Abstract
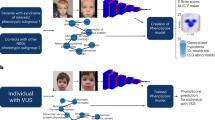
Distinctive facial phenotypes serve as crucial diagnostic markers for many rare genetic diseases. Although AI-driven image recognition achieves high diagnostic accuracy, it often fails to explain its predictions. In this study, we present the Facial phenotype-Gene-Disease Dataset (FGDD), an explainable dataset collected from 509 research publications. It contains 1,147 data records encompassing 197 disease-causing genes, 437 facial phenotypes, and 211 disease entities, with 689 records having disease labels. Each data record represents a patient group and includes demographic information, variation information, and phenotype information. Baseline and explainability validations conducted on FGDD confirmed the dataset’s effectiveness. FGDD supports the training of diagnostic models for rare genetic diseases while delivering explainable results, and provides a foundation for exploring intricate connections between genes, diseases, and facial phenotypes.
Similar content being viewed by others
Background & Summary
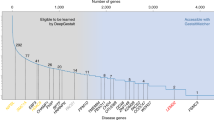
More than 6% of the world’s population is affected by rare genetic diseases1, and the latest Orphanet2 and OMIM3 databases show that there are currently at least 7000 rare genetic diseases. Among these, many rare genetic diseases have recognizable facial phenotypic features, and facial phenotypes are often used as a basis for diagnosis4,5,6,7. In recent studies, the diagnosis of rare genetic diseases through computer vision techniques has reached the level of clinical experts8,9,10,11,12,13,14.
DeepGestalt9, trained on a private dataset comprising over 17,000 images, demonstrates the ability to identify the correct disease among 502 different images, achieving an impressive top-10 accuracy of 91%. However, like many AI models, DeepGestalt lacks the capability to explicitly explain its predictions or provide insights into which specific facial features contributed to the diagnosis. AI models rely heavily on data, and the GestaltMatcher Database (GMDB)15 is the only publicly available dataset in the field, containing 10,189 frontal images of 7,695 patients with 683 diseases. Despite its utility, models trained on GMDB share the same interpretability shortcomings as DeepGestalt. In the medical field, where explainability is paramount16,17,18, the lack of transparent and explainable datasets poses a significant barrier.
Compared to the GMDB image format, the tabular format is naturally more explainable. Tabular dataset with clear meanings and units, the relationship between data is more direct and easier to be understood by humans19. For instance, when we see “age” and “disease” in the table, we can intuitively understand the correlation between them. Moreover, the tabular dataset consists of numerical values and categories that can be directly understood by humans, while GMDB image data consists of pixel values that do not have direct semantic information. Furthermore, some logical rules can be found in tabular data, such as “c.6726_6730del; p.Leu2243Serfs*8 in Exon 20 cause Coffin-Siris syndrome 1”, whereas for image data, what the model learns tends to be high-level feature combinations.
Therefore, We propose a new tabular dataset, FGDD, which contains 1147 data records, 197 associated genes, 437 associated phenotypes, and 211 associated diseases, of which 689 data records have disease labels. FGDD was constructed by retrieving publications from Human Phenotype Ontology (HPO20)-generated terms, and then identifying facial phenotypes-gene-disease associations from these publications.
The FGDD is primarily used for facial phenotype analysis of rare genetic diseases. It serves multiple purposes, including training explainable diagnostic models, conducting in-depth analysis of the complex relationships between genes, diseases, and facial phenotypes, and uncovering additional potential associations and patterns.
Our contributions can be summarized as follows:
-
We propose a new dataset, FGDD, for facial phenotype analysis of rare genetic diseases, which can be used not only for training explainable diagnostic models but also for in-depth analysis of the complex gene-disease-facial phenotype relationships and for mining more potential associations and patterns.
-
We have conducted extensive benchmarking on our dataset, and commonly used algorithms can achieve up to 80.19% accuracy and provide clinical support.
-
We performed feature importance evaluation from both coarse- and fine-grained perspectives to explain relative contributions across feature categories and individual features within each category.
An overview of this study is shown in Fig. 1. FGDD dataset is available at figshare21. All codes are publicly available at https://github.com/zhelishisongjie/FGDD and can be audited, copied, and reused.
The overall process of this study.
Methods
Data retrieval
We used a systematic search strategy to build the publications list as follows: firstly, we built the base search terms based on the concept of facial phenotype and its synonyms in the Human Phenotype Ontology (HPO20), and then logically combined them with the terms related to genetic variation (including “genetic”, “variation”, “deletion”), and finally, we generated 595 composite search terms, with a deadline of 1 April 2023.
An initial search of the PubMed database yielded 26,814 publications, the details of these publications were obtained through the Entrez programming utilities22, and 11,304 publications were retained after the de-duplication process. Subsequently, through a stratified screening process: excluding reviews, commentaries, book chapters and animal experiments (n=6,045), the remaining 5,259 publications entered the full-text assessment stage, and those with insufficient relevance or unclear conclusions were excluded from the full-text screening (n=4,750), and 509 high-quality publications were finally included for data extraction. The complete search strategy (including the search term construction code, the search terms list, and the final included publications list) has been made publicly available at our figshare21 and Github repository to ensure the reproducibility of the research.
Data extraction
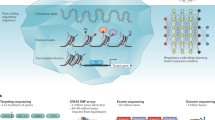
In the data extraction phase, there are four types of information to be extracted for each publication, including demographic information, phenotype information, variation information, and disease information. We constructed a hybrid automated-manual workflow: firstly, we implement basic information extraction by PhenoTagger23 (a human phenotype entity recognition tool) and PubTator24 (a biomedical entity recognition tool), where PhenoTagger is responsible for extracting the standardized phenotype information, and PubTator is responsible for capturing the variation information and disease information. However, existing automated tools have significant technical limitations: 1) although PhenoTagger can recognize phenotypic entities in text, it is unable to distinguish which patient the phenotype belongs to; 2) PubTator lacks standardized support for variant forms described by HGVS nomenclature (e.g., c.898C>T; p.Arg300Cys); 3) demographic information (e.g., age, gender, ethnicity) requires manual parsing due to the highly heterogeneous representation. The complete extraction of the above three types of information relies on the researcher’s in-depth reading of the text.
Finally, we manually integrated the raw data extracted from each publication according to the current format of FGDD, and to ensure the data quality, the publications were read again for data checking to make sure that there were no correspondence mismatches or data misentry. (After M.H. completed the raw data collection, data checking was performed by S.J.). The overall process of data collection is shown in Fig. 2.
The process of data collection.
Data Records
FGDD, available for download on figshare21, provides tabular data in .csv format and a knowledge graph in .dump format. The dataset has a Creative Commons Attribution 4.0 International (CC BY 4.0) license. The figshare repository is organized as shown in Fig. 3. “Search terms” directory stores the data generated during the search and screening process to ensure the reproducibility of the study. “Raw data” directory stores data extracted from research publications, covering facial phenotypes, genes, disease information, and associations with patients, of which data with clear associations with patients are used for subsequent integration (some genes and facial phenotypes are only mentioned in the publications, and are not directly associated with the patients), and “FGDD” directory stores the standardized dataset and its knowledge graph format based on the final systematic integration of the raw data. Table 1 provides an overview of the files stored in figshare21.
Organization of dataset.
Technical Validation
Overview of facial phenotype-gene-disease data in FGDD
The data distribution is illustrated in Fig. 4, providing a clear visual representation of the dataset’s key characteristics. Table 2 systematically presents the data completeness analysis for each feature dimension. Based on the feature attributes, the variables in this study were classified into four categories: demographic features, facial phenotype features, variation features, and disease features. notably, the group of disease-associated features presented the highest missing rate (39.93%). This is mainly attributed to the fact that most genetic articles have focused on the functional association of facial dysmorphisms with specific genetic variants, with an emphasis on the molecular and mechanistic mechanisms, and less on the association with clinical diseases.
(a) Patient’s Regional Distribution on a World Map, where Asia has the highest number of patients. (b) Patient’s Regional Distribution Bar Chart, a bar chart quantifying differences in the number of patients by region. (c) Gene Distribution, Chr is an abbreviation for Chromosome. (d)Age and Gender Distribution, age distribution (upper figure), with the majority of patients aged 10 years or older in years. Gender distribution (lower figure), where Male&Female indicates that this group of patients contains more than or equal to 2 individuals of different genders. (e) Disease Distribution, a bar chart showing the distribution of diseases. (f) Facial Phenotypes Distribution, with the most common facial phenotype being hypertelorism.
To better visualize the facial phenotype-gene-disease relationships, we present FGDD in a format of knowledge graph, the schema of the knowledge graph is shown in Fig. 5(a), and a specific visualization example is shown in Fig. 5(b). Python script and tutorial for transforming FGDD tabular into knowledge graph are given in https://github.com/zhelishisongjie/FGDD.
(a) Knowledge graph schema, which includes four types of nodes with four types of relationships. (b) A Knowledge graph visualization example, displayed through the yworks neo4j explorer.
Common algorithms performance validation
We tested the performance of common classification algorithms by splitting the training and test sets using a ratio of 7:3. All algorithms are in regular configuration and can be simply reproduced using our codes. The results are shown in Table 3. TabNet, Node, TabTransformer, FTTransformer are the popular deep learning algorithms for tabular data, Node achieves the highest Top-1 accuracy (80.19%), FTTransformer achieves the highest Macro-F1 score (0.59).
Macro-F1 is a commonly used evaluation metric in multiclassification tasks that reflects the model’s balanced performance on all categories. Macro-F1 is defined as shown in equation (1). TPi, FPi, FNi represent the true positive, false positive, and false negative cases of disease i, respectively.
Explainability validation
Explainability validation is concerned with understanding the logic of the whole model and tries to explain how the model is obtained through learning. Here we focus on feature importance analysis from both coarse-grained and fine-grained perspectives.
Coarse-grained features: We classify features into three main categories: patients, variations, and phenotypes. Coarse-grained features analyze the overall importance of these three categories of features, aiming to understand the contribution of each category to the model’s decision-making at a macro level. Notably, patient metadata demonstrates limited influence as shown in Fig. 6.
Coarse-grained features importance. This suggests the disease pathogenesis is primarily driven by genetic rather than environmental/lifestyle factors. Genetic variants and facial phenotypes are central to diagnosis; integrating genomic analysis with phenotypic evaluation (e.g., distinct facial features) enhances diagnostic precision.
Fine-grained features: Unlike coarse-grained features, fine-grained features analyze the three categories of patients, variations, and phenotypes individually, aiming to reveal the importance of specific features within each category at the micro level. Genetic variation features emerge as the predominant diagnostic determinants as shown in Fig. 7.
Fine-grained features importance. Key genomic features (e.g., exon count, chromosomal location) and distinctive facial phenotypes are critical diagnostic parameters. This underscores the necessity of integrating genomic profiling with phenotypic evaluation for precise diagnosis. Furthermore, patient-specific factors including ethnicity and geographic origin must be considered, as certain diseases exhibit population-stratified prevalence patterns requiring individualized diagnostic frameworks.
Limitations and future work
Mechanistic interpretability
The current dataset also lacks data on the biological mechanisms that underlie diagnostic decisions at a deeper level. Future datasets should include proteins, complexes, pathways, and biological processes to deepen biological insights. For example, in COFFIN-SIRIS SYNDROME, the most common genetic cause is a mutation in the ARID1B gene25,26.
ARID1B encodes a subunit of the Brg1/Brm associated factor (BAF) complex (a core component of the SWI/SNF chromatin remodeling complex), regulating gene expression in cell differentiation, neural development, and DNA repair27. Pathogenic mutations block the ARID1A-to-ARID1B subunit switch in BAF, causing sustained pluripotency gene activation (NANOG/SOX2)28. This disrupts cranial neural crest cell (CNCC) differentiation/migration and neuroectodermal maturation, leading to craniofacial anomalies (short philtrum, thick eyebrows, abnormal lips) and intellectual disability. Pathogenic mutations in SWI/SNF (e.g., ARID1B) is central to COFFIN-SIRIS SYNDROME29,30,31. Systematic incorporation of such molecular mechanisms will bridge the gap between genetic variants, disease, and phenotype.
Generalizability to different populations
There is a notable population region bias in the collected data, with certain racial or ethnic groups being overrepresented while others are underrepresented. This imbalance in distribution poses significant challenges in ensuring the model’s generalizability across diverse populations. To address this issue, several strategies can be implemented. First, to increase the size of the dataset and ensure a balanced distribution of racial patients; second, to use data augmentation/generation techniques32,33 to generate diverse data and to develop weighted learning algorithms34 to reweight the imbalanced population. In addition, multimodal learning can be used in conjunction with other image datasets to combine demographic, phenotypic, and genetic variation information with visual information to enhance generalization. For instance, a multimodal knowledge graph35 can link image features to diseases, phenotypes, and variations, enabling effective fusion through graph neural networks for tasks like diagnosis and phenotype prediction.
Code availability
The code and its description, required packages, installation tutorials, data preprocessing, and specific parameters are publicly available at https://github.com/zhelishisongjie/FGDD.
References
Ferreira, C. R. The burden of rare diseases. American journal of medical genetics Part A 179, 885–892, https://doi.org/10.1002/ajmg.a.61124 (2019).
Weinreich, S. S., Mangon, R., Sikkens, J., Teeuw, M. E. & Cornel, M. Orphanet: a european database for rare diseases. Nederlands tijdschrift voor geneeskunde 152, 518–519 (2008).
Hamosh, A., Scott, A. F., Amberger, J. S., Bocchini, C. A. & McKusick, V. A. Online mendelian inheritance in man (omim), a knowledgebase of human genes and genetic disorders. Nucleic acids research33, D514–D517, https://doi.org/10.1093/nar/gki033 (2005).
Helman, S. N., Badhey, A., Kadakia, S. & Myers, E. Revisiting crouzon syndrome: reviewing the background and management of a multifaceted disease. Oral and maxillofacial surgery18, 373–379, https://doi.org/10.1007/s10006-014-0467-0 (2014).
Donnai, D. & Karmiloff-Smith, A. Williams syndrome: From genotype through to the cognitive phenotype. American journal of medical genetics 97, 164–171 (2000).
Vergano, S. S. & Deardorff, M. A. Clinical features, diagnostic criteria, and management of coffin–siris syndrome. In American Journal of Medical Genetics Part C: Seminars in Medical Genetics, vol. 166, 252–256, https://doi.org/10.1002/ajmg.c.31411 (Wiley Online Library, 2014).
Hunt, J. A. & Hobar, C. P. Common craniofacial anomalies: the facial dysostoses. Plastic and reconstructive surgery 110, 1714–1725, https://doi.org/10.1097/01.prs.0000033869.10382.91 (2002).
Thevenot, J., López, M. B. & Hadid, A. A survey on computer vision for assistive medical diagnosis from faces. IEEE journal of biomedical and health informatics22, 1497–1511, https://doi.org/10.1109/JBHI.2017.2754861 (2017).
Gurovich, Y. et al. Identifying facial phenotypes of genetic disorders using deep learning. Nature medicine 25, 60–64, https://doi.org/10.1038/s41591-018-0279-0 (2019).
Bannister, J. J. et al. A deep invertible 3-d facial shape model for interpretable genetic syndrome diagnosis. IEEE journal of biomedical and health informatics26, 3229–3239, https://doi.org/10.1109/JBHI.2022.3164848 (2022).
Hallgrímsson, B. et al. Automated syndrome diagnosis by three-dimensional facial imaging. Genetics in medicine22, 1682–1693, https://doi.org/10.1038/s41436-020-0845-y (2020).
Jin, B., Cruz, L. & Gonçalves, N. Deep facial diagnosis: deep transfer learning from face recognition to facial diagnosis. IEEE Access8, 123649–123661, https://doi.org/10.1007/978-3-030-33128-3_1 (2020).
Pooch, E. H. P., Alva, T. A. P. & Becker, C. D. L. A computational tool for automated detection of genetic syndrome using facial images. In Intelligent Systems: 9th Brazilian Conference, BRACIS 2020, Rio Grande, Brazil, October 20–23, 2020, Proceedings, Part I 9, 361–370, https://doi.org/10.1007/978-3-030-61377-8_25 (Springer, 2020).
Singh, A. & Kisku, D. R. Detection of rare genetic diseases using facial 2d images with transfer learning. In 2018 8th International Symposium on Embedded Computing and System Design (ISED), 26–30, https://doi.org/10.1109/ISED.2018.8703997 (IEEE, 2018).
Lesmann, H. et al. Gestaltmatcher database-a global reference for facial phenotypic variability in rare human diseases. medRxiv 2023–06, https://doi.org/10.1101/2023.06.06.23290887 (2023).
Yoon, C. H., Torrance, R. & Scheinerman, N. Machine learning in medicine: should the pursuit of enhanced interpretability be abandoned? Journal of Medical Ethics48, 581–585, https://doi.org/10.1136/medethics-2020-107102 (2022).
Vellido, A. The importance of interpretability and visualization in machine learning for applications in medicine and health care. Neural computing and applications32, 18069–18083, https://doi.org/10.1007/s00521-019-04051-w (2020).
Teng, Q., Liu, Z., Song, Y., Han, K. & Lu, Y. A survey on the interpretability of deep learning in medical diagnosis. Multimedia Systems28, 2335–2355, https://doi.org/10.1007/s00530-022-00960-4 (2022).
Rudin, C. et al. Interpretable machine learning: Fundamental principles and 10 grand challenges. statistics surveys 16 (none): 1–85, https://doi.org/10.1214/21-SS133 (2022).
Köhler, S. et al. The human phenotype ontology in 2021. Nucleic acids research49, D1207–D1217, https://doi.org/10.1093/nar/gkaa1043 (2021).
Song, J., He, M., Ren, S. & Shen, B. An explainable dataset linking facial phenotypes and genes to rare genetic diseases. figsharehttps://doi.org/10.6084/m9.figshare.28516604.v3 (2025).
Buchmann, J. P. & Holmes, E. C. Entrezpy: a python library to dynamically interact with the ncbi entrez databases. Bioinformatics35, 4511–4514, https://doi.org/10.1093/bioinformatics/btz385 (2019).
Luo, L. et al. Phenotagger: a hybrid method for phenotype concept recognition using human phenotype ontology. Bioinformatics37, 1884–1890, https://doi.org/10.1093/bioinformatics/btab019 (2021).
Wei, C.-H., Kao, H.-Y. & Lu, Z. Pubtator: a web-based text mining tool for assisting biocuration. Nucleic acids research41, W518–W522, https://doi.org/10.1093/nar/gkt441 (2013).
van der Sluijs, P. J. et al. The arid1b spectrum in 143 patients: from nonsyndromic intellectual disability to coffin–siris syndrome. Genetics in Medicine21, 1295–1307, https://doi.org/10.1038/s41436-018-0330-z (2019).
Vasko, A., Drivas, T. G. & Schrier Vergano, S. A. Genotype-phenotype correlations in 208 individuals with coffin-siris syndrome. Genes12, 937, https://doi.org/10.3390/genes12060937 (2021).
Aref-Eshghi, E. et al. Bafopathies’ dna methylation epi-signatures demonstrate diagnostic utility and functional continuum of coffin–siris and nicolaides–baraitser syndromes. Nature communications9, 4885, https://doi.org/10.1038/s41467-018-07193-y (2018).
Pagliaroli, L. et al. Inability to switch from arid1a-baf to arid1b-baf impairs exit from pluripotency and commitment towards neural crest formation in arid1b-related neurodevelopmental disorders. Nature communications12, 6469, https://doi.org/10.1038/s41467-021-26810-x (2021).
Santen, G. W. et al. Mutations in swi/snf chromatin remodeling complex gene arid1b cause coffin-siris syndrome. Nature genetics44, 379–380, https://doi.org/10.1038/ng.2217 (2012).
Tsurusaki, Y. et al. Mutations affecting components of the swi/snf complex cause coffin-siris syndrome. Nature genetics44, 376–378, https://doi.org/10.1038/ng.2219 (2012).
Wieczorek, D. et al. A comprehensive molecular study on coffin–siris and nicolaides–baraitser syndromes identifies a broad molecular and clinical spectrum converging on altered chromatin remodeling. Human molecular genetics22, 5121–5135, https://doi.org/10.1093/hmg/ddt366 (2013).
Hernandez, M., Epelde, G., Alberdi, A., Cilla, R. & Rankin, D. Synthetic data generation for tabular health records: A systematic review. Neurocomputing493, 28–45, https://doi.org/10.1016/j.neucom.2022.04.053 (2022).
Nik, A. H. Z., Riegler, M. A., Halvorsen, P. & Storås, A. M. Generation of synthetic tabular healthcare data using generative adversarial networks. In International Conference on Multimedia Modeling, 434–446, https://doi.org/10.1007/978-3-031-27077-2_34 (Springer, 2023).
Huang, L. et al. Gradient attention balance network: Mitigating face recognition racial bias via gradient attention. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, 38–47, https://doi.org/10.48550/arXiv.2304.02284 (2023).
Zhu, X. et al. Multi-modal knowledge graph construction and application: A survey. IEEE Transactions on Knowledge and Data Engineering36, 715–735, https://doi.org/10.1109/TKDE.2022.3224228 (2022).
Arik, S. Ö. & Pfister, T. Tabnet: Attentive interpretable tabular learning. In Proceedings of the AAAI conference on artificial intelligence, vol. 35, 6679–6687, https://doi.org/10.1609/aaai.v35i8.16826 (2021).
Popov, S., Morozov, S. & Babenko, A. Neural oblivious decision ensembles for deep learning on tabular data. arXiv preprint arXiv:1909.06312,https://doi.org/10.48550/arXiv.1909.06312 (2019).
Huang, X., Khetan, A., Cvitkovic, M. & Karnin, Z. Tabtransformer: Tabular data modeling using contextual embeddings. arXiv preprint arXiv:2012.06678,https://doi.org/10.48550/arXiv.2012.06678 (2020).
Gorishniy, Y., Rubachev, I., Khrulkov, V. & Babenko, A. Revisiting deep learning models for tabular data. Advances in Neural Information Processing Systems34, 18932–18943 (2021).
Acknowledgements
The research was supported by National Supercomputing Center in Chengdu.
Author information
Authors and Affiliations
Contributions
Jie Song: Data Curation, Formal Analysis, Validation, Writing - Original Draft, Writing - Review & Editing; Mengqiao He: Data Curation, Writing - Review & Editing; Shumin Ren: Writing - Review & Editing; Bairong Shen: Conceptualization, Supervision, Writing - Review & Editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Song, J., He, M., Ren, S. et al. An explainable dataset linking facial phenotypes and genes to rare genetic diseases. Sci Data 12, 634 (2025). https://doi.org/10.1038/s41597-025-04922-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-04922-z
This article is cited by
-
WDR26-related Skraban–Deardorff syndrome: clinical, genetic and pathomechanistic insights
European Journal of Medical Research (2025)
-
Face-based machine learning diagnostics: applications, challenges and opportunities
Artificial Intelligence Review (2025)