Abstract
Weather radars detect more than weather, they also continuously register the movements of billions of animals aloft in the lower atmosphere. This makes archived, unfiltered weather radar data a goldmine for biological monitoring purposes, providing coverage of the aerial habitat in a way no other method can. Here we present two datasets of biological data extracted from European weather radar data, obtained through a collaboration with the Operational Programme for the Exchange of Weather Radar Information (OPERA) and three national meteorological services. The datasets were created by processing weather radar data with methods optimized for extracting bird targets, resulting in vertical profiles of biological targets. The datasets collectively cover 141 radar stations in 18 countries, from 2008 to 2023. Data quality and coverage differs between years, countries, and radar stations, so care must be taken when evaluating data for each specific use case. Despite these challenges the datasets are currently the most comprehensive of their kind in Europe and open new avenues in understanding continental scale movements of aerial animals.
Similar content being viewed by others
Background & Summary
Since the second world war, the airspace across the world has been monitored by an ever-growing network of silent sentinels, constantly interrogating the skies. Specialized radars monitor all forms of movement in the air, and one of the most widespread networks globally is that of weather surveillance radars. Their main purpose is to quantify meteorological phenomena such as precipitation and monitor storm systems, but weather radars also measure other things aloft in the lower atmosphere. As our abilities to analyze and store the huge amounts of data gathered by weather radar networks increase, new possible applications arise.
One application that has seen a steep rise in use is the extraction of biological targets from weather radar data, information that is immensely useful for a broad variety of fundamental and applied biological and ecological questions1,2. As weather radars detect all aerial fauna, from insects3 to birds4 and bats5, they can continuously monitor the movements of animals in the air. Such continuous biodiversity monitoring is increasingly recognized as essential for detecting biodiversity loss and testing the efficacy of management measures and policies6,7, yet difficult and costly to obtain with traditional observation methods1.
Probably the greatest asset of using weather radar data in biological studies is the large temporal and spatial scopes that can be investigated6. For the first time we can directly measure several large scale phenomena, such as the continent-wide migration of birds8,9. Billions of birds traverse the earth each year, with far-reaching consequences for conservation, ecosystem health, disease spread, aviation safety and infrastructure development6. One limitation of using weather radar data for biological monitoring is the low taxonomic resolution, as targets often cannot be identified beyond their class (for example “bird” or “insect”). However, citizen science and other field observations can complement radar data with species-specific information10. There are also some noteworthy examples where further identification is possible due to specific animal behaviors, such as emergence of Brazilian free-tailed bats (Tadarida brasiliensis) at Bracken cave5, roosting swallows in the US11, and twilight flights of Common swifts (Apus apus)12. The overall low taxonomic resolution means that applications have been largely macro-ecological13 in nature, focused on aspects such as reducing threats along migratory flyways14 or monitoring changes to ensembles of species9,15,16.
The benefits of monitoring large parts of migration flyways and different habitats crucially relies on combining data from a network of weather radars, ideally over long periods. This is comparatively straightforward in overall homogeneous radar networks and where data are managed by a single agency. In the US, for example, NOAA (National Oceanic and Atmospheric Administration) provides public access to archived Level II data (radar sweeps for base and dual-polarization quantities) that can be used to derive meteorological and biological products17. Open access to this archive has greatly facilitated the use of weather radar data in biology and led to the emergence of a vibrant field of aeroecology, the study of life in the air18,19. In contrast, the European network of weather radars is heterogeneous in hardware, data processing and data availability, with national meteorological services operating with varying policy priorities and under very different conditions, which has long hampered the development of a joint dataset of biological weather radar data from across Europe20. However, with careful consideration and quality control, pan-European comparisons can be made8,21,22.
Here we present two public datasets of animal movement data derived from European weather radars. They are the result of long-term interdisciplinary collaborations through the COST action ENRAM (European Network for the Radar surveillance of Animal Movement)23 and BiodivERsA projects GloBAM and HiRAD. The datasets were created by processing radar data with dedicated and validated methods for detecting birds and especially bird migration, resulting in summaries of movement characteristics (e.g. density, direction and speed) at regular altitude intervals across the lower atmosphere. Despite heterogeneity in coverage and data quality, they provide the largest and most complete datasets of their kind in Europe. By creating, standardizing and publicly archiving these data, we reduce computing and access difficulties to facilitate aeroecology research and applications. The datasets are deposited on Zenodo and can also be explored in the Aloft data portal (see Aloft data portal).
Methods
The data were created by processing polar volume data (PVOL) to vertical profiles of biological targets (VP). These are then combined into vertical profile time series (VPTS).
PVOL data access
PVOL data result from the scanning routine of a radar (also known as volume coverage pattern or VCP). It consists of a number of scans (also called sweeps) collected at different elevation angles (angle between radar scan and horizon), with quantities observed by the radar such as reflectivity factor (DBZ) and radial velocity (VRAD) (see for example24 for details). Scanning routines are typically conducted every 5 to 15 minutes and generally result in one PVOL file per routine.
To exchange such data, national meteorological services in Europe collaborate in OPERA, the Operational Programme for the Exchange of Weather Radar Information25, under the umbrella of the European Meteorological Services Network, EIG EUMETNET. OPERA member countries submit PVOL data (or scans) to OPERA in real-time, using the ODIM HDF5 format26. Data are sometimes cleaned (i.e. filtered) for meteorological purposes prior to distribution (see Usage Notes and20). These data are subsequently used in OPERA production centers to produce meteorological products. They are also forwarded to the OPERA development environment (ODE) server, where software called the BALTRAD Toolbox27 does additional processing and archiving. This server is maintained by the Swedish Meteorological and Hydrological Institute (SMHI) and further referred to as “BALTRAD”.
European PVOL data are generally not publicly available. Access is granted to members of the ENRAM consortium (including authors of this paper) by meteorological institutes of many OPERA member countries (i.e. the 18 countries covered by the data, as well as Austria and the United Kingdom) through a license agreement. It states that PVOL data cannot be distributed further and can only be used “for the purpose of extracting animal migration information for scientific research”. These restrictions do not apply to derived VP/VPTS data. For the BALTRAD_VPTS28 dataset, we obtained PVOL data from BALTRAD. For the UVA_VPTS29 dataset, the University of Amsterdam (UvA) obtained PVOL data as part of several research projects related to aviation safety and wind energy, from three national meteorological services: the Royal Meteorological Institute of Belgium (RMI), German Meteorological Service (DWD) and Royal Netherlands Meteorological Institute (KNMI). In contrast with (recent) data provided to BALTRAD, these data are unfiltered and often contain dual polarization quantities, which results in higher quality VP data (see Usage Notes).
PVOL to VP data processing
We created VP data by processing PVOL data with the vol2bird algorithm30,31, using the default settings. A VP consists of estimated density, speed and direction of biological targets such as birds, bats and insects within a weather radar volume per altitude layer. The vol2bird algorithm uses a combination of local variation in radial velocity (VRADH) and reflectivity (DBZH) thresholds to extract animal (mainly bird) signals. Each PVOL file results in a corresponding VP file, a HDF5 file expressed in the ODIM bird profile format.
For BALTRAD_VPTS, data from 2012 to 2017 were obtained from the BALTRAD archive and processed to VP data once, while data from 2018 onward are created by an operational service at BALTRAD (see Fig. 1). For UVA_VPTS, data were processed once. All processing was done with vol2bird v0.3 or v0.5. The vol2bird version and used parameters are recorded in the VP metadata.
Schematic diagram of the processing steps used to create the datasets, showing the main data storage locations (boxes), processes (arrows) and access methods (screenshots). A number of operational services (purple) process incoming PVOL to VP and VP to VPTS on a daily basis.
VP to VPTS data processing
We then extracted the tabular data and relevant metadata from the VP files (55 772 262 files), combined these into time series and converted those to the VPTS CSV (v1.0) format32. This format defines all measurements, facilitating standardized data exchange. While this process removes some metadata, it does not reduce the amount of data records, nor does it statistically aggregate data. BALTRAD_VPTS data from 2018 to 2023 were converted using the Python package vptstools33 as part of an operational service. All other data were converted using the R package bioRad34,35 (see Fig. 1).
We first created daily CSV files per radar (298 498 files) and then concatenated and gzipped these into monthly csv.gz files (10 991 files). The VPTS files are more storage efficient than the VP files, allowing faster downloads and analyses. For the same time period, daily files were on average 25% of the original VP file size, monthly files 6%.
Long-term archiving
Finally, we performed a data format validation on all VPTS data using the Python library frictionless-py36 (see Technical Validation) and archived the monthly-packaged data on Zenodo, a free-to-use, general-purpose research repository. One deposit was created for each source (BALTRAD and UvA).
Operational services
At the time of writing, data derived from BALTRAD are updated on a daily basis. Since 2015, SMHI has maintained a service that processes incoming PVOL to VP data and provides access to these via a restricted FTP server, where data older than two days are deleted. To facilitate access and prevent data loss, the Research Institute for Nature and Forest (INBO) maintains a service since 2017 that downloads the VP data from the FTP server to a publicly accessible Amazon S3 cloud object storage (further referred to as “bucket”). Since 2023, an additional service generates daily and monthly VPTS data from VP data (see Fig. 1). The bucket is gradually growing in size, providing public access within 48 hours after raw data were collected. Although the bucket and services are offered for free, their maintenance, computing and hosting is not. Those costs have been covered by a number of funding sources, but, unfortunately, do not guarantee long-term availability.
Data Records
A static version of the data was deposited on 2025-01-21 on Zenodo as two datasets:
-
BALTRAD_VPTS - Vertical profiles of biological targets derived from European weather radars: https://doi.org/10.5281/zenodo.1471102428
-
UVA_VPTS - Vertical profiles of biological targets derived from weather radars in Belgium, Germany and the Netherlands: https://doi.org/10.5281/zenodo.1471124429
For other data formats and daily updates, see Aloft data portal.
File properties, organization, naming and format
Both datasets contain vertical profile time series (VPTS). A VPTS expresses the density, speed and direction of biological signals (here predominantly birds) within a weather radar volume, grouped into altitude bins (here 200 m resolution/binsize) and measured over time (typically every 5 or 15 min). It can be expressed as tabular data containing three grouping properties (radar, datetime, height), calculated measurements (18 columns), radar metadata as found in the source VP file (4 columns) and the name of that source file (source_file). See Table 1 for a description of all properties.
The files are organized as follows:
-
country: a <country>.tgz file containing directories.
-
radar: a <radar> directory.
-
year: a <yyyy> directory.
-
month: a <radar>_vpts_<yyyy><mm>.csv.gz file.
Where:
-
<country>: 2-letter ISO 3166-1 alpha-2 code.
-
<radar>: 5-letter ODIM NOD radar identifier, where the two first letters signify the country (ISO 3166-1 alpha-2 code) and the last three the radar station (e.g. SEANG for the Swedish radar at Ängelholm). See also Radar metadata.
-
<yyyy>: year.
-
<mm>: month (with leading zero).
Each monthly file uses the VPTS CSV v1.0 format (see Methods). The Table Schema for this format (vpts-csv-table-schema.json) is deposited with the data as technical documentation. All timestamps are in UTC.
License
The data are released under a Creative Commons Zero waiver.
Geographic coverage
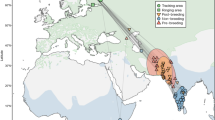
BALTRAD_VPTS28 contains data from 151 radars at 140 locations in 18 countries, ranging from the Spanish radar ESLPA on Gran Canaria in the south (latitude 28.0) to NOHAS in arctic Norway in the north (latitude 70.6) and longitudinally, from -15.6 (ESLPA) to 34.8 (ILTLV) in Israel. UVA_VPTS29 contains data from 24 radars at 24 locations in three countries, with only BEZAV not included in BALTRAD_VPTS. Collectively, the datasets cover 152 radar stations at 141 locations (see Fig. 2).
Maps of radar locations showing how many days with data are available (color) and the area used to extract biological data (size of circle, 25 km radius shown). Radars that changed code, but not location, are combined. (A) The 140 radar locations included in BALTRAD_VPTS. (B) The 24 radar locations included in UVA_VPTS.
Note that the datasets do not include all radar stations in Europe, but only those that are 1) included in the OPERA network, 2) where the managing meteorological service signed the license agreement with ENRAM and sends data to OPERA, and 3) where data contain the necessary properties to be processed to VP/VPTS data. Even if a radar station has data associated with it, not all locations can be reliably compared due to heterogeneity in data acquisition, cleaning and collation procedures before the biological data are extracted20. In recent years, many radars were upgraded from single polarization to dual polarization37. In some cases, this opportunity was used to move the radar to a more suitable location in close proximity to the original radar. For 11 radars in Germany and Sweden the radar code was also changed (see Table 2), resulting in a location being associated with two codes.
For each radar, data collected between 5 and 25 km from the radar was used to calculate VP data. This was updated to 5 and 35 km on 2019-09-23 for BALTRAD_VPTS and 2020-01-02 for UVA_VPTS to improve vertical profile coverage at higher altitudes while not compromising altitudinal resolution. The area used for biological monitoring is thus much smaller than the maximum range of the radar, which extends well beyond 200 km for most radars. Although biological signals can be detected further from the radar, the curvature of the earth and the elevation angle of the lowest beams cause the radar to “overshoot” low-flying fauna at distances further than 35 km (see e.g.38). The beam width also rapidly increases with range, leading to an effective loss in altitudinal resolution. The area closest to the radar (<5 km) is excluded as it does not give reliable data due to radar sidelobes and strong reflections from objects on the ground39 (ground clutter) obscuring bird movements.
Temporal coverage
The datasets span 16 years, from 2008 to 2023 (2012 to 2023 for BALTRAD_VPTS28), covering 4 877 out of 5 844 possible days (83%). Depending on the scanning strategy, data are generally collected at 5 or 15 min intervals, resulting in 1 379 816 unique timestamps (floored to 5 min intervals). The number of days with data at each radar station varies greatly, spanning from almost full coverage at 4 684 days (NLDHL) to only 23 days (ESLPA). See the Supplementary Information for a visual overview of the number of days with data per source, radar and year.
Gaps in the temporal coverage of data occur due to a variety of reasons. Power outages or radars being offline for maintenance can lead to missing data at single locations40. BALTRAD or other processing service issues can lead to missing data across locations, also for extended periods of time. For most years in UVA_VPTS only data during migration seasons (February-May, August-November) are retained.
Taxonomic coverage
Except for very specific circumstances (see Background & Summary), biological signals in weather radar data typically offer limited opportunities for species identification. The vol2bird algorithm uses different aspects of the return echos and the movement of objects in relation to the radar and to prevailing wind to extract biological signals30. However, some ambiguity will remain and background knowledge of the species composition and movements at different locations will be required to correctly interpret the data. Comparative analysis across different monitoring systems have been used to validate the method in for example4,30,41,42. The vol2bird algorithm is optimized for broad-scale nocturnal migration of small birds, and out of the box will work best in these situations30.
During the vol2bird processing a threshold is applied to the standard deviation of the residuals in the volume velocity profiling procedure, a method used to determine speed and direction of targets34. This threshold (sd_vvp_threshold) is used to separate self-propelled animals flying at heterogeneous airspeeds relative to the radar from rain and insects moving more uniformly with the wind. This threshold creates a classification trade-off: when set too low, rain may be included as animals; when set too high, animals may be excluded when movement directions and speeds are uniform. The default threshold (sd_vvp_threshold) in bioRad is set at 2 m/s for C band30 and 1 m/s for S band and can be adjusted after processing. By default, animal densities are converted from radar reflectivity using a radar cross section (rcs) of 11 cm2, which represents the average size of a nocturnal bird migrant in Europe, estimated in a field campaign in the Netherlands, Belgium and France that collocated a bird radar with several weather radars30. The rcs can be adjusted ad hoc, also after processing, using the bioRad function rcs(). Gaussian mixture models fit to VPTS can further refine the distinction between birds and insects22, especially in cases where increasing the sd_vvp_threshold would interfere with the bird signal extraction (see Gaps caused by uniform movements).
Technical Validation
We validated the format of the datasets28,29, by comparing the data with the VPTS CSV v1.0 Table Schema (deposited as vpts-csv-table-schema.json with the data), which defines the expected columns, required values, data types and constraints. Except for 26 cases of dbz_all values above the expected 100 (in BALTRAD_VPTS’s demem_vpts_202103.csv.gz), all 1 353 818 545 rows of data were technically valid and should be readable without issue. Note that duplicate radar, timestamp, height combinations may occur: these are the result of duplicate source files or rounded timestamps in the PVOL data.
The European weather radar network is a heterogeneous network, with national meteorological services using radar hardware, signal processing software, scanning strategies, data cleaning and data storage protocols that vary across Europe. Data quality and interoperability therefore varies, mainly between countries but in some cases also between locations, and over time. Such differences can influence the VPTS data and care must be taken when comparing different countries, locations and time periods. We have performed several data quality assessments on the PVOL and VP data (mainly for BALTRAD_VPTS) and summarized the results by country in https://github.com/aloftdata/data-repository/wiki. Our manual quality and consistency checks do not cover the entire dataset. We therefore encourage users to evaluate qualitative and quantitative requirements for their use case and welcome them to report quality issues in the wiki. Some of the regularly encountered issues are listed below.
Rain contamination
Even though the vol2bird algorithm can separate biological and meteorological targets with high accuracy30, some rain contamination may persist in the biological data. Rain has a very high reflectivity, resulting in a disproportionately large effect on the derived biological density when miss-classified as biological targets. This will often occur either directly before or after precipitation events, and may be caused by miss-classifying a single rain-filled range gate (a small volume of airspace measured by the radar) as birds (see Fig. 3). Rain contamination is usually easy to visually identify in VPTS plots, as high total reflectivity (dbz_all, also referred to as DBZH) extending from the ground to several km above ground or as isolated altitude bins with very high biological densities (dens). Rain contamination can also be spotted by plotting and comparing animal density (dens) and total reflectivity (dbz_all) (see Fig. 3) and events may be excluded based on characteristic altitudinal patterns of dbz_all (see e.g.8,15).
(A–C): Example of residual rain contamination in the biological density data, derived from radar SEANG in Ängelholm, Sweden, in August 2020. (A) Total reflectivity factor (dbz_all) over height. Profiles appear truncated above 1600 m. A rain event (black) is seen starting before midnight. (B) Reflectivity attributed to biological targets over the same period. Nocturnal migration is seen before the rain event, with a typical gradual increase and decrease. Later in the night there is a clear sign of rain contamination (between dotted red lines), with two altitude bins temporarily showing very high densities. (C) Vertical integration of the above biological density. The large peak in vertically integrated density is due to the high density of the rain contamination event. (D–F): Example of gaps due to a high radial velocity standard deviation threshold (sd_vvp_threshold), derived from radar SEHEM in Hemse, Sweden, in August 2021. (D) Total reflectivity factor (dbz_all) over a period of intense migration. (E) Animal volume density using the default sd_vvp_threshold of 2. This high threshold filters out animals moving uniformly in the same direction, resulting in gaps (white). (F) Animal volume density using a lower sd_vvp_threshold of 0.5, retaining more of the migration dynamics. Lowering the threshold retains more nocturnal biological scattering (likely a mixture of birds and insects), but also retains substantial diurnal reflectivity, which are likely due to insects, especially in this early migration season example when insects are still relatively active.
Gaps caused by uniform movements
Especially during very intense and highly uniform movements of migratory birds, and during bird migration events with strong insect contamination, detected bird speeds and directions will become more spatially homogeneous and radial velocity standard deviation (sd_vvp, see30,34) might approach that of wind-borne insect movements. A default sd_vvp_threshold (2 m/s) will filter those out, resulting in characteristic “holes” when plotting the VPTS data (see Fig. 3E,F, for details see Taxonomic coverage). The sd_vvp_threshold can be adjusted in bioRad with the function sd_vvp_threshold(). Nussbaumer et al.22 specifically addressed the gaps by omitting the sd_vvp_threshold on rain-free VPs and instead applied Gaussian mixture models to correct for the presence of insects.
Truncated vertical profiles
VPs may appear truncated, with biological intensity artificially cut off at higher altitudes above the radar, resulting in vertically integrated densities being underestimated (see Fig. 3). This issue was noted especially in several radars in Poland and France. Truncation at higher altitudes appears when the PVOL data only includes relatively low elevation scans, and thus the beam does not reach higher altitudes within the range used to extract VP data. If no additional elevation scans are available, extending the maximum range of PVOL processing can increase the covered altitudinal range. This approach was for example used in Nilsson et al.8.
Filtering of biological signals prior to data exchange
In recent years, several countries have used dual-polarization to clean PVOL data of non-meteorological signals prior to data exchange with OPERA, leading to largely empty VPTS data even when biological activity is present20. Cleaning policies differ between national meteorological services and may result in systematic differences in the magnitudes of biological signals. For example, most Spanish radar stations are located along a major migratory flyway but show almost no bird movement, and Estonian data shows relatively low bird densities compared to its neighboring country Finland.
Noise from moving (man-made) objects
Reflections caused by truly static objects in the landscape, such as buildings and mountains, can be reliably removed using Doppler filtering by radar signal processors based on their velocity being close to 0 m/s. However, moving objects, such as waves or spinning wind turbines, are difficult to filter out using the same approach and might show a radar signature similar to animals, causing them to be classified as such. Depending on the nature of these objects and their movements, this might result in occasional spikes or higher background noise in the animal densities. With rapid expansion of wind energy production across Europe the amount of noise generated by wind parks is likely to increase over time, requiring careful consideration of its effects on temporal patterns derived from VPTS data for radar stations located close to wind parks.
Usage Notes
Aloft data portal
The easiest way to explore and visualize the data is through the Aloft data portal (https://aloftdata.eu). It provides access to the bucket that currently contains both datasets and daily updates of the BALTRAD data (see Operational services). The bucket organizes data in hierarchical directories: source (e.g. baltrad, uva), format (three formats: hdf5 for VP, daily/monthly for VPTS), radar, year, month, and day. Files can be downloaded in the browser or by using a software tool that supports S3, such as AWS CLI or rclone, which allows users to maintain a synchronized local copy of the bucket’s (sub)directories. BALTRAD data can be visualized directly in the browser at https://crow.aloftdata.eu43. By selecting a date (or three-day interval) and radar, it visualizes both bird density at different heights and migration traffic rate (MTR).
Recommended software
Data on Zenodo can be downloaded as one .tgz file per country, which contains the VPTS data as .csv.gz files (see Data Records). Once decompressed, data can be read with any CSV reader. For expected data types, see the associated vpts-csv-table-schema.json file, which can be read automatically with software supporting the Data Package standard.
For data analysis, we recommend using the R package bioRad34, which can directly read (compressed) VPTS data with read_vpts() and provides functionality to filter, time-regularize and plot data (filter_vpts(), regularize_vpts() and plot(), respectively). It can also vertically integrate data with integrate_profile() and calculate migration traffic rate (MTR) and vertically integrated density (VID) (see34 for an overview of these properties).
Radar metadata
A list of radars can be found at https://aloftdata.eu/radars/, which is derived from a database maintained by OPERA and indicates radar code (ODIM identifier), location, coordinates, characteristics and status. This list can change over time as new radars are installed and old radars decommissioned (indicated with status = 0). Codes are issued by OPERA member countries for new radars and sometimes replace those of existing radars after major hardware upgrades (see Geographic coverage, Table 2).
Use cases
The data presented here have the potential to be used in a wide range of applied and fundamental ecological studies. When the heterogeneity in data is appropriately addressed (see Technical Validation) and the limited taxonomic resolution recognized (see Taxonomic coverage) these data can be used to map the mass movements of animals in the air across Europe8,21,22. Applications include investigating anthropogenic disturbances44,45, disease spread46, aviation safety47,48, fundamental characteristics of animal migration15,49 and reactions to extreme events50,51. For more details on possible applications, see for example2,6.
Future developments
OPERA is currently changing how radar data are archived at the European level, which will likely lead to changes in the data flow to BALTRAD and the bucket. Should these changes by OPERA also include that only heavily cleaned data are archived centrally, it would become impossible to extract biological signals20 across the European network without working with each national meteorological service individually, and a unique tool for large scale biodiversity monitoring will be lost. However, with a willingness to accommodate data use beyond meteorology, the current adjustments to the OPERA processing pipeline could lead to more homogeneous products with better access to uncleaned data, which would greatly benefit biological monitoring, applications such as aviation safety or wind farm operations and society as a whole. The value of long-term monitoring datasets in biology cannot be stressed enough. It is our hope that the increased spread and use of the data described here will encourage more policy makers to appreciate the value of reusing already collected radar data, which is only made possible by archiving and sharing uncleaned radar products. See Shamoun-Baranes et al.1,20 for more details.
References
Shamoun-Baranes, J. et al. Weather radars’ role in biodiversity monitoring. Science 372, 248, https://doi.org/10.1126/science.abi4680 (2021).
Nilsson, C., Dokter, A., Bauer, S., Horton, K. & Farnsworth, A. Advances in weather radar monitoring of bird movement. In Bringi, V., Mishra, K. V. & Thurai, M. (eds.) Advances in Weather Radar. Volume 3: Emerging applications, 241–280, https://doi.org/10.1049/sbra557h_ch7 (Institution of Engineering and Technology, 2023).
Hu, G. et al. Mass seasonal bioflows of high-flying insect migrants. Science 354, 1584–1587, https://doi.org/10.1126/science.aah4379 (2016).
Nilsson, C. et al. Field validation of radar systems for monitoring bird migration. Journal of Applied Ecology 55, 2552–2564, https://doi.org/10.1111/1365-2664.13174 (2018).
Stepanian, P. M. & Wainwright, C. E. Ongoing changes in migration phenology and winter residency at Bracken Bat Cave. Global Change Biology https://doi.org/10.1111/gcb.14051 (2018).
Bauer, S. et al. The grand challenges of migration ecology that radar aeroecology can help answer. Ecography 42, 861–875, https://doi.org/10.1111/ecog.04083 (2019).
Moersberger, H. et al. Europa Biodiversity Observation Network: User and Policy Needs Assessment. ARPHA Preprints 3, e84517, https://doi.org/10.3897/arphapreprints.e84517 (2022).
Nilsson, C. et al. Revealing patterns of nocturnal migration using the European weather radar network. Ecography 42, 876–886, https://doi.org/10.1111/ecog.04003 (2019).
Horton, K. G. et al. Phenology of nocturnal avian migration has shifted at the continental scale. Nature Climate Change 10, 63–68, https://doi.org/10.1038/s41558-019-0648-9 (2020).
Weisshaupt, N., Lehtiniemi, T. & Koistinen, J. Combining citizen science and weather radar data to study large-scale bird movements. Ibis 163, 728–736 https://doi.org/10.1111/ibi.12906 (2021).
Kelly, J. F. & Pletschet, S. M. Accuracy of swallow roost locations assigned using weather surveillance radar. Remote Sensing in Ecology and Conservation 4, 166–172, https://doi.org/10.1002/rse2.66 (2018).
Dokter, A. M. et al. Twilight ascents by common swifts, Apus apus, at dawn and dusk: Acquisition of orientation cues? Animal Behaviour 85, 545–552, https://doi.org/10.1016/j.anbehav.2012.12.006 (2013).
Kelly, J. F. & Horton, K. G. Toward a predictive macrosystems framework for migration ecology. Global Ecology and Biogeography 25, 1159–1165, https://doi.org/10.1111/geb.12473 (2016).
Horton, K. G. et al. Bright lights in the big cities: migratory birds’ exposure to artificial light. Frontiers in Ecology and the Environment 17, 209–214, https://doi.org/10.1002/fee.2029 (2019).
Dokter, A. M. et al. Seasonal abundance and survival of North America’s migratory avifauna determined by weather radar. Nature Ecology & Evolution 2, 1603–1609, https://doi.org/10.1038/s41559-018-0666-4 (2018).
Rosenberg, K. V. et al. Decline of the North American avifauna. Science 366, 120–124, https://doi.org/10.1126/science.aaw1313 (2019).
Ansari, S. et al. Unlocking the potential of NEXRAD data through NOAA’s Big Data Partnership. Bulletin of the American Meteorological Society 99, 189–204, https://doi.org/10.1175/BAMS-D-16-0021.1 (2018).
Kunz, T. H. et al. Aeroecology: Probing and Modeling the Aerosphere. Integrative and Comparative Biology 48, 1–11, https://www.jstor.org/stable/25160115 (2008).
Shamoun-Baranes, J., Nilsson, C., Bauer, S. & Chapman, J. Taking radar aeroecology into the 21st century. Ecography 42, 847–851, https://doi.org/10.1111/ecog.04582 (2019).
Shamoun-Baranes, J. et al. Meteorological data policies needed to support biodiversity monitoring with weather radar. Bulletin of the American Meteorological Society 103, 1234–1242, https://doi.org/10.1175/bams-d-21-0196.1 (2022).
Kranstauber, B., Bauer, S. & Shamoun-Baranes, J. Geographic barriers and season shape the nightly timing of avian migration. Global Ecology and Biogeography 32, 1928–1936, https://doi.org/10.1111/geb.13742 (2023).
Nussbaumer, R. et al. Quantifying year-round nocturnal bird migration with a fluid dynamics model. Journal of the Royal Society Interface 18, 20210194, https://doi.org/10.1098/rsif.2021.0194 (2021).
Shamoun-Baranes, J. et al. Continental-scale radar monitoring of the aerial movements of animals. Movement Ecology 2, 9, https://doi.org/10.1186/2051-3933-2-9 (2014).
Stepanian, P. M., Chilson, P. B. & Kelly, J. F. An introduction to radar image processing in ecology. Methods in Ecology and Evolution 5, 730–738, https://doi.org/10.1111/2041-210X.12214 (2014).
Saltikoff, E. et al. OPERA the Radar Project. Atmosphere 10, 1–13, https://doi.org/10.3390/atmos10060320 (2019).
Michelson, D. et al. EUMETNET OPERA weather radar information model for implementation with the HDF5 file format Version 2.4, https://www.eumetnet.eu/wp-content/uploads/2021/07/ODIM_H5_v2.4.pdf. (2021).
Michelson, D. et al. BALTRAD Advanced Weather Radar Networking. Journal of Open Research Software 6, 12, https://doi.org/10.5334/jors.193 (2018).
Desmet, P. et al. BALTRAD_VPTS - Vertical profiles of biological targets derived from European weather radars. Zenodo, https://doi.org/10.5281/zenodo.14711024 (2025).
Shamoun-Baranes, J. et al. UVA_VPTS - Vertical profiles of biological targets derived from weather radars in Belgium, Germany and the Netherlands. Zenodo https://doi.org/10.5281/zenodo.14711244 (2025).
Dokter, A. M. et al. Bird migration flight altitudes studied by a network of operational weather radars. Journal of the Royal Society, Interface 8, 30–43, https://doi.org/10.1098/rsif.2010.0116 (2011).
Dokter, A. M. et al. vol2bird: Vertical profiling of biological scatterers from weather radar data. Zenodo, https://doi.org/10.5281/zenodo.3675621 (2020).
Desmet, P., Noé, N. & Dokter, A. M. VPTS CSV. Zenodo https://doi.org/10.5281/zenodo.13959554 (2023).
Van Hoey, S., Noé, N., Huybrechts, P. & Desmet, P. vptstools: Transfer and convert vertical profile time series data. Zenodohttps://doi.org/10.5281/zenodo.13959710 (2023).
Dokter, A. M. et al. bioRad: biological analysis and visualization of weather radar data. Ecography 42, 852–860, https://doi.org/10.1111/ecog.04028 (2019).
Dokter, A. M., Desmet, P., Kranstauber, B., Nilsson, C. & Van Hoey, S. bioRad: Biological Analysis and Visualization of Weather Radar Data. Zenodo https://doi.org/10.5281/zenodo.10027671 (2023).
Karev, E. et al. frictionless: Python library for Data Packages. Zenodo https://doi.org/10.5281/zenodo.8135154 (2023).
Stepanian, P. M., Horton, K. G., Melnikov, V. M., Zrnić, D. S. & Gauthreaux, S. A. Dual-polarization radar products for biological applications. Ecosphere 7, e01539, https://doi.org/10.1002/ecs2.1539 (2016).
Kranstauber, B. et al. High-resolution spatial distribution of bird movements estimated from aweather radar network. Remote Sensing 12, 635, https://doi.org/10.3390/rs12040635 (2020).
Holleman, I. Quality control and verification of weather radar wind profiles. Journal of Atmospheric and Oceanic Technology 22, 1541–1550, https://doi.org/10.1175/JTECH1781.1 (2005).
Saltikoff, E., Kurri, M., Leijnse, H., Barbosa, S. & Stiansen, K. Maintenance Keeps Radars Running. Bulletin of the American Meteorological Society 98, 1833–1840, https://doi.org/10.1175/BAMS-D-16-0095.1 (2017).
Weisshaupt, N., Hervo, M. & Haest, B. Comparison of bird migration in a radar wind profiler and a dedicated bird radar. Remote Sensing in Ecology and Conservation n/a, https://doi.org/10.1002/rse2.350 (2023).
Gasteren, H. V., Holleman, I., Bouten, W., Loon, E. V. & Shamoun-Baranes, J. Extracting bird migration information from C-band Doppler weather radars. Ibis 150, 674–686, https://doi.org/10.1111/j.1474-919X.2008.00832.x (2008).
Noé, N., Reyniers, M., Van Hoey, S. & Desmet, P. CROW - Online tool to visualize birds detected by weather radars. Zenodo https://doi.org/10.5281/zenodo.7372679 (2022).
Weisshaupt, N., Leskinen, M., Moisseev, D. N. & Koistinen, J. Remote Sensing 14, 1–13, https://doi.org/10.1002/rse2.350 (2022).
Cohen, E. B. et al. Using weather radar to help minimize wind energy impacts on nocturnally migrating birds. Conservation Letters 15, e12887, https://doi.org/10.1111/conl.12887 (2022).
McCuen, M. M. et al. A comparison of amplification methods to detect Avian Influenza viruses in California wetlands targeted via remote sensing of waterfowl. Transboundary and Emerging Diseases 68, 98–109, https://doi.org/10.1111/tbed.13612 (2021).
Kranstauber, B., Bouten, W., Van Gasteren, H. & Shamoun-Baranes, J. Ensemble predictions are essential for accurate bird migration forecasts for conservation and flight safety. Ecological Solutions and Evidence 3, e12158, https://doi.org/10.1002/2688-8319.12158 (2022).
van Gasteren, H. et al. Aeroecology meets aviation safety: early warning systems in Europe and the Middle East prevent collisions between birds and aircraft. Ecography 42, 899–911, https://doi.org/10.1111/ecog.04125 (2019).
Horton, K. G., Doren, B. M. V., Stepanian, P. M., Farnsworth, A. & Kelly, J. F. Where in the air? Aerial habitat use of nocturnally migrating birds. Biology letters 12, 20160591 https://doi.org/10.1098/rsbl.2016.0591 (2016).
Shamoun-Baranes, J. et al. Birds flee en mass from New Year’s Eve fireworks. Behavioral Ecology 22, 1173–1177, https://doi.org/10.1093/beheco/arr102 (2011).
Nilsson, C., Horton, K. G., Dokter, A. M., Van Doren, B. M. & Farnsworth, A. Aeroecology of a solar eclipse. Biology Letters 14, 20180485, https://doi.org/10.1098/rsbl.2018.0485 (2018).
Acknowledgements
The datasets are the result of long-term interdisciplinary collaborations that initiated in ENRAM and continued through GloBAM and other projects. We would like to thank the many people and organizations that were involved. Special thanks go those involved in all aspects of the operation of the European weather radars, in particular EUMETNET and OPERA. Thanks to Maarten Reyniers (RMI) and Klaus Stephan (DWD) for providing data to UvA, Anders Henja for support in implementing vol2bird in BALTRAD and Io Deflem for support in using the Flemish Supercomputer Center (VSC). Data were processed on infrastructure provided by SMHI, INBO, UvA, Dutch national e-infrastructure, SURF Cooperative, Ghent University, and VSC. Financial support was provided by the European Cooperation in Science and Technology (COST) for action ES1305 “ENRAM”, the 2017-2018 Belmont Forum and BiodivERsA joint call for research proposals, under the BiodivScen ERA-Net COFUND programme, for “GloBAM” (funded by the Swiss National Science Foundation (SNF 31BD30_184120), Belgian Federal Science Policy Office (BelSPO BR/185/A1/GloBAM-BE), Netherlands Organisation for Scientific Research (NWO E10008), Academy of Finland (aka 326315) and National Science Foundation (NSF 1927743)), the European Union’s Horizon 2020 research and innovation programme for “EuropaBON” (101003553), the European Union’s Horizon 2020 Marie Skłodowska-Curie grant (844360), the Swedish Research Council for Sustainable Development (Formas, 2021-01622), the Belgian Science Policy Office for the valorisation project “CROW”, the 2022-2023 Biodiversa+ BiodivMon call for research proposals for “HiRAD” (funded by the Swiss National Science Foundation (SNF 31BD30_216840), Belgian Federal Science Policy Office (BelSPO RT/24/HiRAD), Netherlands Organisation for Scientific Research (NWO EP.1512.22.003) and Academy of Finland (aka 359864)), the Research Foundation - Flanders (FWO) as part of the Belgian contribution to LifeWatch, the European Space Agency Integrated Applications Program for “Flysafe”, the Netherlands Ministry of Defense for “Birds Avoidance System”, the Netherlands Organisation for Scientific Research (LifeWatch & 2021.030), the Netherlands Organisation for Scientific Research, in collaboration with Rijkswaterstaat and Gemini (17083), the Province of Groningen (IBED22026), and the Ministry of Agriculture, Fisheries, Food Security and Nature (IBEDO-LNV20220428).
Author information
Authors and Affiliations
Contributions
P.D., B.S., J.S., A.D., S.B., S.V.H. and C.N. conceived the initial idea for the datasets (conceptualization); P.D., J.S., A.D., N.W., S.B. and C.N. acquired funding to support these (funding-acquisition); P.D. supervised and coordinated their processing (supervision); P.D., A.D., G.H., B.K., N.N., S.V.H., B.W., H.L., B.H., P.H. and C.N. developed software to process, visualize or explore the data (software); P.D., G.H., and S.V.H implemented the operational data flow (data-curation); P.D., A.D., B.K., B.S., N.W., B.H., P.H. and C.N. reviewed data for issues (validation); P.D. and C.N. created the figures (visualization), C.N. and P.D. wrote the initial draft of the manuscript (writing-original-draft). All authors contributed critically to the drafts and gave final approval for publication (writing-review-editing).
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Desmet, P., Shamoun-Baranes, J., Kranstauber, B. et al. Biological data derived from European weather radars. Sci Data 12, 361 (2025). https://doi.org/10.1038/s41597-025-04641-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-04641-5