw data files have been processed and reproduce or modify the processing pipeline. Whenever possible, raw data files under 5 MB in size will also be included directly in the archive, ensuring availability and accessibility.
Data Records
All datasets are available to download at [10.5281/zenodo.13820408]18, and through the companion python library, and python, R, and REST APIs. They can also be accessed directly on GitHub at http://github.com/pyrfume/pyrfume-data. Pyrfume is intended for non-commercial purposes under FAIR use principles, except where licenses permit commercial use. More information, including full documentation and links to source code, can be found at http://pyrfume.org.
File structure within Pyrfume is designed to align multiple curated datasets and optimize them for data parsing. Each data source is structurally organized into separate files, which may include detailed subject, stimuli, and experimental behavior information. In cases where stimuli are administered to distinct subjects, the subject.csv files provide information specific to each subject, including unique identifiers. For studies involving human participants, these files also include demographic details such as race, ethnicity, and age. Testing stimuli are arranged into two files. The first file, referred to as stimuli.csv, contains experiment specific stimulus identifiers, and lists where applicable, PubChem compound identifiers (CIDs), concentrations, ratios, and solvents. This file can include single molecules, mixtures, or unique substances. The second file, known as molecules.csv, provides detailed information about all tested molecules, which may include CIDs, odor names, Chemical Abstracts Service (CAS) numbers, SMILES, and molecular weights. The behavior.csv file contains all observed experimental measures. It is organized by stimulus, subject, experimental measures, and the values of those measures. The measures column specifies the variable being recorded, while its corresponding value is found in the experimental values column. An example of this layout can be seen in Fig. 2.
Schematic showing important data-formatting standards of Pyrfume. In a hypothetical and simplified experiment, 2 odorants were tested on two animals, and data were collected across a total of 5 glomeruli. The Pyrfume archive is organized around a subject-object distinction. The molecules.csv file (lower left) contains chemical descriptors, which can be used to index experimental measurements. The stimuli.csv file (lower right) provides a mapping between stimulus conditions and chemical information. The subjects.csv file (upper left) catalogs all experimental levels for all subjects. Lastly, behavior.csv (upper right) is indexed on the stimuli, and contains the actual experimental measurements, with one measurement per row.
Technical Validation
All curated datasets were derived from a wide body of published and commercial olfactory data and evaluated both prior to compilation in the Pyrfume repository by the corresponding data collectors and/or distributors, and reviewed and cross-checked by the authors of this manuscript.
Usage Notes
In addition to the repository files18, Pyrfume is available to access via [http://github.com/pyrfume]. Pyrfume is a live repository in GitHub and will be updated as new datasets are added.
Sample use cases
The basic Pyrfume ecosystem is described in Figs. 1, 2, and is defined by two ways of interacting with data, which we call bottling and unbottling. Bottling involves investigators readying their experimental data for machine learning applications, and the goal of this process is essentially to make life easy for downstream users (data scientists, computational neuroscientists, machine learning engineers, etc.) through standardization. In a typical workflow, an investigator would compile an inventory of all odorants used in their experiments, and then use the Pyrfume functions get_cids() and from_cids() to programatically create the molecules.csv file. Examples of these functions, and others, can be found in Table 2. Creation of the behavior.csv file is more idiosyncratic to the particular experiment under consideration, but is rarely more than an hour’s work. The critical step, as alluded to above, involves defining the measurements that will comprise individual cell values of the dataframe (e.g. “each cell = peak deltaF/F measured in one glomerulus, for one odorant”, or “each cell = an EC50 value reported for one receptor responding to one odorant”, or “each cell = a perceptual rating applied for one descriptor, for one odorant”, etc). As of the writing of this manuscript, there are >40 bottled experiments, comprising data for >20,000 unique odorants. In addition to data assembled from supplemental materials of published research, Pyrfume contains cleaned and digitized versions of several large databases, which are shown in Table 1.
Pyrfume offers scientists, engineers, and trainees the opportunity to discover, test, and explore the world of olfactory experience through access to an unprecedented volume and diversity of data linked through a standardized format. Standardization allows for cross-modal analyses, meta-analyses, and benchmark construction for the next generation of predictive models, an example of which can be found in Supplementary Figure 2. The next step is up to the broader research community, whom we welcome to utilize this resource and, where applicable, to contribute their own datasets to increase their visibility and utility. In the past, olfactory research has faced significant constraints posed by the scarcity and inadequacy of available data, making it challenging to draw robust conclusions or develop sophisticated theories. As such, we hope that this resource will provide benchmark datasets for a variety of models and raise the bar for theoretical efforts.
Code availability
All code is also available at [https://github.com/pyrfume] or can be accessed using packages which are available for python via pypi (pip install pyrfume) and R via CRAN (install.packages(“rfume”)).
References
Young, T. II. The Bakerian Lecture. On the theory of light and colours. Philos Trans R Soc Lond 92, 12–48 (1802).
Bell, A. G. Discovery and Invention. (Press of Judd & Detweiler, 1914).
Gerkin, R. C. Parsing Sage and Rosemary in Time: The Machine Learning Race to Crack Olfactory Perception. Chem Senses 46 (2021).
Schrimpf, M. et al. Integrative Benchmarking to Advance Neurally Mechanistic Models of Human Intelligence. Neuron 108, 413–423 (2020).
Deng, L. The MNIST database of handwritten digit images for machine learning research. IEEE Signal Process Mag 29, 141–142 (2012).
Deng, J. et al. ImageNet: A large-scale hierarchical image database. 248–255, https://doi.org/10.1109/CVPR.2009.5206848 (2010).
Kearnes, S. Pursuing a Prospective Perspective. Trends Chem 3, 77–79 (2021).
Wu, Z. et al. MoleculeNet: a benchmark for molecular machine learning. Chem Sci 9, 513–530 (2018).
Garg, N. et al. FlavorDB: a database of flavor molecules. Nucleic Acids Res 46 (2018).
Kumar, Y. et al. AromaDb: A database of medicinal and aromatic plant’s aroma molecules with phytochemistry and therapeutic potentials. Front Plant Sci 9 (2018).
The PubChem Project. https://pubchem.ncbi.nlm.nih.gov/.
Dravnieks, A. Atlas of Odor Character Profiles. Atlas of Odor Character Profiles, https://doi.org/10.1520/DS61-EB (1992).
Wysocki, C. J. & Gilbert, A. N. National Geographic Smell Survey. Effects of age are heterogenous. Ann N Y Acad Sci 561, 12–28 (1989).
Weininger, D., Weininger, A. & Weininger, J. L. SMILES. 2. Algorithm for Generation of Unique SMILES Notation. J Chem Inf Comput Sci 29, 97–101 (1989).
RDKit. https://www.rdkit.org/.
Moriwaki, H., Tian, Y. S., Kawashita, N. & Takagi, T. Mordred: a molecular descriptor calculator. J Cheminform 10, 4 (2018).
Wickham, H. Tidy Data. J Stat Softw 59, 1–23 (2014).
Hamel, E. A. et al. Pyrfume: A window to the world’s olfactory data, Zenodo., https://doi.org/10.5281/zenodo.13820408 (2024).
Abraham, N. M., Guerin, D., Bhaukaurally, K. & Carleton, A. Similar Odor Discrimination Behavior in Head-Restrained and Freely Moving Mice. PLoS One 7, 51789 (2012).
Ahmed, L. et al. Molecular mechanism of activation of human musk receptors OR5AN1 and OR1A1 by (R)-muscone and diverse other musk-smelling compounds. Proc Natl Acad Sci USA 115, E3950–E3958 (2018).
Arshamian, A. et al. The perception of odor pleasantness is shared across cultures. Current Biology 32, 2061–2066.e3 (2022).
Bolding, K. A. & Franks, K. M. Recurrent cortical circuits implement concentration-invariant odor coding. Science 361 (2018).
Burton, S. D. et al. Mapping odorant sensitivities reveals a sparse but structured representation of olfactory chemical space by sensory input to the mouse olfactory bulb. 11, 80470 (2022).
Bushdid, C., Magnasco, M., Vosshall, L. & Keller, A. Humans can Discriminate more than one Trillion Olfactory Stimuli HHS Public Access. Science (1979) 343, 1370–1372 (2014).
Chae, H. et al. Mosaic representations of odors in the input and output layers of the mouse olfactory bulb. Nat Neurosci 22, 1306 (2019).
Arn, H. & Acree, T. Flavornet: a database of aroma compounds based on odor potency in natural products (1998).
FooDB. www.foodb.ca.
Manach, C. FoodComEx a new chemical library for rare food-derived compounds. https://www.researchgate.net/publication/289522373_FoodComEx_a_new_chemical_library_for_rare_food-derived_compounds (2016).
Mobley, D. L. & Guthrie, J. P. FreeSolv: a database of experimental and calculated hydration free energies, with input files. J Comput Aided Mol Des 28, 711–720 (2014).
The Good Scents Company Information System. https://www.thegoodscentscompany.com/index.html.
CFR - Code of Federal Regulations Title 21. https://www.accessdata.fda.gov/scripts/cdrh/cfdocs/cfcfr/CFRSearch.cfm?CFRPart=184&showFR=1.
Haddad, R., Carmel, L., Sobel, N. & Harel, D. Predicting the receptive range of olfactory receptors. PLoS Comput Biol 4, 18 (2008).
IFRA Fragrance Ingredient Glossary. https://ifrafragrance.org/priorities/ingredients/glossary.
Iurilli, G. & Datta, S. R. Population Coding in an Innately Relevant Olfactory Area. Neuron 93, 1180–1197.e7 (2017).
Johnson, B. A., Xu, Z., Ali, S. S. & Leon, M. Spatial representations of odorants in olfactory bulbs of rats and mice: Similarities and differences in chemotopic organization. Journal of Comparative Neurology 514, 658–673 (2009).
Jones, E. M. et al. A Scalable, Multiplexed Assay for Decoding GPCR-Ligand Interactions with RNA Sequencing. Cell Syst 8 (2019).
Keller, A., Hempstead, M., Gomez, I. A., Gilbert, A. N. & Vosshall, L. B. An olfactory demography of a diverse metropolitan population. https://doi.org/10.1186/1471-2202-13-122 (2012).
Keller, A. & Vosshall, L. B. Olfactory perception of chemically diverse molecules. BMC Neurosci 17, 55 (2016).
ChemInfo.org. Knapsack. https://www.cheminfo.org/Chemistry/Database/Knapsack/index.html.
Sanchez-Lengeling, B. et al. Leffingwell Odor Dataset, https://doi.org/10.5281/zenodo.4085098 (2020).
Ma, L. et al. Distributed representation of chemical features and tunotopic organization of glomeruli in the mouse olfactory bulb. Proc Natl Acad Sci USA 109, 5481–5486 (2012).
Ma, Y., Tang, K., Xu, Y., Thomas-Danguin, T. & Thomas, T. A dataset on odor intensity and odor pleasantness of 222 binary mixtures of 72 key food odorants rated by a sensory panel of 30 trained assessors. Data Brief 36, 107143 (2021).
Mainland, J. D., Li, Y. R., Zhou, T., Liu, W. L. L. & Matsunami, H. Human olfactory receptor responses to odorants. Sci Data 2 (2015).
Manoel, D. et al. Deconstructing the mouse olfactory percept through an ethological atlas. Current Biology 31 (2021).
Mayhew, E. J. et al. Transport features predict if a molecule is odorous. Proc Natl Acad Sci USA 119, e2116576119 (2022).
Nagappan, S. & Franks, K. M. Parallel processing by distinct classes of principal neurons in the olfactory cortex. Elife 10 (2021).
Nakayama, H., Gerkin, R. C. & Rinberg, D. A behavioral paradigm for measuring perceptual distances in mice. Cell reports methods 2 (2022).
NHANES 2013-2014: Taste & Smell Data Documentation, Codebook, and Frequencies. https://wwwn.cdc.gov/Nchs/Nhanes/2013-2014/CSX_H.htm.
Ravia, A. et al. A measure of smell enables the creation of olfactory metamers. Nature 588 (2020).
Scott, J. W., Sherrill, L., Jiang, J. & Zhao, K. Tuning to Odor Solubility and Sorption Pattern in Olfactory Epithelial Responses. https://doi.org/10.1523/JNEUROSCI.3736-13.2014 (2014).
Sharma, A., Kumar, R., Ranjta, S. & Varadwaj, P. K. SMILES to Smell: Decoding the Structure-Odor Relationship of Chemical Compounds Using the Deep Neural Network Approach. J Chem Inf Model 61, 676–688 (2021).
Sharma, A., Kumar Saha, B., Kumar, R. & Kumar Varadwaj, P. OlfactionBase: a repository to explore odors, odorants, olfactory receptors and odorant-receptor interactions. Nucleic Acids Ress 50 (2022).
SAFC® Sigma Flavors & Fragrances Catalog, (2014).
Slone, J. D. et al. Functional characterization of odorant receptors in the ponerine ant, Harpegnathos saltator. Proc Natl Acad Sci USA 114, 8586–8591 (2017).
Snitz, K. et al. Predicting Odor Perceptual Similarity from Odor Structure. PLoS Comput Biol 9, e1003184 (2013).
Snitz, K. et al. SmellSpace: An Odor-Based Social Network as a Platform for Collecting Olfactory Perceptual Data. Chem Senses 44, 267–278 (2019).
Soh, Z. et al. A Comparison Between the Human Sense of Smell and Neural Activity in the Olfactory Bulb of Rats. Chem. Senses 39, 91–105 (2014).
Dunkel, M. et al. SuperScent—a database of flavors and scents. Nucleic Acids Res 37, D291–D294 (2009).
The Toxin and Toxin Target Database (T3DB). http://www.t3db.ca/.
Wakayama, H., Sakasai, M., Yoshikawa, K. & Inoue, M. Method for Predicting Odor Intensity of Perfumery Raw Materials Using Dose-Response Curve Database. Ind Eng Chem Res 58, 15036–15044 (2019).
Weiss, T. et al. Perceptual convergence of multi-component mixtures in olfaction implies an olfactory white. Proc Natl Acad Sci USA 109, 19959–19964 (2012).
Yu, Y. et al. Responsiveness of G protein-coupled odorant receptors is partially attributed to the activation mechanism. Proc Natl Acad Sci USA 112, 14966–14971 (2015).
Acknowledgements
We thank all of those who contributed datasets to the project and NIH for support under R01DC018455, U19NS112953, and R01DC017757. Further support for this work was provided by NSF Grant 1553270 (to JBC), NIH Grant F32DC020380 (to RP), and NIH Grant T32DC000014 (to EAH and RP).
Author information
Authors and Affiliations
Contributions
R.C.G. conceived of the overall project, wrote the Pyrfume library, curated some Pyrfume datasets, wrote the manuscript, and wrote the grant application. J.D.M. helped with implementation of the project and co-wrote the grant application. J.B.C. and T.J.G. curated and standardized Pyrfume datasets, and wrote the manuscript. R.P., Z.L., L.A.C., F.P., D.S.W., T.B.: Contributed to the Pyrfume codebase. E.A.H.: Wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hamel, E.A., Castro, J.B., Gould, T.J. et al. Pyrfume: A window to the world’s olfactory data. Sci Data 11, 1220 (2024). https://doi.org/10.1038/s41597-024-04051-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-04051-z
This article is cited by
-
A comparative study of machine learning models on molecular fingerprints for odor decoding
Communications Chemistry (2025)