Abstract
Observational health research often relies on accurate and complete race and ethnicity (RE) patient information, such as characterizing cohorts, assessing quality/performance metrics of hospitals and health systems, and identifying health disparities. While the electronic health record contains structured data such as accessible patient-level RE data, it is often missing, inaccurate, or lacking granular details. Natural language processing models can be trained to identify RE in clinical text which can supplement missing RE data in clinical data repositories. Here we describe the Contextualized Race and Ethnicity Annotations for Clinical Text (C-REACT) Dataset, which comprises 12,000 patients and 17,281 sentences from their clinical notes in the MIMIC-III dataset. Using these sentences, two sets of reference standard annotations for RE data are made available with annotation guidelines. The first set of annotations comprise highly granular information related to RE, such as preferred language and country of origin, while the second set contains RE labels annotated by physicians. This dataset can support health systems’ ability to use RE data to serve health equity goals.
Similar content being viewed by others
Background & Summary
Many areas of observational health research and clinical informatics research rely on accurate and complete race and ethnicity (RE) patient information, particularly for estimating disease risk1,2,3, assessing quality and performance metrics4, and identifying health disparities5,6,7,8. The electronic health record (EHR) provides a rich source of patient health data, but while RE is often stored in easily accessible structured EHR fields, this format often suffers from missing, inadequate, or inaccurate information9,10,11,12,13,14,15,16. For example, Polubriaginof et al. found missing RE data affected 25% of patients with data in large observational health databases and 57% of patients at a large academic medical center in New York City9. Finally, while there has been an acknowledged need for more granular information such as preferred language, this data is often not recorded in EHR systems17. Overall, missing RE data decreases a patient’s visibility within research and healthcare systems and can affect the allocation of resources in hospitals and health systems to best serve health equity goals. When done carefully and thoughtfully, the ability to supplement missing or inaccurate RE data is an important step toward increasing the diversity of patients represented in observational health research and supporting health equity, by filling in a common unobserved confounder. Further, the Affordable Care Act and other federal laws now include non-discrimination clauses18, compliance with which can only be assessed with adequate RE data at the patient- and provider-levels. Similarly, algorithmic bias, fairness, and recourse can trickle down into clinical guidelines starting from reported differences between races in epidemiological statistics such as prevalence19. Finally, given the rise of large language models in clinical care, operations, and revenue cycle management20, the training data available can lead to biased output that can impact clinical care and a hospital’s revenue. It is well known in the algorithmic fairness research field that there is no way to create risk scores that are fair with respect to the intersection of multiple legally-protected classes (such as a readmission risk score prediction algorithm that is fair with regard to subgroups defined using both race and sex21). Given that labelled training data is difficult to come by and federal laws such as HIPAA restrict the transmission of large language model parameters, reference standard RE annotations with high inter-rater reliability are a significant opportunity to ensure informed consent for clinicians, patients, hospitals, and health systems that use tools such as large language models to inform triage, decision-making, resource allocation, and revenue.
Inadequate RE categories can also mask important subgroup differences, in part due to a lack of sufficient granularity13,22,23,24,25,26,27,28. While still important for federal data reporting, concerns of inadequate data have generally revolved around the Office of Management and Budget’s (OMB) five race categories (American Indian or Alaska Native, Black or African American, Asian, Native Hawaiian or Other Pacific Islander, white) and two ethnic categories (Hispanic or Latino and Not Hispanic or Latino)29. The Institute of Medicine’s landmark report, Unequal Treatment: Confronting Racial and Ethnic Disparities in Health Care, expressed concern over the lack of more granular RE categories and how this hinders health disparities research13. Furthermore, research has called for adapting RE categories to a multiracial and multi-ethnic U.S. population. Without access to more granular information, current OMB category standards obscure subpopulations that can have distinct healthcare needs30,31,32,33. More granular information can highlight country or geographic region of origin and level of language proficiency and help distinguish differences within these broader categories. For example, major differences have been observed among people of Asian descent in the U.S. with respect to access to mental healthcare31 and cancer incidence32 based on differences in English language proficiency and country of origin, respectively. More granular race information can also be used to uncover disparities in clinical risk scores that would otherwise be concealed with coarse race groups34.
Clinical text often provides a rich, unstructured source of granular information related to RE, such as immigration status35, country of origin36, and preferred language17. Natural language processing (NLP) models can be trained to identify RE in clinical text to supplement and/or complement structured RE data that is missing, inaccurate, or lacking in granularity. For example, Sholle et al. developed a rule-based approach to extract RE categories from clinical text and achieved excellent performance for identifying Black and Hispanic patients37. Within the setting of a hospital in an affluent neighborhood of New York City, Sholle et al. found that clinical notes could increase positive documentation of RE data by upwards of 20% for Black and/or Hispanic patients with previously missing RE data37. One major challenge to training NLP models to identify RE data from clinical text, however, is the need for reference standard annotations, which can be costly and time-consuming to create. Publicly available reference standard annotations can support these tasks and future research on patterns of clinical documentation of RE in clinical text.
We present the Contextualized Race and Ethnicity Annotations for Clinical Text (C-REACT) dataset, two sets of publicly available reference standard annotations on 17,281 sentences from 12,000 patients from the MIMIC-III dataset, a corpus extracted from critical care units at Beth Israel Deaconess Medical Center between 2001 and 201238. The first set of annotations provides granular detail on RE at the span level within sentences. The second set of annotations are physician-assigned RE labels at the sentence level. Both annotation sets and their guidelines are made available to the research community to enable widespread use of more granular RE-related information in clinical notes and demonstrate how NLP can be leveraged to infer RE using clinical text.
While other datasets for RE exist, such as such as The Home Mortgage Disclosure Act data (https://www.consumerfinance.gov/data-research/hmda/historic-data/), race imputation using name information39, or aggregate clinical trials data (ClinicalTrials.gov), clinical note datasets for RE are difficult to make public given the privacy risks involved for individual patients. Research through the National NLP Clinical Challenges does offer access to clinical text annotations for variety of tasks but does not include RE annotations at the level provided by C-REACT. Given that MIMIC-III is the only publicly accessible clinical dataset (with the required credentials) that combines patient data including clinical notes, labs, diagnosis codes, demographics, medications, and procedures on 59,652 patients, the addition of C-REACT to MIIMIC-III can greatly enhance NLP research into RE extraction from clinical notes.
Methods
In this section, we describe our approach to annotate clinical text in two ways (1) at the span level for RE-related information (i.e., RE indicators) and (2) at the sentence level for RE assignment. We then describe our analysis of the presence of indicators within sentences in relation to RE assignments.
Data and pre-processing
We extracted all sentences from 59,652 discharge summary clinical notes for 41,127 patients from the MIMIC-III dataset38 (version 1.4). We used NLTK40 to extract sentences and heuristics to handle clinical lists such as medication and condition lists. Sentences likely to contain RE information were identified using keywords related to patient demographics (e.g., “male”, “female”, “patient”) and section headings (e.g., “Past Medical History”, “PMH”, “Social History”, “SHX”). Case was ignored for these keywords. From the entire set of discharge summary sentences, those with demographic keywords and/or section headers were extracted as the candidate corpus in Table 1 (n=794,841). The comprehensive list of section header keywords used is “sshx, “social history”, “social hx”, “pmh”, “past medical history”, “pmhx”, “hpi”, and “history of present illness”. Sentences with RE keywords (e.g., “Black”, “AA”, “Native American”, “Hispanic”, “Spanish”) were prioritized for explicit indicator span annotation (Table 2). Section heading, patient demographic, and RE keyword identification were not case sensitive. We conducted two separate annotation processes (one for RE indicators, one for labels) using 17,281 sentences sampled from the 794,841 sentences. This corpus comprised 13,507 notes for 12,000 patients. We refer to the corpus of 17,281 sentences as the central corpus since it is used in both RE indicator and label annotation phases. Table 1 provides more details on the central corpus.
Sentence sampling process
Of the 794,841 sentences extracted from MIMIC-III, 17,281 were sampled to create the central corpus. All sentences extracted contained section headers and/or demographic keywords. Sentences can be split into three main categories: (1) those with RE keywords (RE matches), (2) those with section headers (headers), and (3) those with only demographic keywords (dems). Sentences were sampled randomly within each category iteratively until sentences with RE keywords or headers were exhausted. More specifically, RE matches were randomly sampled at a rate of 50%, headers at 25%, and dems at 25%. Given that dems is the only mutually exclusive category, headers and matches have significant overlap and thus the percentages in Table 1 do not exactly reflect the sampling percentages. As the RE matches and headers categories overlap, we differentiate between sentences with RE keywords AND NO section headers, RE keywords AND section headers, and section headers AND NO RE keywords (Table 1). Sentences containing RE keywords and/or section headers were prioritized for sampling as we hypothesized that these were likely to contain RE discussions. While RE discussions were hypothesized to be rare in sentences with only demographic keywords, we randomly sampled from this subset to reduce bias in our dataset. More specifically, sentences likely to have positive RE labels are important for training, but a diversity of sentences without positive labels are also important as these are the most common sentences researchers will encounter in real-world settings.
Explicit span-level indicator annotation process
The explicit indicator annotation process was performed by non-physicians (n=4), and focused on identifying spans of text that explicitly convey RE-relevant information. All 17,281 sentences in the central corpus were annotated for RE indicators. We chose four categories of indicators to capture explicit spans of text potentially describing RE: (1) spans of text that discuss country/nation of origin or geographic ancestry (country); (2) spans of text that discuss primary, preferred, or spoken language (language); (3) spans of text that discuss direct race mentions (race); and (4) spans of text that discuss direct ethnicity mentions (ethnicity). It is important to note that the race and ethnicity indicators follow U.S.-centric definitions of race and ethnicity29. Examples of these four indicators can be found in Table 3. The annotation guidelines for indicators are available in the PhysioNet dataset under the file name “File_1_Annotation_Guidelines_for_Race_and_Ethnicity_Indicators.docx”. Annotations were conducted using the software Prodigy(https://prodi.gy/)41.
Macro F1 was used to measure inter-annotator agreement instead of the more traditional Cohen’s kappa, given that the annotation task is at the span-level42,43. The F1-score is defined as the harmonic mean between precision and recall. Given that the number of negative cases is unknown, the F1-score is more appropriate than Cohen’s kappa when annotating spans of text42. Additionally, this measure was computed for exact span matches rather than across tokens. Aggregating the F1-score across indicator classes was performed using macro F1 as it is more sensitive to imbalanced data than micro F1. When calculating macro F1 scores for a pair of annotators, one annotator was treated as the reference standard and the other annotator was compared to their annotations. For a pair of annotators, macro F1 scores are the same regardless of which annotator is chosen as the reference standard annotator. All sentences were double-coded and annotators iteratively updated guidelines until sufficient agreement was present (>0.85 macro F1). While iterating, all annotators converged as a group to discuss any sentences with different annotations and settle on the correct annotation to be used. If necessary, the annotation guidelines would be updated to prevent the potential for similar disagreements in the future. When a pair of annotators reached sufficient agreement, they were allowed to annotate independently and continue to provide input on the annotation guidelines. The annotation guidelines are publicly available to support reproducibility.
Assigning race and ethnicity labels to sentences
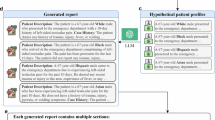
The RE labeling process was conducted by physicians (n=10) with a medical degree and at least one year of post-graduate residency experience. All sentences from the central corpus were annotated by at least one physician for RE labels. Physicians were provided with two subsets of sentences to annotate: one subset (n=5,834) contained sentences that all physicians annotated independently (shared annotation subset), and the other subset (n=11,447) contained the remainder of the sentences split evenly among physicians to annotate (single annotation subset). For a graphical depiction of this information please see Fig. 1. Each physician annotated approximately 1,144 sentences in their single annotation subset. The shared annotation subset contained 4,834 sentences with at least one positive RE indicator span annotation and 1,000 sentences randomly sampled from sentences without any RE indicator. It was later determined that one of these 1,000 sentences contained a positive indicator span that was previously missed; the final data reflects this change. None of the sentences in the single annotation subset contained any positive indicators based on physician review during the indicator annotation step. It was later determined that five sentences in the single annotation subset contained positive indicator spans that were previously missed; the final data reflects this change. The annotation guidelines for RE categories are available in the PhysioNet dataset under the file name “File_2_Annotation_Guidelines_for_Race_and_Ethnicity_Assignments.docx”. We divided sentences such that the shared annotation subset contained all the sentences with known indicators because we hypothesized these sentences to have the vast majority of the positive RE labels and thus require multiple annotators to confirm a positive label. The single annotation subset contained no known indicators and we hypothesized that it would have very few positive RE labels. Our hypotheses were confirmed (see the Technical Validation section for more details).
Subsets of sentences to be annotated for race and ethnicity labels from the central corpus. More green means that more race and ethnicity (RE) indicator spans are included in the subset.
In the interest of exploring the relationship between RE labels and explicit indicator annotations, we only provided physicians with sentences from the central corpus and did not include explicit indicator annotations performed by non-physicians. We also provided physicians with sentence-level RE labeling guidelines, which emphasized that the entire content of the sentence could be used to infer RE labels, and physicians could rely on any previously acquired knowledge related to the sentence (e.g., clinical training or life experience) to infer RE if they so desire. Physicians were not provided with other information about the patient (e.g., patient identifiers). The RE label set consists of positive and negative labels. The positive labels included the U.S. census categories and an additional label, “Not Covered”, to signal the presence of a RE category that falls outside of the census categories (Table 4). A negative label, “No Information Indicated”, was included to explicitly convey that a sentence did not contain sufficient, if any, RE information to make a positive assignment. Multiple labels per sentence were permitted, and each sentence had to be assigned at least one race and at least one ethnicity label (either positive or negative). More specifically, annotators could assign one or more labels, excluding the negative label. The negative label “No Information Indicated” could be used when annotators felt there was not enough information to make a positive assignment. This follows the structured data in MIMIC-III, which also allows for more than one RE category to be documented for a patient. The negative category “No Information Indicated” allows annotators to abstain from assigning a positive label for any sentence. In addition to the 10 individual annotations per sentence in the shared annotation subset, we also kept track of global annotation labels composed of the majority vote (n >=5) among the 10 physicians. We assigned a RE label to a sentence if five or more physicians agreed on the assignment through their annotations. Multiple RE categories were allowed for each sentence. We measured physician agreement using Cohen’s Kappa for each category in the RE labeling task. Agreement for each physician’s annotations was measured against a leave one out (LOO) majority vote corpus, created using the remaining nine annotators.
During our analysis, we also consolidated our sentence-level RE assignments up to the patient level to compare structured and unstructured sources of RE information. All sentence-level assignments were combined for a patient using a union operation over all sentence-level RE labels. We assigned a patient an RE label if any of their sentences were assigned that label (through the majority vote for the shared annotation sentences and a single vote for the single annotation sentences). Similarly, patient-level structured data in MIMIC-III were combined for each patient using the union operation. MIMIC-III only provides “Ethnicity” information on patients, which captures both race and ethnicity categories, and was mapped to the RE categories used in this work. MIMIC-III Ethnicity category mappings follow the definitions provided in Table 4. A complete mapping can be found in the MIMIC-III mapping table in the PhysioNet dataset under the file name “File_3_MIMIC_Ethnicity_Category_Mapping.docx”.
We acknowledge the nuance, diversity, and history that the terms in Table 4, and their respective abbreviations, are unable to convey. In particular, we want to highlight the terms Hispanic, Latino, Latina, and Latinx. We originally used these terms during the annotation process to be as inclusive as possible with this category, though we recognize that not everyone assigned to this category would equally identify or agree with each term44. Each term has its own unique limitations. For example, the term, Hispanic, ignores ancestries of non-Spanish origin45; use of Latino/Latina implies a gender binary46,47; and the umbrella term, Latinx, is an Anglicization47. In this work we have opted to use the term “Latino” in the interest of brevity to refer to the broader grouping that includes Hispanic, Latino, Latina, and Latinx (and non-Latino for its compliment, which includes non-Hispanic, non-Latino. non-Latina, and non-Latinx) as Latino is the preferred term from the AP Stylebook and can refer to people from (directly or ancestrally) Spanish-speaking lands or cultures as well as Latin America48. We acknowledge that this term may not be the preferred term among all patients49. While this concern applies to all racial and ethnic categories used in this research, it is particularly important for the Latino category as we are actively choosing to use one term over others.
In this research, we used federally recognized RE categories while performing sentence-level RE label annotation. Federally recognized RE categories in the U.S. census have recognized limitations13. However they continue to be widely used in research and hospital quality reporting metrics and thus they continue to have value for research. Additionally, given that each sentence is annotated for RE labels and indicators, the indicators can provide information on nuance that the U.S. census-derived RE labels are missing.
Data Records
Both sets of reference standard annotations are available on PhysioNet50. While PhysioNet51 data are freely accessible, users are required to register, complete a credentialing process, and sign a data use agreement. The C-REACT Dataset project page on PhysioNet provides further details on the dataset and the access application process(https://doi.org/10.13026/t9ka-6k29)50.
There is one main folder containing two jsonl files with span-level RE indicators and sentence-level RE assignments as well as a subdirectory with raw RE assignment files. All records contain a sentence identifier (sentence_id) that can link to sentences in other files. The rest of this section outlines the columns of each file.
The ‘indicators_df.jsonl’ file (17,281 sentences from the entire central corpus) contains information from the original MIMIC-III NOTEEVENTS file, including identifiers for visits (visit_id), patients (patient_id), and notes (note_id). Output from the annotation software Prodigy (https://prodi.gy/) (spans) is also available in the file. Finally, tokenized sentence (text) and tokens (tokens) are presented.
The ‘all_re_assignments_df.jsonl’ file (17,281 sentences from the entire central corpus) contain sentence (text) and sentence identifier (sentence_id) columns. Additionally, the files contain binarized columns for all race and ethnicity categories described in Table 4. Each racial category has ‘RACE’ as a prefix, while ethnicity categories have the prefix ‘ETH’. The final two binary columns (shared_subset, single_subset) indicate the source of the RE assignments as either the shared or single annotation subset respectively. All shared annotation subset assignments are majority vote assignments.
We also provide the raw RE assignment files for each annotator in a second folder. The folder structure for each annotator is the same. Each annotator’s folder contains six xlsx files with ‘all_clinician_sentences’ as a prefix and numbered 0–5 (shared annotation subset). Within this folder there is another folder containing the single annotation subset xlsx file. All xlsx files follow the format outline in Fig. 1 of the File_2_Annotation_Guidelines_for_Race_and_Ethnicity_Assignments.docx file on PhysioNet and contain information on the sentence identifier (ID), sentence text (Sentence), and all RE categories previously described. Annotators marked their annotations by simply adding any character (often ‘x’) to a cell in the xlsx files.
Technical Validation
We present validation results for the RE labels (sentence-level) and indicator annotations (span-level). For RE category annotations, we measured physician-annotator agreement for the RE labels and concordance between structured and unstructured sources for patient-level RE information. For RE indicator annotations, we measured inter-annotator agreement and the proportion of sentences with RE positive labels but no positive RE indicator annotation.
Validating race and ethnicity indicator span-level annotations
Indicators were double coded until all annotator pairs reached sufficient agreement of >0.85 macro F1. Until sufficient agreement was reached, all disagreements were adjudicated through discussion and the annotation guidelines were iteratively updated. Then, we measured how often the majority vote RE labels were assigned to sentences that did not contain at least one indicator annotation. Out of all 12,411 sentences without positive indicator annotations, only six were assigned a positive race and ethnicity label. In other words, six out of 4,811 sentences (0.1%) sentences were assigned a positive RE label but did not contain a positive indicator. All six sentences occurred in the individual annotated subset and contained spans of text that were not considered indicative of race and/or ethnicity in this work, i.e., cuisine, occupation, immigration status, and wars/conflicts. These results provide evidence for high agreement on indicator annotations and confirm that our indicators covered a vast majority of RE discussions in the corpus.
Validating race and ethnicity label annotations
In the sentence-level RE labeling task on the 5,834 shared annotation subset sentences, physicians had moderate to strong agreement (Cohen’s kappa >0.61) when compared to the LOO majority vote assignments. When averaging an annotator’s kappa scores for RE categories with more than 300 assignments, half the annotators had almost perfect agreement (Cohen’s kappa >0.81) agreement and all annotators had substantial agreement (0.61–0.80)52. Perfect agreement is indicated using bold font. Overall, clinical annotators had the lowest agreement for the non-Latino and the “No Information Indicated” categories for both race and ethnicity (Table 5). Another table with identical information and shaded agreement scores is included in Supplementary Table 1 in the supplementary file.
The majority vote sentence-level RE assignments represented 4,575 patients, whose RE labels could be compared to structured RE sources data in MIMIC. This subset of patients will be used to compare RE data in MIMIC-III. For this analysis, race and ethnicity categories were collapsed to match MIMIC-III’s single “Ethnicity” column that contains race and ethnicity categories. Merging RE categories from the MIMIC-III demographics table found in the “File_3_MIMIC_Ethnicity_Category_Mapping.docx” from PhysioNet, a total of 3,931 (85.9%) patients had at least one positive race or ethnicity category in structured data, and 527 (11.5%) patients had race or ethnicity information recovered through unstructured data leading to 4,458 patients (97.4%) with at least one positive race or ethnicity category. Looking specifically at positive race categories, a total of 4,114 (89.9%) patients had race data from structured and/or unstructured sources, with 772 (16.9%) patients being unique to structured and 489 (10.7%) being unique to unstructured (left-hand side of Fig. 2). Of the 489 patients with race information recovered through text, most patients had race information related to being white (n=402), Black or African American (Black/AA) (n=40), and Asian (n=39). Positive ethnicity categories were missing more often than positive race categories, with only 390 (8.5%) patients with positive ethnicity information from any source, 38 (0.8%) patients unique to structured, and 80 (1.7%) unique to unstructured (right-hand side of Fig. 2).
Venn diagram comparing structured and unstructured data sources in their overlap of labels associated with patient race (left) and ethnicity (right) data. Labels concerning race (i.e., Black or African American, Native American or Alaskan Native, Native Hawaiian or Other Pacific Islander, Asian, white, Not Covered) were noted among 4,114 patients, while labels related to ethnicity (i.e., Latino, Not Covered) were attributed to 390 patients.
From the same subset of 4,575 patients with majority vote sentence-level assignments, a significant number of patients had both structured and unstructured data for either ethnicity or race, for which there was high concordance. Of those patients who had structured and unstructured race data (n=2,853), 98.9% had at least partial agreement between the two sources (n=2,821), with the vast majority (n=2,819) of those agreements being perfect. All 272 patients with structured and unstructured data for the Latino category had perfect agreement between the two data sources. This agreement provides evidence RE inferences align with other sources for RE information in MIMIC-III. A small number of patients (n=5) had multiple RE categories documented from the structured and/or unstructured data. Our analysis allows for multiple categories to be documented for a patient much like the structured RE data in MIMIC-III.
We examined the discharge summaries of patients without RE data derived from clinical notes but had structured race or ethnicity data. For the 772 patients missing unstructured race data, most notes examined did not have any mentions of race indicators in the clinical notes or the sentences annotated from those notes. There were some cases of misspellings that were not handled by our regular expressions (e.g., “intertpreter [sic]”), and there were 55 patients who had de-identified country (e.g., “[**country 456**]”) information that was not used to assign patients any RE categories. Finally, there were patients who had race data from a few annotators, but not enough to meet the majority vote requirement and thus received no positive assignment. Similarly, for the 38 patients with only structured ethnicity data, nine had sentences with annotator assignments that failed to meet the majority vote and 12 had de-identified country mentions. For both race and ethnicity assignments, patients had sentences where most annotators assigned a positive race or ethnicity label, but did not agree on which label to use and so no label was assigned in the majority vote. For example, annotators labeled a sentence identifying a patient as Egyptian in the clinical note, as a positive indicator for Black/AA, Asian, white, and Not Covered.
Validating low annotator agreement sentences
To better understand why certain sentences had low agreement between annotators for RE assignments, OBDW manually sampled and inspected low agreement sentences for each RE category. Within an RE category, all low agreement sentences were used if there were fewer than 100, otherwise, sentences were sampled with priority given to sentences with lower agreement (Fig. 3).
Sampling tiers for sentences with low annotator agreement. Within the triangle are the number of votes in each tier, with the sampling percentage on the left side. For each tier, sentences were sampled up to 10% of the total low agreement sentences for a given RE category. Tiers are symmetric, represented by the left and right-side numbers within the triangle. For example, a sentence with 3 votes for a given category has 7 annotators who did not vote for that category, while a sentence with 7 votes for a category has 3 annotators who did not vote for that category.
Table 6 summarizes general observations for sentences with low agreement across RE categories. RE categories with no low agreement sentences are excluded from the table. Generally, certain indicators specific to each category commonly occurred in low agreement sentences. The reasons for which indicators could lead to low agreement are likely specific to each category. Another potential reason for diverse labeling opinions was how de-identified country mentions were interpreted or used. While specific RE categories could sometimes be inferred, No Information Indicated and Not Covered were common choices for sentences with de-identified indicators.
As previously noted, the two ethnicity categories non-Latino and No Information Indicated had lower agreement than other RE categories (Table 5). From the low agreement sentences for non-Latino we observed that annotators were split in how to use direct race mentions (e.g., “black”, “white”, “AA”) and certain language and country indicators (e.g., “French Creole”, “Chinese”) as either uninformative for or indicative of a patient being non-Latino. The category No Information Indicated often contained votes from annotators who might not have used the previously discussed direct race mentions to infer that someone is non-Latino. Additionally, there were multiple de-identified mentions under no information indicated, which could point to the need for a modified “Not Covered” category that is explicitly designed for de-identified text. For these two lower agreement categories (non-Latino and No Information Indicated), annotators seem to be split on how to use information to infer ethnicity that are not considered direct ethnicity mentions or language mentions like “Spanish”.
Low agreement examples from the non-Latino category, could indicate that there is room for interpretation on what kinds of phrases can be used to infer that someone is non-Latino. Previous research has noted that Latino patients often do not identify with OMB-defined race categories used in this work9,53, and it is possible that a similar phenomenon is occurring here for the category non-Latino. More specifically, certain physician annotators could have different views on how OMB-defined race and ethnicity categories should inform one another and view Latino/non-Latino as incongruent with certain racial information or just fluidly defined53,54,55,56. This can happen when hospital workers do not feel adequately trained to collect RE data54. The raw annotation data provides insight into the spectrum of potential interpretations by physicians.
Validating potential false negative RE assignment annotations
To validate potential false negatives, we first examined sentences with no majority vote positive RE assignment that included an indicator. Second, we examined false positive mistakes made by deep learning models trained to identify sentences with RE information. All sentences were examined by OBDW.
When examining sentences with indicators but no majority vote positive RE assignment, it was observed that de-identified tokens and diverse opinions on using indicators to infer RE were most common. De-identification issues aren’t necessarily false negatives, but rather limitations of the data and the absence of a category that explicitly handles de-identified RE information. The closest false negatives are the diverse opinions between physicians on how to use country or language indicators to infer RE. Examples of indicators include “Canada”, “Cantonese”, and “Portuguese”. Some of these examples may not actually be false negatives. For example, one sentence contained “Canada” as the only indicator and most annotators agreed that this did not convey any RE information. However, other examples do seem to reflect some of the variety in which physicians interpret indicators, such as a sentence with the indicator “Portuguese” that had votes for all RE categories except Asian and Native Hawaiian or Other Pacific Islander.
For the second approach examining false positive modelling mistakes, these models are further described in Bear Don’t Walk et al., and were trained, validated, and tested on the C-REACT dataset to identify sentences with information for the RE categories Black/AA, Asian, white, and Latino57. All modelling false positive sentences and vote counts were inspected. Upon examination, most false positives by the model were modelling mistakes, and did not indicate any mistakes by the annotators. In one case, the model assigned a label and our examination revealed that there could be a positive label by annotators. In this case, only four annotators had assigned the label Asian to a sentence with the indicator “Sri Lankan”.
Overall, these analyses into lower agreement sentences indicates that there are likely nuances with how certain textual data is used to infer RE labels by physicians. Additionally, we have attempted to manually identify and correct potential errors, where appropriate, to ensure high quality data. However, given the potential for rich interpretation by annotators there may be assignments that not all researchers will agree with. These different interpretations may be a signal to pause and look further into potential assumptions leading to an RE assignment. The voting data for each assignment provides both nuance and potential limitations, allowing users to investigate lower agreement labels.
Usage Notes
Data are publicly available through PhysioNet and are subject to their credentialing process and data use agreement. The credentialing process includes training on HIPAA, human subjects research, privacy and confidentiality, and principles to support the ethical conduct of research. Furthermore, users must sign a data use agreement to openly share code related to publications using MIMIC-III data while protecting data security and patient privacy. The authors affirm that they have followed these data use and ethical guidelines as well. Approved users can download data from the C-REACT dataset on PhysioNet project (https://doi.org/10.13026/t9ka-6k29)through PhysioNet.
We recommend using the Python library pandas to work with the provided files (e.g., the jsonl files may be read using the ‘read_json(‘file_name.jsonl’, lines=True)’ function. We provide code to work with the raw RE label annotation files and functions to determine the majority vote assignments. Additionally, we provide examples for working with indicator span data in ‘indicators_df.jsonl’. Working with the raw RE assignment files can be accomplished using the scripts and functions provided in the GitHub repository discussed in the Code Availability section below. While we provide a single jsonl file for the RE labeling, researchers should be aware that sentences from the shared annotation and single annotation subsets are differentiated using the “shared_subset” and “single_subset” columns. The columns “patient_id”, “visit_id” map to columns within the original MIMC-III data. The mappings from C-REACT to MIMIC-III are “patient_id” to “SUBJECT_ID”, and “visit_id” to “HADM_ID”.
If this corpus is used to train models to infer RE from clinical text, we suggest that researchers split training and test sets along patient ID rather than visit ID to limit data leakage and better emulate real-world settings. Finally, it should be noted that these sentences are not drawn randomly from MIMIC-III clinical notes and that we prioritized sentences with likely documentation of RE labels and indicators, while also drawing clinical notes without this information. This was done to balance identifying positive labels while limiting sampling bias. Please see Table 1 for more information on how these sentences are distributed in MIMIC-III clinical notes. Finally, researchers using the C-REACT dataset should be aware that while MIMIC-III offers a great opportunity to train models on real-world clinical data (often difficult to obtain given security and privacy concerns), MIMIC-III comes from a non-representative health organization in Boston, Massachusetts. While the indicators in this work likely sufficiently covered this population, they might not generalize to other populations or discussion of race and ethnicity-specifically for people from Native American, Alaskan Native, Hawaiian, and Pacific Islander populations, of which there is limited representation in our corpus.
Beyond technical usage notes, there are also notable ethical concerns. The C-REACT dataset is intended to inform future research about how granular RE-related information manifests in clinical notes and can be used to infer RE labels through NLP. While creating this dataset we balanced the importance of granular information with the established use of broader RE categories from the U.S. census. We encourage future researchers to be intentional and transparent about their assumptions and definitions for RE categories when using this dataset for whatever level of granularity. Importantly, researchers should consider if the federally recognized categories provided in this dataset are appropriate or if the more granular information from RE indicators are needed. Finally, self-reported RE data is still the reference standard and we cannot guarantee that what is reported in the clinical note is reflective of a patient’s self-reported racial and ethnic identity. Still, RE information derived from clinical notes can be used to complement self-reported RE information and mitigate missingness, while potentially providing more nuanced information. Because we strongly believe that granular information can provide key insights on discussions of broad RE categories and that RE categories are dependent on the research question at hand, we do not provide pre-defined training, validation, and test sets for benchmarking purposes.
RE labels played a dominant role in the analysis presented here. While RE labels can be used for sentence classification and RE indicators can be used for span level tasks, there are many nuances in how these two sets of annotations can be leveraged. Thus, we provide a non-exhaustive list of research projects that can make use of the indicator and/or label data. As previously mentioned, Bear Don’t Walk et al., used the RE labels to train models to identify sentences with positive RE mentions and assessed learned associations between textual inputs and model classifications57. C-REACT’s combination of sentence-level labels and span-level indicators allowed Bear Don’t Walk et al., to assess how well model-derived salient features aligned with the manually identified indicator spans and found that high classification performance may mask potentially concerning learned associations. Future research may leverage span-level features to augment label classification training while pushing models to use certain features and feature types58. Additionally, different interpretations between physician RE label annotations can be incorporated into model training to estimate uncertainty while improving task performance59. Researchers may leverage only the RE label data to train a model and interrogate differences between structured and text-based sources for RE data within the EHR. In the case that researchers choose to forgo the sentence-level labels, spans can be used for named entity recognition and leveraged in downstream analyses such as large-scale analysis into patterns of RE discussion for various patient groupings.
Code availability
All code is made publicly available through GitHub (https://github.com/elhadadlab/MIMIC_race_ethnicity_dataset). The code was run in an environment with Python version 3.9.4. and pandas60 version 1.2.4. Versions for all other libraries used can be found in the environment.yml the GitHub repository.
References
Stevens, L. A., Coresh, J., Greene, T. & Levey, A. S. Assessing Kidney Function — Measured and Estimated Glomerular Filtration Rate. N Engl J Med 354, 2473–2483 (2006).
Levey, A. S. et al. A New Equation to Estimate Glomerular Filtration Rate. Ann Intern Med 150, 604–612 (2009).
Gail, M. H. et al. Projecting individualized probabilities of developing breast cancer for white females who are being examined annually. J Natl Cancer Inst 81, 1879–1886 (1989).
Blumenthal, D. & Tavenner, M. The ‘meaningful use’ regulation for electronic health records. N Engl J Med 363, 501–504 (2010).
LaVeist, T. A., Gaskin, D. & Richard, P. Estimating the Economic Burden of Racial Health Inequalities in the United States. Int J Health Serv 41, 231–238 (2011).
Dorsey, R. et al. Implementing health reform: improved data collection and the monitoring of health disparities. Annu Rev Public Health 35, 123–138 (2014).
Douglas, M. D., Dawes, D. E., Holden, K. B. & Mack, D. Missed Policy Opportunities to Advance Health Equity by Recording Demographic Data in Electronic Health Records. Am J Public Health 105, S380–S388 (2015).
Kressin, N. R. Race/Ethnicity Identification: Vital for Disparities Research, Quality Improvement, and Much More Than “Meets the Eye”. Medical Care 53, 663–665 (2015).
Polubriaginof, F. C. G. et al. Challenges with quality of race and ethnicity data in observational databases. J Am Med Inform Assoc 26, 730–736 (2019).
Chakkalakal, R. J., Green, J. C., Krumholz, H. M. & Nallamothu, B. K. Standardized data collection practices and the racial/ethnic distribution of hospitalized patients. Med Care 53, 666–672 (2015).
Kressin, N. R., Chang, B.-H., Hendricks, A. & Kazis, L. E. Agreement between administrative data and patients’ self-reports of race/ethnicity. Am J Public Health 93, 1734–1739 (2003).
Institute of Medicine (US) Subcommittee on Standardized Collection of Race/Ethnicity Data for Healthcare Quality Improvement. Race, Ethnicity, and Language Data: Standardization for Health Care Quality Improvement. (National Academies Press (US), Washington (DC), 2009).
Nelson, A. Unequal treatment: confronting racial and ethnic disparities in health care. J Natl Med Assoc 94, 666–668 (2002).
Hasnain-Wynia, R., Pierce, D. & Pittman, M. A. Who, When, and How: The Current State of Race, Ethnicity, and Primary Language Data Collection in Hospitals. New York: Commonwealth Fund 42 (2004).
Magaña López, M., Bevans, M., Wehrlen, L., Yang, L. & Wallen, G. R. Discrepancies in Race and Ethnicity Documentation: a Potential Barrier in Identifying Racial and Ethnic Disparities. J Racial Ethn Health Disparities https://doi.org/10.1007/s40615-016-0283-3 (2016).
Zingmond, D. S. et al. Improving Hospital Reporting of Patient Race and Ethnicity—Approaches to Data Auditing. Health Serv Res 50, 1372–1389 (2015).
Hatef, E. et al. Assessing the Availability of Data on Social and Behavioral Determinants in Structured and Unstructured Electronic Health Records: A Retrospective Analysis of a Multilevel Health Care System. JMIR Med Inform 7, e13802 (2019).
Rights (OCR), O. for C. Section 1557 of the Patient Protection and Affordable Care Act. https://www.hhs.gov/civil-rights/for-individuals/section-1557/index.html (2010).
Sun, T. Y., Bhave, S. A., Altosaar, J. & Elhadad, N. Assessing Phenotype Definitions for Algorithmic Fairness. AMIA Annu Symp Proc 2022, 1032–1041 (2023).
Wornow, M. et al. The shaky foundations of large language models and foundation models for electronic health records. npj Digit. Med. 6, 1–10 (2023).
Kleinberg, J., Mullainathan, S. & Raghavan, M. Inherent Trade-Offs in the Fair Determination of Risk Scores (2016).
Gordon, N. P., Lin, T. Y., Rau, J. & Lo, J. C. Aggregation of Asian-American subgroups masks meaningful differences in health and health risks among Asian ethnicities: an electronic health record based cohort study. BMC Public Health 19, 1551 (2019).
Griffith, D. M., Moy, E., Reischl, T. M. & Dayton, E. National Data for Monitoring and Evaluating Racial and Ethnic Health Inequities: Where Do We Go From Here? Health Education & Behavior 33, 470–487 (2006).
Ford, C. L. & Harawa, N. T. A new conceptualization of ethnicity for social epidemiologic and health equity research. Soc Sci Med 71, 251–258 (2010).
Wang, K. et al. Information Loss in Harmonizing Granular Race and Ethnicity Data: Descriptive Study of Standards. J Med Internet Res 22, e14591 (2020).
Islam, N. S. et al. Methodological issues in the collection, analysis, and reporting of granular data in Asian American populations: historical challenges and potential solutions. J Health Care Poor Underserved 21, 1354–1381 (2010).
Bilheimer, L. T. & Klein, R. J. Data and Measurement Issues in the Analysis of Health Disparities. Health Serv Res 45, 1489–1507 (2010).
Mays, V. M., Ponce, N. A., Washington, D. L. & Cochran, S. D. Classification of race and ethnicity: implications for public health. Annu Rev Public Health 24, 83–110 (2003).
Revisions to the Standards for the Classification of Federal Data on Race and Ethnicity. The White House https://obamawhitehouse.archives.gov/node/15626.
Kaneshiro, B., Geling, O., Gellert, K. & Millar, L. The Challenges of Collecting Data on Race and Ethnicity in a Diverse, Multiethnic State. Hawaii Med J 70, 168–171 (2011).
Sentell, T., Shumway, M. & Snowden, L. Access to mental health treatment by English language proficiency and race/ethnicity. J Gen Intern Med 22(Suppl 2), 289–293 (2007).
Gomez, S. L. et al. Cancer incidence trends among Asian American populations in the United States, 1990-2008. J Natl Cancer Inst 105, 1096–1110 (2013).
Villarroel, M. A., Clarke, T. C. & Norris, T. Health of American Indian and Alaska Native Adults, by Urbanization Level: United States, 2014–2018. NCHS Data Brief 1–8 (2020).
Movva, R. et al. Coarse race data conceals disparities in clinical risk score performance. In: Proceedings of the 8th Machine Learning for Healthcare Conference. PMLR, pp 443–472 (2023).
Ross, J., Hanna, D. B., Felsen, U. R., Cunningham, C. O. & Patel, V. V. Emerging from the database shadows: characterizing undocumented immigrants in a large cohort of HIV-infected persons. AIDS Care 29, 1491–1498 (2017).
Farber-Eger, E., Goodloe, R., Boston, J., Bush, W. S. & Crawford, D. C. Extracting Country-of-Origin from Electronic Health Records for Gene- Environment Studies as Part of the Epidemiologic Architecture for Genes Linked to Environment (EAGLE) Study. AMIA Jt Summits Transl Sci Proc 2017, 50–57 (2017).
Sholle, E. T. et al. Underserved populations with missing race ethnicity data differ significantly from those with structured race/ethnicity documentation. J Am Med Inform Assoc 26, 722–729 (2019).
Johnson, A. E. W. et al. MIMIC-III, a freely accessible critical care database. Sci Data 3, 160035 (2016).
Rosenman, E. T. R., Olivella, S. & Imai, K. Race and ethnicity data for first, middle, and surnames. Sci Data 10, 299 (2023).
Bird, S., Klein, E. & Loper, E. Natural Language Processing with Python: Analyzing Text with the Natural Language Toolkit. (O’Reilly Media, Beijing; Cambridge Mass., 2009).
Montani, I. & Honnibal, H. Prodigy: A new annotation tool for radically efficient machine teaching. Artificial Intelligence to appear, (2018).
Hripcsak, G. & Rothschild, A. S. Agreement, the F-Measure, and Reliability in Information Retrieval. J Am Med Inform Assoc 12, 296–298 (2005).
Deleger, L. et al. Building Gold Standard Corpora for Medical Natural Language Processing Tasks. AMIA Annu Symp Proc 2012, 144–153 (2012).
Noe-Bustamante, L., Mora, L. & Lopez, M. H. About One-in-Four U.S. Hispanics Have Heard of Latinx, but Just 3% Use It. Pew Research Center’s Hispanic Trends Project https://www.pewresearch.org/hispanic/2020/08/11/about-one-in-four-u-s-hispanics-have-heard-of-latinx-but-just-3-use-it/ (2020).
Rumbaut, R. G. The Making of a People. https://papers.ssrn.com/abstract=1877405 (2006).
What does ‘Latinx’ mean? A look at the term that’s challenging gender norms. Complex https://www.complex.com/life/2016/04/latinx/.
García, I. Cultural Insights for Planners: Understanding the Terms Hispanic, Latino, and Latinx. Journal of the American Planning Association 86, 393–402 (2020).
The Associated Press. The Associated Press Stylebook 2019: And Briefing on Media Law. (Basic Books, an imprint of Perseus Books, LLC., New York, NY, 2019).
Flanagin, A., Frey, T., Christiansen, S. L. & Manual of Style Committee, A. M. A. Updated Guidance on the Reporting of Race and Ethnicity in Medical and Science Journals. JAMA 326, 621–627 (2021).
Bear Don’t Walk, O. J. et al. C-REACT: Contextualized Race and Ethnicity Annotations for Clinical Text. physionet.org https://doi.org/10.13026/C2JT1Q (2023).
Goldberger, A. L. et al. PhysioBank, PhysioToolkit, and PhysioNet: components of a new research resource for complex physiologic signals. Circulation 101, E215–220 (2000).
McHugh, M. L. Interrater reliability: the kappa statistic. Biochem Med (Zagreb) 22, 276–282 (2012).
Bhalla, R., Yongue, B. G. & Currie, B. P. Standardizing Race, Ethnicity, and Preferred Language Data Collection in Hospital Information Systems: Results and Implications for Healthcare Delivery and Policy. Journal for Healthcare Quality 34, 44–52 (2012).
Berry, C., Kaplan, S. A., Mijanovich, T. & Mayer, A. Moving to patient reported collection of race and ethnicity data: implementation and impact in ten hospitals. Int J Health Care Qual Assur 27, 271–283 (2014).
Robbin, A. The problematic status of u.s. statistics on race and ethnicity: An “imperfect representation of reality”. Journal of Government Information 26, 467–483 (1999).
Cook, L. A., Sachs, J. & Weiskopf, N. G. The quality of social determinants data in the electronic health record: a systematic review. J Am Med Inform Assoc 29, 187–196 (2021).
Bear Don’t Walk, IV et al. Auditing Learned Associations in Deep Learning Approaches to Extract Race and Ethnicity from Clinical Text. AMIA Annu Symp Proc 2023, 289–298 (2024).
Ross, A. S., Hughes, M. C. & Doshi-Velez, F. Right for the Right Reasons: Training Differentiable Models by Constraining their Explanations. in Proceedings of the Twenty-Sixth International Joint Conference on Artificial Intelligence 2662–2670, https://doi.org/10.24963/ijcai.2017/371 (International Joint Conferences on Artificial Intelligence Organization, Melbourne, Australia, 2017).
Davani, A. M., Díaz, M. & Prabhakaran, V. Dealing with Disagreements: Looking Beyond the Majority Vote in Subjective Annotations. Transactions of the Association for Computational Linguistics 10, 92–110 (2022).
Mckinney, W. Data Structures for Statistical Computing in Python. Proceedings of the 9th Python in Science Conference (2010).
Acknowledgements
The authors are grateful for the guidance and feedback from Adler Perotte and Karthik Natarajan. The authors would also like to thank Alistair Johnson and Travis Goodwin for their help with the publication process. This work was supported by grants from the National Library of Medicine (OBDW, HRN, TS, AP: T15LM007079, LRR: R01 LM006910), Agency for Healthcare Research and Quality (BR: T32HS026121), Intramural Research Program of the National Library of Medicine and National Institutes of Health (BDW, DDF), and a Computational and Data Science Fellowship from the Association for Computing Machinery Special Interest Group in High Performance Computing (HRN). The study funders had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication. The content is solely the responsibility of the authors and does not necessarily represent the official views of a funder. OBDW completed much of this work at Columbia University, but finished writing and editing while at the University of Washington, their current institution.
Author information
Authors and Affiliations
Contributions
All authors have contributed to the drafting and revision process and have approved the final manuscript. Oliver J. Bear Don’t Walk IV – Study concept and design, data acquisition, data validation. Adrienne Pichon – Study concept and design, data acquisition. Harry Reyes Nieva – Study concept and design, data acquisition. Tony Sun – Study concept and design, data acquisition. Jaan Altosaar – Study concept and design. Josh Joseph – Data acquisition. Sivan Kinberg – Data acquisition. Lauren R. Richter – Data acquisition. Salvatore Crusco – Data acquisition. Kyle Kulas – Data acquisition. Shaan A. Ahmed – Data acquisition. Daniel Snyder – Data acquisition. Ashkon Rahbari – Data acquisition. Benjamin L. Ranard – Data acquisition. Pallavi Juneja – Data acquisition. Dina Demner-Fushman – Supervision, study concept and design. Noémie Elhadad – Supervision, study concept and design.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bear Don’t Walk, O.J., Pichon, A., Reyes Nieva, H. et al. Contextualized race and ethnicity annotations for clinical text from MIMIC-III. Sci Data 11, 1332 (2024). https://doi.org/10.1038/s41597-024-04183-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-024-04183-2