Abstract
Motor dysfunction is one of the most significant sequelae of stroke, with lower limb impairment being a major concern for stroke patients. Motor imagery (MI) technology based on brain-computer interface (BCI) offers promising rehabilitation potential for stroke patients by activating motor-related brain areas. However, developing a robust BCI-MI system and uncovering the underlying mechanisms of neural plasticity during stroke recovery through such systems requires large-scale datasets. These datasets are particularly needed for accurate lower limb MI in stroke patients and for longitudinal data reflecting the rehabilitation process. This study addresses this gap by collecting EEG data from 27 stroke patients, covering two enhanced paradigms and three different time points. The dataset includes raw EEG signals, preprocessed data, and patient information. An initial analysis using CSP-SVM on the dataset yielded an average classification accuracy of 80.50%. We anticipate that this dataset will facilitate research into brain neuroplasticity in stroke patients, aid in the development of decoding algorithms for lower limb stroke, and contribute to the establishment of comprehensive stroke rehabilitation systems.
Similar content being viewed by others
Background & Summary
Each year, more than 12.2 million new cases of stroke are reported globally1, with motor dysfunction being the most prominent and disabling sequela2,3. This loss of motor function severely impacts the independence and social participation of stroke survivors. Consequently, the restoration of lower limb motor function and walking ability is one of the most pressing needs for stroke patients.
Brain-computer interface (BCI) have demonstrated significant potential in the field of neurorehabilitation4,5. In particular, motor imagery (MI) based on BCI has been increasingly applied to stroke rehabilitation in recent years6,7,8,9. BCI-MI has become an effective therapeutic aid in stroke rehabilitation, facilitating neural plasticity10 and improving motor function11. However, the rehabilitation success of BCI-MI largely depends on its decoding accuracy12, and achieving stable high recognition rates for MI signals has become a key area of focus in the field13. To improve decoding algorithm, high-quality open datasets are indispensable14.
Although several publicly available MI datasets from healthy participants exist for researchers15,16,17,18,19, these datasets may not be reliable for real clinical applications, as stroke patients exhibit significantly weaker neural activation during MI tasks compared to healthy individuals20. Targeted datasets focusing on stroke patients are also essential for better understanding their brain activation patterns. Such data could help improve training paradigms, enhance the overall usability and stability of MI systems, and potentially improve rehabilitation outcomes. However, there is currently a scarcity of MI datasets specifically for stroke patients. Liu et al.21 established the first MI dataset for acute stroke patients, which included the conventional paradigm for upper limb MI tasks. They highlighted the importance of having a dataset with high classification accuracy. However, compared to upper limb MI, lower limb MI responses are more challenging to detect due to the corresponding brain regions being located deep within the sulci22, resulting in weaker characteristic responses and poorer classification performance, making the data less effective. To address this, enhanced MI paradigms have been proposed to improve data quality. These include composite limb MI paradigms22,23, visually guided MI paradigms24, and somatosensory-stimulated MI paradigms25,26. Among these, somatosensory-stimulated MI paradigms show unique potential for future clinical applications, as they do not overly consume the participant’s attention or occupy visual or auditory channels. Hence, we27,28 have built three MI paradigms in this dataset: gait-phase-encoded sequential electrical stimulation paradigm (SES), invariable electrical stimulation paradigm (IES), and a conventional paradigm without electrical stimulation. The goal is to facilitate the development of highly effective lower limb MI paradigms and algorithms for stroke patients. There is also a lack of publicly available longitudinal patient MI datasets, which are crucial for understanding the neural plasticity mechanisms during the rehabilitation process. Therefore, to address the gaps in BCI datasets and to facilitate the development of high-performance EEG decoding algorithms, it is of great importance to establish a high-quality, multi-paradigm, longitudinal lower limb MI dataset specifically for stroke patients.
In this study, we collected data from 27 stroke recovery patients, including EEG recordings of MI for the affected leg and idle-state data under two types of sensory stimulation-enhanced paradigms, across three distinct time periods. 4,260 lower limb MI EEG trials were collected. The public dataset comprises three components: raw EEG data, preprocessed data, and patient information. The preprocessed data can be used to generate result plots and perform classification directly using the provided code. Researchers may also reuse the dataset according to their specific needs. To our knowledge, this is the first study to provide a large-scale MI dataset for stroke patients that simultaneously includes lower limb MI, multiple enhancement paradigms, and longitudinal data.
Using this dataset, we conducted a preliminary comparison between the EEG data obtained during lower limb MI and the idle state in stroke patients. We performed a binary classification task using baseline data and classical classification methods, achieving an average classification accuracy of 80.50%. This accuracy is comparable to recent studies of lower limb MI decoding in healthy subjects (RCM: 82.29%29, TRCA: 81.07%30) and in stroke patients (LDA: 79.17%31), demonstrating that the collected EEG data can be classified based on the execution of MI tasks. We expect that our dataset will help address the challenges in recognizing lower limb MI features, enhance the understanding of neural plasticity mechanisms underlying lower limb rehabilitation in stroke patients, and support the development of robust, long-term brain-computer interface systems.
Methods
Patients
This study recruited stroke patients from Tianjin Huanhu Hospital. The inclusion criteria were as follows: (1) Patients met the diagnostic criteria for stroke and were confirmed by head CT or MRI; (2) Stable vital signs, normal cognitive function, and unimpaired communication ability; (3) Disease duration of 1 to 12 months and age between 30 and 70 years.
Exclusion criteria included: (1) Patients with motor dysfunction caused by other diseases; (2) Patients with significant cognitive, language, or communication impairments that prevented them from cooperating with the study; (3) Patients with Achilles tendon contracture or inability to place their heels on the ground; (4) Patients with severe cardiac conditions or implanted cardiac pacemakers.
A total of 27 post-stroke patients in the recovery phase were recruited for the study. The patients comprised 23 males (85%) and 4 females (15%), aged between 33 and 68 years, with an average age of 51.44 years (SD = 9.31). Among the patients, 18 (67%) had right hemiplegia, and 9 (33%) had left hemiplegia. All participants were thoroughly informed about the experimental procedures and provided written consent for the collection and anonymized publication of their data. The study was approved by the Ethics Committee of Tianjin Huanhu Hospital (IEC-B-013-V3.0).
Experimental Paradigms
This dataset includes five distinct experiments: the initial assessment with a conventional paradigm prompted by text (Pre), initial assessment with an invariable electrical stimulation paradigm (IES), initial assessment with a gait-phase-encoded sequential electrical stimulation paradigm (SES), post-treatment assessment with a conventional paradigm (Post), and follow-up assessment with a conventional paradigm (Follow). Each experiment comprised two tasks: gait MI and a idle-state task, where participants were instructed to focus on kinesthetic motor imagery to perform the gait MI task.
In the conventional paradigm, patients receive fixed monochromatic circles as visual cues, and in the MI tasks, patients are presented with gait phase text prompts numbered 1-4 without stimulation. On top of this, the IES paradigm applies constant electrical stimulation to the affected leg, while the SES paradigm delivers gait-phase-encoded stimulation27. Both paradigms aim to enhance patients’ attention to their legs. Before initiating the experiment, electrodes were placed on the sural and superficial peroneal nerves of the affected leg, and the pulse width of the electrical stimulation was calibrated to ensure the patient could distinguish between strong and weak stimuli without experiencing pain. In the IES paradigm, constant strong stimulation was applied to both nerves simultaneously during the MI task, starting from the cue appearance until the task ended. In the SES paradigm, time-varying stimulation was delivered during the task in the following sequence: strong stimulation to the sural nerve for 1 second, followed by strong stimulation to both nerves for 0.6 seconds, strong stimulation to the superficial peroneal nerve for 1.8 seconds, and finally weak stimulation to both nerves for 1.6 seconds. The Pre, Post, and Follow sessions serve as longitudinal control groups using a traditional paradigm, differing only in the timing of data collection. Although monochromatic circles and numeric cues were used as visual prompts, these cues were not designed to flicker at fixed frequencies capable of eliciting steady-state visual evoked potentials (SSVEPs); rather, they were utilized as static cues. Additionally, the frequency band of MI features (8-25 Hz) differs from the low-frequency range that these visual prompts might influence. This ensures that our paradigms facilitate MI in patients rather than inducing SSVEP.
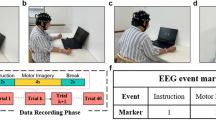
The experimental scenario is shown in Fig. 1. Each patient participated in at least three paradigm experiments or three longitudinal experiments. In the following it is noted as the paradigm group and the longitudinal group. Experiments in which each patient participated are shown by Table 1.
Experimental equipment and scenarios. (a) The Scene of the experiment. (b) Position of the stimulating electrodes.
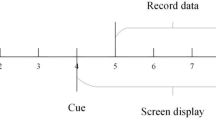
Each trial lasted approximately 12 seconds. At the beginning of each trial (t = 0s), a fixation cross appeared on the screen to direct the participant’s attention. During this preparation phase, MI tasks were labeled as “1” and idle-state tasks as “7.” At t = 3s, the task text prompt was displayed. For the SES and IES tasks, a weak electrical stimulation at weak level was applied at the electrode sites. Stimulus levels have been measured in preparation for the experiment. During this cueing phase, MI tasks were labeled as “2” and idle-state tasks as “8.” At t = 4s, participants were asked to imagine the corresponding state for 5 seconds according to the task prompt. For gait MI tasks, a number encircled in orange appeared on the screen, indicating the phase of gait imagery required. During the MI phase, the labels “3” to “6” were used sequentially according to the gait phase. For the idle-state tasks, a black circle was displayed, and the stimulation method was the same as in the MI task. Participants were instructed to disregard the stimulation and clear their minds, with the idle-state phase labeled sequentially as “9” to “12.” After each task, a 3-second relaxation period allowed participants to rest before the next trial. In each paradigm, participants performed 40 trials each of gait MI and idle tasks, presented in a randomized order, as depicted in Fig. 2.
The experiment paradigm.
Data Collection & Preprocessing
Before the experiment, participants watched a gait task video to familiarize themselves with the paradigm and experimental procedures. EEG data were collected using a NeuSenW amplifier (Neuracle, Inc.) with a 64-channel Ag/AgCl electrode cap, sampled at 1000 Hz. During the recording, impedance was kept below 10 kΩ. A 50 Hz notch filter was applied to remove power line noise, and electrodes were placed according to the international 10/20 system. Given the weak signal and the presence of artifacts such as eye movements and electromyography (EMG) noise, rigorous preprocessing was necessary to minimize the impact of artifacts. The specific preprocessing steps included: (1) downsampling the signals to 250 Hz and applying a 3-35 Hz bandpass filter to isolate EEG signals within the MI task-relevant frequency band; (2) re-referencing the filtered data using a common average reference; (3) manually removing data segments with obvious motion artifacts; (4) performing ICA to manually remove eye movement and EMG artifacts based on the temporal-frequency characteristics and topographical maps of the components, aided by the ICLabel plugin in the EEGLAB toolbox; (5) extracting data segments corresponding to the target labels.
The header of the table of specific information for all patients is shown in Table 2.
Data Records
Our dataset is available in the Figshare repository32, organized according to the Brain Imaging Data Structure (BIDS)33, a widely-used format for open EEG datasets.
The dataset structure is illustrated in Fig. 3. The repository is divided into four main sections: (1) Preprocessed data, stored as ‘.set’ files; (2) Raw data, containing the original EEG recordings for each subject throughout the entire experiment, stored as ‘.set’ files; (3) Additional information, including patient details, electrode locations, and EEG event markers; and (4) Code, provided as MATLAB scripts.
The dataset structure.
Preprocessed Data
This section contains segmented data produced by preprocessing scripts. The data for each subject is stored in a folder named ‘sub-xx,’ where ‘xx’ represents the subject ID. The files are named according to the following convention:
where ‘xx’ is the subject ID, ‘group’ indicates the experiment (Pre, IES, SES, Post and Follow), and ‘y’ specifies the run number. Loading these files with the EEGLAB toolbox generates a structure variable named ‘EEG.’ Within this variable, ‘EEG.data’ stores the EEG recordings for the trial in a “channels × time samples” format, with 40 channels and 50,000 time samples. ‘EEG.chanlocs’ contains the location information for each channel, and ‘EEG.event’ records event labels and their corresponding timestamps, following the labeling conventions described in the paradigm explanation.
Raw Data
This section includes the unprocessed, unsegmented EEG data recorded for each subject throughout the entire experiment. The original data files already include the EEG channel location information, so no additional import is necessary. These files can be loaded using the EEGLAB MATLAB plugin to obtain the ‘EEG’ structure variable. The naming convention is identical to that of the preprocessed data as follow:
Code
The “code” folder contains all the scripts and functions used for data processing and for generating the result figures.
Technical Validation
ERD/ERS Analysis
Event-Related Spectral Perturbation (ERSP) is a widely utilized time-frequency analysis method for evaluating MI tasks34. It reflects changes in spectral power in the EEG relative to the average event-related stimulus across the time-frequency domain. The ERSP value, computed through the averaging of multiple trials, is defined as follows:
where n represents the number of trials, and Fk(f, t) denotes the spectral estimate at frequency f and time t for the kth trial. The ERSP values are extracted using short-time Fourier transform (STFT) with a Hamming window of 1024 points. The baseline is set to 1 second before the task onset, and the ERSP is calculated by subtracting the mean spectral values during the baseline from the spectral values after the task onset. This study primarily analyzes ERSP values within the 5-35 Hz frequency range, focusing on the time window from -1 to 6 seconds.
Topographical maps provide a visual representation of the brain regions involved in Event-Related Desynchronization (ERD) during different tasks. To further quantify the spatial distribution of cortical activation during MI tasks, the mean ERSP values were computed for 40 channels (excluding peripheral ones) in the 1-6 second window after task cue onset. Based on these, the average ERD values were calculated for the alpha (8-13 Hz) and beta (13-30 Hz) bands, as defined by the following formula:
where band represents the frequency band of interest, F denotes the specific frequency range within the band, T is the number of time sampling points within the calculation window, and N is the total number of data points in the time-frequency domain. The average ERD values were computed across the 40 EEG channels for the three experimental paradigms. The grand average ERD topographical maps were then generated by averaging the results across all subjects.
Figure 4 presents the average ERD topographical maps for the two groups, and Fig. 5 shows the average time-frequency plots from the Cz electrode and its surrounding electrodes (C1, C2, FCz and CPz), representing the primary foot motor area35. The areas outlined by dashed lines in Fig. 4 correspond to the electrodes selected for analysis in Fig. 5. In both figures, blue regions indicate ERD phenomena, signifying the activation of the sensorimotor cortex during lower limb MI. In all experiments, MI tasks triggered alpha and beta band activation in the sensorimotor cortex shortly after task onset. In contrast, no discernible activation was observed during the Idle task in any of the groups, as shown in the dashed regions of Fig. 4 and in Fig. 5.
The average ERD topographical maps for the paradigm experiment group and longitudinal experiment group. The foot-related feature region is delineated with dashed lines.
The average time-frequency plots at the Cz electrode and its surrounding electrodes for the paradigm experiment group and longitudinal experiment group. Task prompts are marked with black lines and the initiation of the MI task is denoted by the red dashed lines.
Classification Accuracy
In this study, each gait MI task lasted 5 seconds. To obtain more training samples and improve classification performance, the 5-second EEG data was divided into two segments, treating each 2.5-second window as a separate sample for feature extraction and classification. The CSP method, widely applied in MI-BCI binary classification, was used to extract MI features. The resulting feature vectors were then classified using SVM. A 10-fold cross-validation was performed on the test dataset for each experimental condition. The test data was randomly divided into 10 groups, with each group serving as the test set while the remaining 9 groups were used as the training set. This process was repeated 10 times, and the average of all results was taken as the final classification accuracy.
Figure 6 displays the classification accuracy results for all participants across the five experiments. The average decoding accuracies for the Pre, IES, SES, Post, and Follow experiments were 78.92%(paradigm: 78.53%, longitudinal: 79.51%), 80.57%, 82.41%, 80.95%, and 80.52%, respectively. Comparing these results with recent studies on lower limb motor imagery (RCM: 82.29%29, TRCA: 81.07%30, LDA: 79.17%31), the classification accuracies obtained using this dataset are consistent with those from other datasets, indicating the reliability of this dataset.
The decoding accuracy for all patients in the Pre, IES, SES, Post, and Follow experiments. Mean values for each group are marked with a blue dash.
Usage Notes
When using this dataset, it is important to consider the following: For stroke patients, completing extended motor imagery tasks can be challenging. Due to individual patient preferences or disease-related factors, not all patients completed every experimental task. Specifically, both sub-03 and sub-08 have only 20 trials each of gait MI and idle tasks in the IES experiment. Additionally, there is also missing information for some patients (e.g., affected area). Given the critical role of MI in neurological rehabilitation for patients, our experiment exclusively collected MI data from the affected leg of the participants. However, MI data from the unaffected leg could provide valuable insights into the neural rehabilitation mechanisms of stroke patients and contribute to the development of robust and versatile BCI rehabilitation systems. To address this limitation, we plan to collect EEG data from the unaffected leg in future studies.
Code availability
All MATLAB scripts used for preprocessing, image generation, and classification can be found in the ‘code’ folder. Researchers need to download the EEGLAB toolbox to run these scripts. Please note that the file path format used during our data processing differs from the BIDS format. To use the code, the original data must be reorganized according to the file path format specified in the provided notes.
References
Feigin, V. et al. Global, regional, and national burden of stroke and its risk factors, 1990-2019: A systematic analysis for the global burden of disease study 2019. The Lancet Neurology 20, 795–820 (2021).
Du, J. et al. One-year disability trajectories and long-term cardiovascular events, recurrent stroke, and mortality after ischemic stroke. Journal of the American Heart Association 13, e030702 (2024).
Wang, P. et al. In-hospital medical complications associated with stroke recurrence after initial ischemic stroke: A prospective cohort study from the china national stroke registry. Medicine 95, e4929 (2016).
Fu, Y., Chen, X. & Hu, Y. Correct understanding of brain–computer interfaces. Journal of Neurorestoratology 12(3), 100139, https://doi.org/10.1016/j.jnrt.2024.100139 (2024).
Sun, Y., Chen, X. & Gao, X. The potential of diverse brain–computer interface signal acquisition techniques in neurorestoratology. Journal of Neurorestoratology 12(3), 100138, https://doi.org/10.1016/j.jnrt.2024.100138 (2024).
Tang, Z. et al. An upper-limb rehabilitation exoskeleton system controlled by mi recognition model with deep emphasized informative features in a vr scene. IEEE Transactions on Neural Systems and Rehabilitation Engineering 31, 4390–4401 (2023).
Jochumsen, M. et al. Induction of neural plasticity using a low-cost open source brain-computer interface and a 3d-printed wrist exoskeleton. Sensors 21 (2021).
Barria, P. et al. Bci-based control for ankle exoskeleton t-flex: Comparison of visual and haptic stimuli with stroke survivors. Sensors 21 (2021).
Huang, J., Lin, M., Fu, J., Sun, Y. & Fang, Q. An immersive motor imagery training system for post-stroke rehabilitation combining vr and emg-based real-time feedback. In 2021 43rd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), 7590–7593 (2021).
Xu, M., Wei, Z. & Ming, D. Research advancements of motor imagery for motor function recovery after stroke. Journal of biomedical engineering 37, 169–173 (2020).
Birinci, T., Mutlu, E. & Altun, S. The efficacy of graded motor imagery in post-traumatic stiffness of elbow: A randomized controlled trial. Journal of shoulder and elbow surgery / American Shoulder and Elbow Surgeons ... [et al.] (2022).
Bundy, D. et al. Contralesional brain-computer interface control of a powered exoskeleton for motor recovery in chronic stroke survivors. Stroke 48 (2017).
Ahn, M. & Jun, S. Performance variation in motor imagery brain-computer interface: A brief review. Journal of neuroscience methods 243 (2015).
Jeong, J.-H. et al. 2020 international brain–computer interface competition: A review. Frontiers in Human Neuroscience 16, 898300 (2022).
Blankertz, B. et al. The bci competition iii: Validating alternative approachs to actual bci problems. IEEE transactions on neural systems and rehabilitation engineering : a publication of the IEEE Engineering in Medicine and Biology Society 14, 153–9 (2006).
Cho, H., Ahn, M., Ahn, S., Kwon, M. & Jun, S. Eeg datasets for motor imagery brain computer interface. GigaScience 6 (2017).
Dreyer, P., Roc, A., Pillette, L., Rimbert, S. & Lotte, F. A large eeg database with users’ profile information for motor imagery brain-computer interface research. Scientific Data 10, 580 (2023).
Ma, J. et al. A large eeg dataset for studying cross-session variability in motor imagery brain-computer interface. Scientific Data 9, 531 (2022).
Gwon, D. et al. Review of public motor imagery and execution datasets in brain-computer interfaces. Frontiers in Human Neuroscience 17, 1134869 (2023).
Platz, T. et al. Multimodal eeg analysis in man suggests impairment-specific changes in movement-related electric brain activity after stroke. Brain 123, 2475–2490 (2000).
Liu, H. et al. An eeg motor imagery dataset for brain computer interface in acute stroke patients. Scientific Data 11 (2024).
Ma, R., Chen, Y.-F., Jiang, Y. & Zhang, M. A new compound-limbs paradigm: Integrating upper-limb swing improves lower-limb stepping intention decoding from eeg. IEEE Transactions on Neural Systems and Rehabilitation Engineering PP, 1–1 (2023).
Yi, W. et al. Eeg oscillatory patterns and classification of sequential compound limb motor imagery. Journal of NeuroEngineering and Rehabilitation 13 (2016).
Lei, B. et al. Walking imagery evaluation in brain computer interfaces via a new deep learning framework. IEEE Transactions on Neural Systems and Rehabilitation Engineering PP, 1–1 (2019).
Shixin, R. et al. Enhanced motor imagery based brain-computer interface via fes and vr for lower limbs. IEEE Transactions on Neural Systems and Rehabilitation Engineering PP, 1–1 (2020).
Shu, X. et al. Tactile stimulation improves sensorimotor rhythm-based bci performance in stroke patients. IEEE transactions on bio-medical engineering PP (2018).
Zhang, Y. et al. Enhanced motor imagery of lower limbs induced by gait phase encoding sensory electrical stimulation. In 2022 IEEE International Conference on Robotics and Biomimetics (ROBIO) (2022).
Gui, Z. et al. Electrical stimulation-based paradigm to enhance lower limb motor imagery: initial validation in stroke patients. In 2024 46th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), 1–4 (2024).
Delisle Rodriguez, D., Silva, L. & Bastos-Filho, T. Eeg changes during passive movements improve the motor imagery feature extraction in bcis-based sensory feedback calibration. Journal of Neural Engineering 20 (2023).
Bian, Y. et al. Improvements in classification of left and right foot motor intention using modulated steady-state somatosensory evoked potential induced by electrical stimulation and motor imagery. IEEE Transactions on Neural Systems and Rehabilitation Engineering 31, 150–159 (2023).
Romero Laiseca, M. et al. A low-cost lower-limb brain-machine interface triggered by pedaling motor imagery for post-stroke patients rehabilitation. IEEE Transactions on Neural Systems and Rehabilitation Engineering PP, 1–1 (2020).
Gui, Z., Liu, Y., Wang, Z., Gao, R. & Han, N. Lower limb motor imagery eeg dataset based on the multi-paradigm and longitudinal-training of stroke patients. figshare (2024).
Pernet, C. et al. Eeg-bids, an extension to the brain imaging data structure for electroencephalography. Scientific Data 6 (2019).
Makeig, S., Debener, S., Onton, J. & Delorme, A. Mining event-related brain dynamics. Trends in cognitive sciences 8, 204–10 (2004).
Pfurtscheller, G., Brunner, C., Schlögl, A. & Lopes da Silva, F. Mu rhythm (de)synchronization and eeg single-trial classification of different motor imagery tasks. NeuroImage 31, 153–159 (2006).
Acknowledgements
This work was supported in part by the National Key R& D Program of China (2023YFC3603800, 2023YFC3603802), National Natural Science Foundation of China (62273251), Research Project of State Key Laboratory of Mechanical System and Vibration (MSV202418), Tianjin Science and Technology Plan Project (21JCYBJC00420, 21JCYBJC00520).
Author information
Authors and Affiliations
Contributions
Yuan Liu conceived the experiments, Zhuolan Gui, De Yan, Ruisi Gao and Ningxin Han conducted the experiments, Zhuolan Gui analysed the data and wrote the code. Zhuolan Gui, Yuan Liu and Zhuang Wang wrote the paper. Junying Chen and Jialing Wu provided the site, recruited patients and provided clinical support. Yuan Liu, Junying Chen and Jialing Wu supervised the experiments. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, Y., Gui, Z., Yan, D. et al. Lower limb motor imagery EEG dataset based on the multi-paradigm and longitudinal-training of stroke patients. Sci Data 12, 314 (2025). https://doi.org/10.1038/s41597-025-04618-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-04618-4
This article is cited by
-
An EEG dataset for handwriting imagery decoding of Chinese character strokes and Pinyin single vowels
Scientific Data (2026)
-
A multi-paradigm EEG dataset for studying upper limb rehabilitation exercises
Scientific Data (2025)
-
Enhancing classification of a large lower-limb motor imagery EEG dataset for BCI in knee pain patients
Scientific Data (2025)