Abstract
Ultrasonography (US) of thyroid nodules is often time consuming and may be inconsistent between observers, with a low positivity rate for malignancy in biopsies. Even after determining the ultrasound Thyroid Imaging Reporting and Data System (TIRADS) stage, Fine needle aspiration biopsy (FNAB) is still required to obtain a definitive diagnosis. Although various deep learning methods were developed in medical field, they tend to be trained using TI-RADS reports as image labels. Here, we present a large US dataset with pathological diagnosis annotation for each case, designed for developing deep learning algorithms to directly infer histological status from thyroid ultrasound images. The dataset was collected from two retrospective cohorts, which consists of 8508 US images from 842 cases. Additionally, we explained three deep learning models used as validation examples using this dataset.
Similar content being viewed by others
Background & Summary
The widespread use of imaging methods has contributed to a 2.4-fold increase in the incidence of thyroid cancer over the last 30 years, the fastest increase of any cancer type1,2, with a correspondingly unchanged or declining mortality rate for thyroid cancer3. This has given rise to the need for accurate diagnosis of thyroid cancer. There are two commonly used methods for diagnosing benign and malignant thyroid nodules: non invasive ultrasonography (US)4 and invasive fine-needle aspiration biopsy (FNAB)5. Currently, ultrasound is the first clinical choice of thyroid nodules screening, because of its non-radioactivity, easy-to-operate, and rapid diagnostic work-up6,7,8,9,10. In actual studies, there are overlapping ultrasound image features of benign and malignant thyroid nodules with blurred nodule appearance and irregular shape11,12, making manual feature extraction and annotation difficult for experience-based clinicians. In addition, the clinical experience of the operator, and different diagnostic criteria can interfere with the physician’s assessment of thyroid nodules13,14. The sensitivity of using US to diagnose thyroid cancer ranges from 27% to 63% only15,16. In clinical practice, ultrasound Thyroid Imaging Reporting and Data System (TIRADS) stage >3 is recommended to perform FNAB or surgical resection17. The use of FNAB can cause trauma to patients and incur additional costs. Moreover, FNAB is not absolutely effective and suffers from ultrasound localization of nodules, failing to provide a definitive diagnosis in at least 20% of patients or requiring repeated FNABs that still do not yield definitive results18,19,20,21. Therefore, to address the exponentially increasing patient demand and reduce the burden on healthcare services, computer-aided diagnosis was introduced to improve diagnostic performance. This approach aims to automate image analysis, providing a robust and reliable diagnosis22.
Deep learning (DL) is a subset of machine learning (ML) and artificial intelligence (AI) and can automatically extract features from images with complex hierarchical structures23. The introduction of DL such as convolutional neural networks in thyroid imaging has achieved better diagnostic results than experienced radiologists24,25,26. However, in practice, thyroid ultrasonography acquires the region of interest by reading dynamic images, the number of which is usually large and uneven. Different nodules of the same patient may have different labels, and the annotation of all images is a tedious and time-consuming task. Furthermore, the previous DL-based methods tend to be trained using TI-RADS reports as image labels. After that, histological examinations must be conducted to obtain the final histological diagnosis. Given the difficulty and inconsistency of dynamic image annotation in US, FNAB remains the gold standard for nodule diagnosis even after ultrasound assessment. Therefore, we have built a US of thyroid nodules dataset with direct histological diagnostic labels, aiming to explore the relationship between ultrasound images and videos and histological diagnosis.
Methods
Subject characteristics
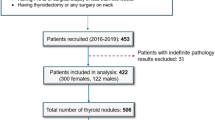
The thyroid nodules ultrasound images from 842 cases were collected at the Second Affiliated Hospital of Jiaxing University in China in 2019–2022 (see Fig. 1). The datasets were prepared according to the following inclusion criteria: (1) hemi‐ or total thyroidectomy, (2) maximum nodule diameter 2.5 cm, (3) examination by conventional US and real‐time elastography (RTE) within 1 month before surgery, and (4) no previous thyroid surgery or percutaneous thermotherapy. This study received approval from the institutional review boards of the Second Affiliated Hospital of Jiaxing University (No. 2022ZFYJ295-01). The requirement of obtaining written informed consent from patients was forgone because retrospective data collection has not impacted the standard diagnostic procedures and all data has been anonymized before being entered into the database. The ethics committee has issued a waiver of consent and approved the open publication of the dataset.
Thyroid image instances.
Image acquisition
Ultrasound acquisitions were performed using a portable machine equipped with a 3.5 MHz probe. The Esaote My Lab was used in the Second Affiliated Hospital of Jiaxing University. During each acquisition, the operator positioned the probe. These original images were cropped to remove sensitive information about patients. Histological labels were obtained according to the histological diagnosis report of the corresponding tissue slides.
Data Records
All the ultrasound images in ‘JPG’ format for each case with pathological diagnosis annotation (benign or malignant) is available at the public figshare repository27. The images in JPG format and their associated meta data in .csv format are stored in .zip files within the repository. The datasets file structure are shown in Fig. 2, the demographic and histological label information can be matched using the corresponding case name in the .csv files.
Overview of the datasets structure.
Technical Validation
To validate the dataset proposed in this study, we introduced a novel dual attention-guided deep learning framework, ThyUS2Path. We assessed its performance against two state-of-the-art multiple instance learning (MIL) based methods, Meanpool and Maxpool. The MIL is a type of weakly supervised learning where training instances are arranged in sets, and a label is provided for the entire set, opposedly to the instance themselves. The dataset was organized into two batches: (1) Batch 1, consisting of 6,005 thyroid images from 601 patients, and (2) Batch 2, consisting of 2,503 thyroid images from 241 patients. The subject demographics of these datasets are shown in Table 1. Model training was conducted using a 5-fold cross-validation strategy. For Batch 1, 90% of the patients were allocated to the training-validation set, while the remaining 10% were reserved for testing. The training-validation set was further split using 5-fold cross-validation, with each fold comprising 4,380 images for training and 1,077 images for validation, ensuring no overlap between validation sets. Finally, 548 images were used for internal testing, while all 2,503 images from Batch 2 were used for external independent validation.
Our model is composed of three interconnected modules (Fig. 3):
-
1)
Backbone Network: We adapted the state-of-the-art ResNet-34 as our backbone network to extract relevant features from each thyroid image. Specifically, we removed the final classification layer, which is the fully connected layer with 1,000 neurons. The backbone network processes thyroid images from each patient to automatically extract tumor-related nodule patterns.
-
2)
Dual Attention Feature Aggregation Module: The features extracted by the backbone network are passed through a dual attention module (Fig. 3). This module comprises two sub-modules: spatial attention and instance attention. The input to the dual attention module is the output from the final convolutional layer of the backbone network. First, the spatial attention sub-module filters the input features of multiple images from each patient along the spatial dimension to capture subtle relationships between adjacent regions within each image. Importance scores are generated for each region to quantify its contribution to the final prediction. These spatially filtered features are then processed by the instance attention sub-module, which assigns attention scores to weight the different images from each case. This process aggregates the features into a patient-level thyroid nodule representation.
-
3)
Fully Connected Layer Classifier: The final module is a fully connected layer with 2 neurons, which converts the patient-level features generated by the dual attention module into a histological diagnosis prediction for the patient.
Overview of the proposed deep learning framework in this study. (a,b) The training strategy, where multiple thyroid images from each patient are input into ThyUS2Path to predict the histological label. (c) The structure of the framework, which is composed of three main components: the backbone network, the dual-attention module, and the fully connected classifier.
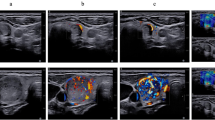
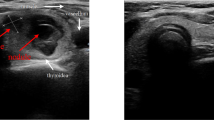
We utilized 6,005 thyroid images from 601 patients to develop the deep learning framework. The dataset characteristics are summarized in Fig. 4. As depicted in Fig. 4a, the training dataset included 601 patients, comprising 218 benign cases and 383 malignant cases, each with corresponding histological label. Since each patient may have multiple thyroid images, we provided a statistical overview of all patients in Fig. 4b. Additionally, we displayed representative thyroid nodule images from both malignant and benign patients for comparison in Fig. 1.
Performance comparison between ThyUS2Path and state-of-the-art MIL-based methods on the internal test set.
Due to variations caused by different operators in thyroid ultrasonography, images from different cohorts can differ in appearance. Ensuring the generalizability of computational ultrasonography algorithms to real-world clinical data is essential. To address this, we collected 2,503 thyroid nodule images from a separate cohort as an external test dataset. As shown in Table 2 and Fig. 5, the results on this dataset were promising, with AUROCs (area under the receiver operating characteristic curve) ranging from 0.70 to 0.80 and AUPRCs (area under the precision-recall curve) from 0.78 to 0.83, demonstrating that ThyUS2Path generalized well to heterogeneous real-world data. Furthermore, the other two MIL-based methods also yielded promising results, further validating the validity and reliability of our dataset. These findings may be used to evaluated the generalization performance of different deep learning algorithms for thyroid nodule identification.
ROC and PR curves of ThyUS2Path and state-of-the-art MIL-based methods on the external test set.
Limitations
The datasets and model have some limitations. Firstly, because the dataset is from retrospective cohorts, there may be some quality problems such as lower resolution in some of these images. Secondly, we validated the datasets only on three different models. For a more comprehensive evaluation, future studies should include a broader range of models to establish a more robust baseline. Despite these limitations, these datasets still provide valuable insights into the thyroid nodules identification and can serve as a foundation for future research.
Code availability
The code used in this study was written in Python3 and is available at GitHub (https://github.com/TomHardy1997/ThyUS2Path). The code is based on PyTorch (version 1.6).
References
Hayat, M. J., Howlader, N., Reichman, M. E. & Edwards, B. K. Cancer statistics, trends, and multiple primary cancer analyses from the Surveillance, Epidemiology, and End Results (SEER) Program. Oncologist. 12(1), 20–37 (2007).
Zhu, C. et al. A birth cohort analysis of the incidence of papillary thyroid cancer in the United States, 1973–2004. Thyroid. 19(10), 1061–6 (2009).
Ahn, H. S. et al. Thyroid Cancer Screening in South Korea Increases Detection of Papillary Cancers with No Impact on Other Subtypes or Thyroid Cancer Mortality. Thyroid. 26(11), 1535–40 (2016).
Brito, J. P. et al. The accuracy of thyroid nodule ultrasound to predict thyroid cancer: systematic review and meta-analysis, The Journal of Clinical Endocrinology & Metabolism 99(4), pp. 1253–1263 (2014).
Cibas, E. S. & Ali, S. Z. The 2017 bethesda system for reporting thyroid cytopathology, Thyroid, vol. 27, no. 11, pp. 1341–1346, (2017).
Chikui, T. et al. Quantitative analyses of sonographic images of the parotid gland in patients with Sjögren’s syndrome. Ultrasound Med Biol. 32(5), 617–22 (2006).
Haugen, B. R. et al. 2015 American Thyroid Association Management Guidelines for Adult Patients with Thyroid Nodules and Differentiated Thyroid Cancer: The American Thyroid Association Guidelines Task Force on Thyroid Nodules and Differentiated Thyroid Cancer. Thyroid. 26(1), 1–133 (2016).
Mahesh, M. The Essential Physics of Medical Imaging, Third Edition. Med Phys. 40(7) (2013).
Jemal, A., Siegel, R., Xu, J. & Ward, E. Cancer statistics, 2010. CA Cancer J Clin. 60(5), 277–300 (2010).
Cooper, D. et al. Revised American Thyroid Association management guidelines for patients with thyroid nodules and differentiated thyroid cancer. Thyroid 19(11), 1167–1214 (2009).
Iannuccilli, J. D., Cronan, J. J. & Monchik, J. M. Risk for malignancy of thyroid nodules as assessed by sonographic criteria: the need for biopsy. J Ultrasound Med. 23(11), 1455–64 (2004).
Wienke, J. R., Chong, W. K., Fielding, J. R., Zou, K. H. & Mittelstaedt, C. A. Sonographic features of benign thyroid nodules: interobserver reliability and overlap with malignancy. J Ultrasound Med. 22(10), 1027–31 (2003).
Tessler, F. N. et al. ACR Thyroid Imaging, Reporting and Data System (TI-RADS): White Paper of the ACR TI-RADS Committee. J Am Coll Radiol. 14(5), 587–95 (2017).
Russ, G. et al. European Thyroid Association Guidelines for Ultrasound Malignancy Risk Stratification of Thyroid Nodules in Adults: The EU-TIRADS. Eur Thyroid J. 6(5), 225–37 (2017).
Frates, M. C. et al. Management of thyroid nodules detected at US: Society of Radiologists in Ultrasound consensus conference statement. Radiology. 237(3), 794–800 (2005).
Remonti, L. R., Kramer, C. K., Leitão, C. B., Pinto, L. C. & Gross, J. L. Thyroid ultrasound features and risk of carcinoma: a systematic review and meta-analysis of observational studies. Thyroid. 25(5), 538–50 (2015).
Hang, J. F., Hsu, C. Y. & Lai, C. R. Thyroid Fine-Needle Aspiration in Taiwan: The History and Current Practice. J Pathol Transl Med. 51(6), 560–4 (2017).
Misiakos, E. P. et al. Cytopathologic diagnosis of fine needle aspiration biopsies of thyroid nodules. World J Clin Cases. 4(2), 38–48 (2016).
Sebo, T. J. What are the keys to successful thyroid FNA interpretation? Clin Endocrinol (Oxf). 77(1), 13–7 (2012).
Theoharis, C. G., Schofield, K. M., Hammers, L., Udelsman, R. & Chhieng, D. C. The Bethesda thyroid fine-needle aspiration classification system: year 1 at an academic institution. Thyroid. 19(11), 1215–23 (2009).
VanderLaan, P. A., Marqusee, E. & Krane, J. F. Clinical outcome for atypia of undetermined significance in thyroid fine-needle aspirations: should repeated fna be the preferred initial approach? Am J Clin Pathol. 135(5), 770–5 (2011).
Doi, K. Computer-aided diagnosis in medical imaging: historical review, current status and future potential. Comput Med Imaging Graph. 31(4-5), 198–211 (2007).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature. 521(7553), 436–44 (2015).
Li, X. et al. Diagnosis of thyroid cancer using deep convolutional neural network models applied to sonographic images: a retrospective, multicohort, diagnostic study. Lancet Oncol. 20(2), 193–201 (2019).
Wei, X. et al. Ensemble Deep Learning Model for Multicenter Classification of Thyroid Nodules on Ultrasound Images. Med Sci Monit. 26, e926096 (2020).
Shen, W., Zhou, M., Yang, F., Yang, C. & Tian, J. Multi-scale Convolutional Neural Networks for Lung Nodule Classification. Inf Process Med Imaging. 24, 588–99 (2015).
Hou, X. W. et al. An ultrasonography of thyroid nodules dataset with pathological diagnosis annotation for deep learning. https://doi.org/10.6084/m9.figshare.27021604.v1 (2024).
Acknowledgements
The study was supported by National Natural Science Foundation of China (NSFC) (82304250, 82273734). Shanghai Three-Year Action Plan for Strengthening Public Health System (GWVI-11.1-26), Cardiovascular Disease - Cardiovascular and Cerebrovascular Diseases, Key Discipline Project. Shanghai Municipal Science and Technology Commission Professional and Technical Service Platform Project (22DZ2292400), “Shanghai Municipal Cardiovascular and Cerebrovascular Disease Biological Sample and Database Professional and Technical Service Platform”. 2023ZZ02021, Shanghai Research Center of Cardiovascular and Cerebrovascular Diseases, Shanghai Municipal Health Commission, Shanghai, China. 2022JC013, “Accurate Early Warning and Intervention of Cardiovascular and Cerebrovascular Diseases Based on the Whole-Life Cohort”, Emerging Cross-cutting Research Program, Shanghai Municipal Health Commission. Health Research Program of Pudong New Area Health Commission (PW2023E-02), “Effects of Sleep Patterns on Cardiovascular and Cerebrovascular Diseases in a Community Cohort in Pudong”.
Author information
Authors and Affiliations
Contributions
M.H., S.S., L.C., X.H., K.L. and L.W. conceived of the project, M.H., X.Z. and J.J. designed the model and developed the algorithms, S.S., X.H. provided and annotated the images, W.Z., H.J., M.L., X.W. and W.Z. participated in workstation environment deployment. M.H., J.J., L.W and X.Z. contributed to the preprocess of the data. M.H. and L.W. wrote the manuscript. All the authors revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hou, X., Hua, M., Zhang, W. et al. An ultrasonography of thyroid nodules dataset with pathological diagnosis annotation for deep learning. Sci Data 11, 1272 (2024). https://doi.org/10.1038/s41597-024-04156-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-04156-5
This article is cited by
-
Training the diagnostic artificial intelligence in thyroid sonography: how well is deep learning truly learning?
Updates in Surgery (2026)
-
TN5000: An Ultrasound Image Dataset for Thyroid Nodule Detection and Classification
Scientific Data (2025)
-
An annotated heterogeneous ultrasound database
Scientific Data (2025)