Abstract
There is currently a paucity of neuroimaging data from the African continent, limiting the diversity of data from a significant proportion of the global population. This in turn diminishes global health research and innovation. To address this issue, we present and describe the first Magnetic Resonance Imaging (MRI) dataset from individuals in the African nation of Nigeria. This dataset contains pseudonymized structural MRI (T1w, T2w, FLAIR) data of clinical quality, with 35 images from healthy control subjects, 31 images from individuals diagnosed with age-related dementia, and 22 from individuals with Parkinson’s Disease. Given the potential for Africa to contribute to the global neuroscience community, this unique MRI dataset represents both an opportunity and benchmark for future studies to share data from the African continent.
Similar content being viewed by others
Background & Summary
Human populations in Africa have among the greatest genetic diversity in the world. Mental and brain health in Africa are also among the most diverse in the world, because of the variety of stress factors, such as child health, the high level of incidence of traumatic brain injury, and the diseases endemic to the region1. Concurrently, Africa is experiencing a demographic shift with rising life expectancy, resulting in an increasing population of individuals aged 60 or above. For example, in Nigeria, approximately 7.76 million citizens out of a total population of 230 million (3.36%) are above 65 years old, and this number is steadily increasing2,3,4. This trend is accompanied by an increased prevalence in age-related diseases2, such as Parkinson’s Disease (PD)5 and Dementia6. As we stand, Africa has great potential for the neuroscience community, by offering both untapped talent, and contributing diversity to global data—unfortunately, the lack of infrastructure limits Africa’s realization of its potential1. Opportunities to increase Africa’s neuroscientific output and neuroimaging data originating from the continent are currently limited due to the lack of such data.
Parkinson’s disease (PD) is a gradually progressive neurodegenerative movement disorder resulting from selective loss of nigral dopaminergic neurons of unknown etiology and is clinically characterized by both motor and non-motor manifestations5. Age-related dementia, on the other hand, is a syndrome characterized by deterioration in cognitive function beyond what might be expected from the usual consequences of biological aging. It is caused by damage to or loss of nerve cells and their connections in the brain regions associated with memory and learning6. Parkinson’s Disease (PD) and Dementia are most prevalent among people 60 and older and are consequently among the leading causes of disability and dependency among older people globally2. PD has a prevalence rate of 6.0% to 8.3% of neurologic hospital admissions/consultations in West Africa, with estimated crude prevalence varying from 15 to 572 per 100,000 people7,8. Dementia, on the other hand, has a prevalence rate of 2.3% to 20.0%, and the incidence rates are 13.3 per 1000 people with increased mortality in parts of rapidly developing Africa6. Despite the rising prevalence of PD and Dementia in Africa, there is relatively little information on them in the global brain data repositories5,9.
Diversity or heterogeneity of datasets is critical to an effective, functional, dependable, and trustworthy health ecosystem. However, datasets from the global south, particularly from Africa, are worryingly missing from the global neuroscience research and innovation space. Furthermore, the availability of Findable, Accessible, Interoperable, and Reusable (FAIR) data10 is a critical factor that drives global health research and innovation. Despite comprising 12.5% of the world’s population, Africa still accounts for less than 1 percent of global research output11. Adequate representation of diverse datasets that reflect global demographics in scientific research is crucial for inclusion and equitable global health research. Our research aims to make FAIR African brain data available using Nigerian brain MRI datasets of patients with Parkinson’s Disease and Dementia as use cases.
To begin addressing these issues, we present and describe the first openly-shared Magnetic Resonance Imaging (MRI) dataset from individuals from the African nation of Nigeria. Study participants were recruited in three diagnostic centers across the South-South and North-Central geopolitical regions of Nigeria, spanning the following states: Delta, Edo, Bayelsa, Cross River, Akwa Ibom, and Rivers States (for the South-South) and Benue, Kogi, Kwara, Nasarawa, Niger, Plateau States and the Federal Capital Territory (for the North-Central). This dataset is unique because of the limited availability of shared data in similar ethnic groups. However, due to the reduced strength of the MRI scanners used in the procurement of the data (0.3 and 1.5 Tesla), the data are marked by diminished quality as they were collected following the best clinical practices in the regions and diagnostic centers without research purposes nor protocol standardization. The diversity in the ethnic group, the data quality, and protocol heterogeneity provide a first opportunity for training local clinicians, radiologists, and researchers. Furthermore, the dataset provides a unique opportunity for medical imaging and artificial intelligence systems interested in developing algorithms for disease detection that are robust to diminished data quality and heterogeneity.
This dataset is publicly available by utilizing the free and secure cloud computing platform brainlife.io12. The goal of sharing this dataset is primarily to demonstrate the potential of the untapped neuroimaging community in Nigeria. Secondly, the dataset can be used for the development and validity testing of FAIR tools13, including pseudo anonymization, preprocessing, and structural segmentation in clinical scanner environments. The data were converted from the DICOM imaging format to the Brain Imaging Data Structure (BIDS) using ezBIDS14, and then uploaded to b12. This approach integrates into a single record, the data and the preprocessing pipeline used, thereby enabling availability, interoperability, and reusability of the data15,16,17.
The processed dataset contains raw data and derivatives across 88 participants (Dementia: 31; Parkinson: 22; Control: 35; see Tables 1, 2 for data sources and demographics, respectively). The total size of the repository is over 15 GB of imaging data and derivatives, comprising T1w, T2w, and FLAIR anatomical sequences, corresponding brain masks, and summary measures18. The processing pipeline for this dataset utilizes mainstream neuroimaging software libraries including FSL19,20,21 and MRIQC18. The corresponding brainlife.io data processing Apps were developed with a lightweight specification and use modern methods for software containerization, making the analyses trackable, reproducible, and reusable on a wide range of computing resources22. The present descriptor describes the repository and pipelines published via brainlife.io mechanisms. These resources provide the broader neuroimaging community additional insights into the pathology of PD and Dementia by exploring high-quality preprocessed neuroimaging data, replicating previous examinations of the data, and examining a wide variety of hypotheses without the barriers above.
Methods
Ethics
Ethics approval was obtained by E.W. and P.F. from the Research Ethics Committee of the University Of Port-Harcourt, Nigeria with the reference code: UPH/CERMAD/REC/MM84/056.
Neuroimaging data sources
Data were collected from three Neuroimaging Diagnostic centers in Nigeria. Two centers (RUSTH lab, and the Intercontinental Lab) are located in the South-South city of Port Harcourt, and one center (the LifeBridge Diagnostic Lab) is in the Federal Capital Territory (FCT; North -Central), Abuja, specifically.
Study participants
Participants or their guardians signed informed consent forms for data collection for diagnostic and research purposes. Participants were not financially compensated for participation. A total of 761 MRI anatomical sequences were collected across 88 Nigerian participants, all of whom reside in Nigeria. Due to the sensitive nature of the data, and to protect the confidentiality and privacy of the subjects, data protection measures such as pseudonymization (defacing, manual brain masking), dedicated access control procedures, and Data Use Agreements (DUA), were developed and utilized. In total, the dataset is comprised of 31 participants with Dementia (age: M = 65.26 years, SD = 14.11, Range = [36–86]), 22 participants with PD (age: M = 61.14 years, SD = 14.60, Range = [36–79]), and 35 healthy control participants (age: M = 34.14 years, SD = 12.75, Range = [12–75]). Table 1 provides a breakdown of the demographics of the participant groups.
Below, we follow the COBIDAS reporting protocol23 to describe subjects’ information, exclusion, and inclusion criteria, and socioeconomic status (SES) information (Table 3). SES was measured using a combination of income, class, and education, consistent with best practices for SES indicators24. Specifically, we categorized the individuals into upper, middle, and lower class based on their economic standing, likely reflecting both income and occupation. Additionally, their highest level of education (e.g., completed tertiary, secondary, or informal education) was recorded. This method aligns with the multiple single indicators approach, where income/class and education are treated as separate, independent indicators of SES, rather than combining them into a composite index. By using this approach, we ensure that each aspect of SES—both economic and educational—is captured distinctly, providing a comprehensive understanding of the participants’ socioeconomic background.
Neuroimaging parameters
Participant data from the RSUTH lab were collected using a GE Signa 1.5 T (Software version SV25.1_R04_1743.a and SV25.1_R07_2127.a) scanner, whereas data from the LifeBridge Diagnostic Lab were collected using a Toshiba MRT200PP3 1.5 T scanner (Software version V9.51*R249), and data from the Intercontinental lab were collected using a Hitachi AIRIS II 0.3 T scanner (Software version V5.1H-1). A 12-channel head coil was used at all sites.
For a subset of participants (32 total participants), multiple runs of data collection were performed (i.e. run-1, run-2, run-3, run-4). For a larger subset (64 total participants), contrast-enhanced T1w images were collected, where the contrast agent used was gadolinium.
Due to the unique data acquisition parameters, not all participants have the same number of images. The type of data orientation, contrast enhancement, or run can be different across subjects. Specifically, 72 participants had T1w images each with separate high-resolution orientations (axial, coronal, sagittal), while some had less than all three high-resolution orientations. This was also the case for T2w images, where only 51 participants had three high-resolution orientations collected. For most participants where a FLAIR image was collected (80 participants total), only 32 participants had more than a single high-resolution orientation collected, but never all 3. Information regarding the breakdowns of the number of images collected across the various acquisition parameters is found in Table 4.
Anatomical data (T1w, T2w, FLAIR) preprocessing
The raw data contain anatomical (T1-weighted, T2-weighted, FLAIR) sequences. These DICOMs were converted to BIDS using ezBIDS. Upon conversion, the NIfTI images were visually inspected using FSL’s19,20,21 slicer functionality implemented in brainlife.app.300, brainlife.app.301, and brainlife.app.689. Following this, quality metrics of the T1w and T2w images were computed using MRIQC18 in brainlife.app.701 and brainlife.app.702.
Before publication, all images were masked to remove non-brain material to preserve participant anonymity. This was performed by generating brain masks using FSL’s Brain Extraction Tool (BET)25 implemented as brainlife Apps brainlife.app.2, brainlife.app.156, and brainlife.app.728. Masks were then manually refined using the FSLeyes26 image viewer in cases in which certain facial features were not removed automatically by the software. This failure of the automated procedure is likely due to the low data resolution. The goal of the manual procedure was to eliminate non-brain tissue without removing brain tissue. Following manual editing, the masks were applied to the anatomical data to extract brain data using FSL functionality implemented as brainlife.app.751, brainlife.app.752, and brainlife.app.753. Only defaced, masked images will be released to preserve participant anonymity.
Table 5 lists all of the brainlife.io Apps, with their associated GitHub repositories and branches, used to process data.
Data Records
Accessing the data
The data are available as a brainlife.io data publication: https://doi.org/10.25663/brainlife.pub.6127, masks of the brain were manually curated and can be found at https://doi.org/10.25663/brainlife.pub.5528. The following video shows how to access the data: https://youtu.be/fEeJwifagUQ.
Data files can either be used on brainlife.io or downloaded and organized using the BIDS standard25. Access to the published data is currently supported via (i) web interface and (ii) Command Line Interface (CLI).
T1-weighted Anatomical
T1w images were collected as 2D images, with one high-resolution and two low-resolution orientations. For some participants, multiple scans were collected in a single session and designated by a “run” tag, with the first run denoted as “run-1”, the second as “run-2”, the third as “run-3”, and the fourth as “run-4”. For the remainder who did not have multiple runs collected, no “run-” tag is specified. Some data were collected using a gadolinium contrast agent, these images are denoted by the “ce-gadolinium” tag.
upload/sub-{}/anat/ sub-{}_acq-[coronal,sagittal,axial]_tag-brainextracted_tag_desc-{}_T1w.json sub-{}_acq-[coronal,sagittal,axial]_tag-brainextracted_tag_desc-{}_T1w.nii.gz
T2-weighted Anatomical
T2w images were collected as 2D images with one high-resolution and two low-resolution orientations. For some participants, multiple runs were also collected, and were designated by a “run-” tag, with the first run being “run-1”, the second as “run-2”, the third as “run-3”, and the fourth as “run-4”. For the others who did not have multiple runs collected, no “run-” tag is specified.
upload/sub-{}/anat/ sub-{}_acq-[coronal,sagittal,axial]_tag-brainextracted_tag_desc-{}_T2w.json sub-{}_acq-[coronal,sagittal,axial]_tag-brainextracted_tag_desc-{}_T2w.nii.gz
Fluid attenuated inversion recovery (FLAIR)
FLAIR images were collected as 2D images with one high-resolution and two low-resolution orientations. For some participants, multiple runs were collected, and were designated by a “run” tag, with the first run as “run-1”, the second as “run-2”, and the third as “run-3”. For the others who did not have multiple runs collected, no “run-” tag is specified.
upload/sub-{}/anat/ sub-{}_acq-[coronal,sagittal,axial]_tag-brainextracted_tag_desc-{}_FLAIR.json sub-{}_acq-[coronal,sagittal,axial]_tag-brainextracted_tag_desc-{}_FLAIR.nii.gz
Brain masks
Brainmasks were generated for each individually collected image. First-pass brain masks were generated using FSL’s Brain Extraction Tool (BET) functionality and then downloaded locally to be manually refined. Upon refinement, data were then re-uploaded to brainlife.io before being applied to the anatomical images.
upload/sub-{}/mask/ sub-{}_acq-[coronal,sagittal,axial]_tag-anat_tag-brain_tag-[t1,t2,flair]_tag_desc-{}_mask.nii.gz
MRIQC
MRIQC was used to compute quality assurance metrics of the raw T1w and T2w images. With this comes a data structure that loosely follows the BIDS standard.
regressors/ regressors.tsv
Technical Validation
Anatomical (T1w, T2w, FLAIR) data
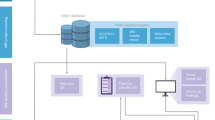
In this section, we provide a qualitative and quantitative evaluation of the data derivatives made available on brainlife.io. Figure 1 describes the workflow used to process and publish the data. Raw DICOM files from the MRI scanners were first converted to BIDS standard formats using ezBIDS and uploaded directly to brainlife.io.
Workflow for releasing the Nigerian Brain Dataset. Diagram describing the workflow performed for getting the Nigerian Brain Dataset ready for publication. Raw DICOMs from the MRI scanners were first converted into BIDS standard format using ezBIDS, and the converted data was uploaded and organized automatically to brainlife.io. Data was then assessed for quality both qualitatively using brainlife.io Apps to generate images of the raw data, and quantitatively using MRIQC. Before release, the data was pseudo-anonymized via defacing by creating brain masks and removing all non-brain material. Defaced data was then published with a digital object identifier (DOI) which can be interacted with via brainlife.io.
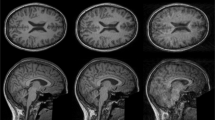
Upon conversion of the data from DICOMS to BIDS, data were masked and visually inspected for quality. Figure 2 exemplifies the quality of the raw anatomical [T1w & T2w (a), FLAIR (b)] images obtained with brainlife.app.300, brainlife.app.301, and brainlife.app.689 in representative participants from each diagnosis group [i.e. Dementia (sagittal T1w & T2w), Parkinson’s (coronal T1w & T2w), Control (axial T1w & T2w)]. Displayed images are the highest resolution orientation of the collected scan for each modality. Note, that for FLAIR images, no high-resolution sagittal orientation was collected.
Images of raw anatomical (T1w, T2w, FLAIR) data. (a) Axial (left), coronal (middle), and sagittal (right) high-resolution orientations for the T1w and T2w anatomical data. The displayed axial image is from a Control participant, the coronal image is from a Parkinson’s patient, and the sagittal image is from a Dementia participant. (b) Axial (top) and coronal (bottom) high-resolution orientations for the FLAIR images collected from a Control participant.
There exists a wide range of quality of the collected data. To assess this, following visual quality assurance, MRIQC18 was performed on the T1w and T2w images for each high-resolution orientation collected. Figure 3 reports the contrast-to-noise ratio (CNR) calculated by MRIQC for each participant, modality, and image orientation.
Range of quality of data (MRIQC data). (a) Contrast-to-noise ratio across all of the T1w and T2w data for each high-resolution orientation (axial: blue, sagittal: orange, coronal: green). Error bars represent ±1 standard deviation.
Usage Notes
Data access conditions/requirements
Access to the dataset is granted on a ‘controlled access’ basis. Users will be fully identified and authenticated. Access conditions are necessary to protect the rights of the participants due to the nature of the dataset (i.e. personal, sensitive health data). The dataset can be classified as special category data and as such requires technical and organizational measures as well as safeguards to ensure the confidentiality and privacy of the participants29,30. Rather than limit access to the dataset, such measures are designed to enable responsible research and innovation (RRI). To achieve this, data protection measures such as pseudonymization, access control, and Data Use Agreement (DUA) have been developed and utilized31. Users are required to agree to the DUA before accessing the data. All shared datasets were pseudonymized using masks generated using FSL19,20,21 that were then manually edited to remove all non-brain material that could be useful in re-identification procedures. All users will be asked to sign a DUA detailing their responsibilities and liabilities while the data is in their possession. This is a legally binding agreement that users are expected to comply with in their role as data controllers of the data they process. The DUA would be signed before providing data access.
The brainlife.io CLI can be installed on most Unix/Linux systems using the following command:
npm install brainlife.io -g
The CLI can be used to query and download partial or full datasets. The following example shows the CLI command to download all T1w datasets from a subject in the publication data:
bl pub query # this will return the publication IDs bl bids download–pub 66614215829dd9abdbb3c51c–subject 01–datatype \ neuro/anat/t1w–tag "acq-coronal"
The following command downloads the data in the entire project (from Release 1) into BIDS format:
bl bids download–pub 64233a2a73d2685502db46a3
Additional information about the brainlife.io CLI commands can be found at https://github.com/brainlife/cli.
Code availability
Table 5 reports the links to each web service and GitHub URL implementing the processing pipeline. The ezBIDS GitHub repository can be found at https://github.com/brainlife/ezbids. Analyses using MRIQC outputs were performed using open-source code hosted by brainlife.io and available as a Jupyter Notebook at https://github.com/bacaron/nigerian_brain_analyses/blob/main/nigerian_brain_analyses.ipynb.
References
Donald, K. A. et al. What is next in African neuroscience? Elife 11, (2022).
Lekoubou, A., Echouffo-Tcheugui, J. B. & Kengne, A. P. Epidemiology of neurodegenerative diseases in sub-Saharan Africa: a systematic review. BMC Public Health 14, 653 (2014).
US Census Bureau. Population Aging in Sub-Saharan Africa: Demographic Dimensions 2006. https://www.census.gov/library/publications/2007/demo/p95-07-1.html.
Nigeria. https://www.cia.gov/the-world-factbook/countries/nigeria/.
Okubadejo, N. U., Bower, J. H., Rocca, W. A. & Maraganore, D. M. Parkinson’s disease in Africa: A systematic review of epidemiologic and genetic studies. Mov. Disord. 21, 2150–2156 (2006).
Akinyemi, R. O. et al. Dementia in Africa: Current evidence, knowledge gaps, and future directions. Alzheimers. Dement. 18, 790–809 (2022).
Quarshie, J. T., Mensah, E. N., Quaye, O. & Aikins, A. R. The Current State of Parkinsonism in West Africa: A Systematic Review. Parkinsons Dis. 2021, 7479423 (2021).
Okubadejo, N. U., Ojo, O. O. & Oshinaike, O. O. Clinical profile of parkinsonism and Parkinson’s disease in Lagos, Southwestern Nigeria. BMC Neurol. 10, 1 (2010).
Blanckenberg, J., Bardien, S., Glanzmann, B., Okubadejo, N. U. & Carr, J. A. The prevalence and genetics of Parkinson’s disease in sub-Saharan Africans. J. Neurol. Sci. 335, 22–25 (2013).
Wilkinson, M. D. et al. The FAIR Guiding Principles for scientific data management and stewardship. Sci Data 3, 160018 (2016).
Elsevier. Africa generates less than 1% of the world’s research; data analytics can change that. Elsevier Connect https://www.elsevier.com/connect/africa-generates-less-than-1-of-the-worlds-research-data-analytics-can-change-that (2018).
Hayashi, S. et al. brainlife.io: a decentralized and open-source cloud platform to support neuroscience research. Nat. Methods https://doi.org/10.1038/s41592-024-02237-2 (2024).
Barker, M. et al. Introducing the FAIR Principles for research software. Sci Data 9, 622 (2022).
Levitas, D. et al. ezBIDS: Guided standardization of neuroimaging data interoperable with major data archives and platforms. Sci Data 11, 179 (2024).
Avesani, P. et al. The open diffusion data derivatives, brain data upcycling via integrated publishing of derivatives and reproducible open cloud services. Sci Data 6, 69 (2019).
Stewart, C. A. et al. Jetstream: A self-provisioned, scalable science and engineering cloud environment. https://doi.org/10.1145/2792745.2792774 (2015).
Towns, J. et al. XSEDE: Accelerating Scientific Discovery. Computing in Science Engineering 16, 62–74 (2014).
Esteban, O. et al. MRIQC: Advancing the automatic prediction of image quality in MRI from unseen sites. PLoS One 12, e0184661 (2017).
Woolrich, M. W. et al. Bayesian analysis of neuroimaging data in FSL. Neuroimage 45, S173–86 (2009).
Smith, S. M. et al. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage 23(Suppl 1), S208–19 (2004).
Jenkinson, M., Beckmann, C. F., Behrens, T. E. J., Woolrich, M. W. & Smith, S. M. FSL. Neuroimage 62, 782–790 (2012).
Merkel, D. Docker: lightweight Linux containers for consistent development and deployment. Linux J. 2014, 2 (2014).
Nichols, T. E. et al. Best practices in data analysis and sharing in neuroimaging using MRI. Nat. Neurosci. 20, 299–303 (2017).
Antonoplis, S. Studying Socioeconomic Status: Conceptual Problems and an Alternative Path Forward. Perspect. Psychol. Sci. https://doi.org/10.1177/17456916221093615 (2022).
Gorgolewski, K. J. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci. Data 3, (2016).
Creators McCarthy, P. FSLeyes. https://doi.org/10.5281/zenodo.10122614.
Wogu, E., Filima P.L., Levitas, D., Caron, B., Pestilli, F. A labeled Clinical-MRI dataset of Nigerian brains - Pseudonymized data. brainlife. https://doi.org/10.25663/brainlife.pub.61.
Caron, B. & Pestilli, F. A labeled Clinical-MRI dataset of Nigerian brains - Brain Masks. brainlife.io https://doi.org/10.25663/brainlife.pub.55 (2024).
Eke, D. O. et al. International data governance for neuroscience. Neuron https://doi.org/10.1016/j.neuron.2021.11.017 (2021).
Ochang, P., Stahl, B. C. & Eke, D. The ethical and legal landscape of brain data governance. PLoS One 17, e0273473 (2022).
Eke, D. et al. Pseudonymisation of neuroimages and data protection: Increasing access to data while retaining scientific utility. Neuroimage: Reports 1, 100053 (2021).
Acknowledgements
National Science Foundation (NSF) awards 1916518, 1912270, 1636893, and 1734853. National Institutes of Health awards (NIH) R01MH126699, R01EB030896, R01EB029272, U24NS140384, and a Microsoft Investigator Fellowship to Franco Pestilli. NIH UM1-NS132207 to K. Ugurbil. A Wellcome Trust award (226486/Z/22/Z) and gifts from the Kavli Foundation to Franco Pestilli and Damian Eke.
Author information
Authors and Affiliations
Contributions
Eberechi Wogu: Conceptualization, Methodology, Investigation, Writing–original draft, Writing-review & editing, Visualization. Patrick Filima: Conceptualization, Investigation, Data Curation. Bradley Caron: Data Curation, Software, Formal analysis, Writing–original draft, Writing-review & editing. Daniel Deabler: Data Curation, Software, Writing–original draft, Writing-review & editing. Peer Herholz: Data Curation, Software, Writing–original draft, Writing-review & editing. Catherine Leal: Formal analysis. Mohammed F. Mehboob: Formal analysis. Sohmee Kim: Formal analysis. Ananya Gosain: Formal analysis. Alisha Flexwala: Formal Analysis Soichi Hayashi: Software. Simisola Akintoye: Conceptualization. George Ogoh: Conceptualization. Tawe Godwin: Investigation. Damian Eke: Conceptualization, Writing–original draft, Writing-review & editing, Supervision, Funding acquisition. Franco Pestilli: Conceptualization, Software, Writing-review & editing, Supervision, Funding acquisition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wogu, E., Filima, P., Caron, B. et al. A labeled Clinical-MRI dataset of Nigerian brains. Sci Data 12, 518 (2025). https://doi.org/10.1038/s41597-025-04743-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-04743-0