Abstract
Cultivated strawberry (Fragaria × ananassa) belongs to the family Rosaceae and is an allo-octoploid species (2n = 8× = 56). Using PacBio Revio long reads of ‘Seolhyang’, we completed telomere-to-telomere phased genome assemblies with a size of 797 Mb with a contig N50 of 27.04 Mb. Benchmarking of the universal single-copy orthologs (BUSCO) analysis detected 99.1% conserved genes in the assembly. In addition, the average long terminal repeat assembly index (LAI) was 17.28, with high genome continuity. In this study, we identified 50 of the possible 56 telomeres across 28 chromosomes. The ‘Seolhyang’ genome was annotated using RNA-Seq data representing various F. × ananassa tissues from the NCBI sequence read archive, which resulted in 129,184 genes.
Similar content being viewed by others
Background & Summary
The cultivated strawberry (Fragaria × ananassa), a perennial plant belonging to the Rosaceae family, is an allo-octoploid species with a highly heterozygous genome that contributes to its genetic complexity and diverse phenotypic traits. This complexity poses a significant challenge for genetic research and breeding programs. Strawberries are a globally crucial crop, with the United Nations Food and Agricultural Organization (UN-FAO) reporting worldwide production of 9.57 million tons in 2022 (https://www.fao.org/faostat/). In South Korea, strawberries are a significant economic crop, with a cultivation area of 5,745 ha and a production volume of 158,807 tons in 20221. The domestic production value of strawberries in South Korea is approximately USD 932 million, accounting for 14.7% of the total vegetable production value in the country2.
Among the various Korean strawberry cultivars, ‘Seolhyang’ (‘Akihime’ × ‘Red Pearl’), developed in 20053, dominates the South Korean market, occupying 82.1% of the total strawberry cultivation area in 20224. ‘Seolhyang’ is favored for its ease of cultivation; large fruit size; high yields5,6,7; and resistance to diseases such as angular leaf spot, anthracnose, and powdery mildew3,8,9,10. In an analysis of 45 representative Korean cultivars and genetic resources, ‘Seolhyang’ was distinguished by having the highest overall concentration of volatile organic compounds (VOCs)11. Various breeding programs have been initiated to harness the desirable traits of the elite cultivar ‘Seolhyang’. However, progress in precision breeding efforts has been hindered by limited genomic research on ‘Seolhyang’.
The availability of reference genomes has substantially affected agricultural research and has driven significant advancements in the understanding of the genetic basis of plant traits. This genomic insight reveals how artificial selection shapes these traits over time. This has deepened the understanding of how genetic characteristics influence interactions within agricultural ecosystems, particularly with pathogens and insects12,13. Recently, the assembly of reference genomes in agriculture has undergone significant advancements, particularly owing to the integration of third-generation sequencing technology14. These developments have enhanced the quality and completeness of plant reference genomes. High-throughput sequencing methods, such as next-generation sequencing (NGS), have enabled the generation of extensive genomic data. However, to overcome the limitations associated with short-read sequences in contigs and scaffolds, long-read sequencing technologies, such as PacBio, BioNano, and Nanopore, have emerged as pivotal tools for third- and fourth-generation sequencing15,16. Pacific Biosciences (PacBio) High-Fidelity (HiFi) sequencing technology generates long reads with an average length ranging from 10 to 25 kb and an error rate of less than 0.5%. This level of accuracy and read length position of HiFi sequencing is the primary source of data for producing high-quality genome assemblies17,18. Advances have addressed some of these challenges, particularly regarding the assembly of telomere-to-telomere (T2T) gap-free reference genomes. Notably, for cultivated and diploid strawberries19,20,21,22,23, there has been the successful assembly of such high-quality genomes for the ‘Hawaii 4’, ‘Benihoppe’ and ‘Florida Brilliance’ cultivars, providing more reliable references in currently available genomic resources.
In this study, a high-quality genome assembly of the strawberry cultivar ‘Seolhyang’ was generated using approximately 100 Gb of HiFi sequencing data obtained from the PacBio Revio platform. Unlike previous assembly methods for octoploid strawberry genomes, this assembly was completed without incorporating data from additional sequencing platforms, resulting in a high-quality reference genome comparable to those of ‘Royal Royce’ and ‘Florida Brilliance.’ We completed a telomere-to-telomere genome assembly with a genome size of 797 Mb and a contig N50 of 27.04 Mb. Benchmarking of the universal single-copy orthologs (BUSCO) analysis detected 99.1% conserved genes in the assembly. In addition, the average of long terminal repeat assembly index (LAI) was 17.28, reflecting the overall high genome continuity based on analysis of intact and total LTR retrotransposons measured using Extensive de novo TE Annotator (EDTA) followed by LTR retriever. Notably, we identified 50 of the possible 56 telomeres across 28 chromosomes. The ‘Seolhyang’ genome was annotated using RNA-Seq data representing various F. × ananassa tissues from the NCBI for Biotechnology Information sequence read archive, which resulted in 129,184 genes. Powdery mildew is a significant disease frequently observed in controlled cultivation environments, such as plastic greenhouses, posing substantial challenges to strawberry production. The strawberry cultivar ‘Seolhyang’ is well known for its resistance to powdery mildew. This study utilized the assembled genome of ‘Seolhyang’ to investigate the genetic basis of its resistance, focusing on the MLO (Mildew Locus O) genes, which have been reported to be associated with powdery mildew resistance. A total of 55 MLO genes were identified in the ‘Seolhyang’ genome. Their structures and domains were systematically compared with 20 MLO genes previously reported in diploid strawberries and 69 MLO genes identified in the octoploid strawberry ‘Camarosa.’ These comparisons provide valuable insights into the unique genetic characteristics underlying the powdery mildew resistance of ‘Seolhyang’, suggesting that the genome of ‘Seolhyang’ will be a promising genetic resource for the identification studies of powdery mildew resistance genes and development of resistant cultivars.
Methods
Materials and DNA sequencing
The cultivated strawberry (F. × ananassa) cultivar ‘Seolhyang’ was used for genome sequencing. Young leaves were covered with black plastic bags and stored in a greenhouse for 14 d. The etiolated leaf tissue was harvested for DNA extraction. The leaves were frozen and subjected to genomic DNA extraction and library preparation by using DNA Link (Seoul, South Korea). The single-molecule real-time sequencing (SMRT) bell library for ‘Seolhyang’ was constructed using a PacBio DNA Template Prep Kit 3.0 (Pacific Biosciences, CA, USA). PacBio’s standard protocol (Pacific Biosciences, CA, USA) was used to build the SMRTbell target-size libraries. The library was sequenced using the PacBio Revio System (DNA Link, Seoul, South Korea).
De Novo genome assembly and validation
Figure 1 illustrates the workflow for the genome assembly and annotation implemented in this study. HiFi reads were used to produce a draft assembly without sequencing the parents by using Hifiasm ver. 0.16.124. Hifiasm was run with the following commands, according to the developer’s recommendations for heterozygous polyploid crops. The contigs were scaffolded and oriented based on the reference genome of ‘Florida Brilliance’ (https://www.rosaceae.org/Analysis/14031408) by using RagTag25.
Workflow implemented for ‘Seolhyang’ genome assembly and annotation.
Genome assembly statistics were calculated using QUAST version 5.0.26626. Merqury version 1.3 was used to measure the assembly consensus quality value (QV) and to evaluate the assembly based on efficient K-mer set operations27. The completeness of the genome assembly and protein-coding gene annotations were assessed using the BUSCO database28. The long terminal repeat (LTR) assembly index (LAI)29 for each sub-genome was calculated using LTR-retriever30 along with whole-genome Transposable elements (TE)-annotations and intact LTR retrotransposons identified using EDTA31.
Genome annotation
The TEs were annotated using EDTA v1.9.6 with default parameters31. A TE annotation library was generated in separate runs by using EDTA. The TE regions of haploid assembly were masked using the ReapeatMasker v4.1.1 provided with the repeat library. Simple sequence repeats (SSRs) or microsatellites were mined using the SSR Finder on the Genome Sequence Annotation Server v6.0 (GenSAS; https://www.gensas.org)32. To increase the accuracy of gene annotation, we generated a transcriptome assembly containing possible sets of transcripts from ‘Seolhyang’ and publicly available F. × ananassa expression data. Read alignments were converted to the Binary alignment map (BAM) format by using SAMTools. Splice junctions for all merged RNA alignments were predicted and trimmed using Portcullis v1.2.233. Genome assemblies were annotated using Braker2. Functions of the predicted transcripts were annotated based on alignment by using BlastP v2.2.2834 in the UniProtKB database35.
Collinearity and synteny
Genomic synteny at the DNA level among F. vesca36, F. × ananassa cultivars ‘Royal Royce’37, and ‘Florida Brilliance’ (https://www.rosaceae.org/Analysis/14031408), and ‘Seolhyang’ was visualized using D-GENIES38 by applying default parameters after alignment with minimap239. Candidate structural variations were explored using SYRI40.
Technical Validation
Details of the sequencing data are shown in Table 1. With one single-molecule real-time cell on the PacBio Revio platform, 103.3 Gb of the sequence was generated in 9.1 M reads. The average read length was 17,668 bp with an N50 of 17,769 bp. The assembly contained 2,140 contigs with an N50 of 27.04 Mb. Fifteen contigs accounted for 50% of the total assembly (Table 2). The largest contig size was 36.27 Mb, which covered 99% of the chromosome length. Before scaffolding, BUSCO was 99.1%, and LTR analysis showed that the LAI score was 17.28, indicating the gold standard of the reference genome. Scaffolded contigs resulted in 796.9 Mb of a final genome size. Notably, only 30 contigs were anchored to the final assembly for ‘Seolhyang’.
Identification and characterization of pectin lyase sequence analysis
The sequences with conserved MLO domains (cl03887) were retrieved on Pfam database41. The physical location of the MLO genes was retrieved from the genome annotation file. The conserved motifs were searched using the MEME42 and visualized with gene structure using TBtools43.
Based on the multiple alignment of MLO proteins obtained by the MUSCLE44, a phylogenetic tree was constructed by using the maximum likelihood method in Geneious Prime. The collinear gene pairs were generated using MCScanX45 software. The analysis was conducted using the default parameters of specific software according to the user instructions.
Data Records
The PacBio HiFi sequencing reads used for genome assembly have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject accession number [PRJNA1148756] (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1148756)45.
The chromosome-level genome assembly has been deposited in GenBank under the accession number [JBKFVU000000000] (https://identifiers.org/ncbi/insdc.gca:JBKFVU000000000.1)46.
In addition, the gene annotation files and supplementary materials are available on FigShare (https://doi.org/10.6084/m9.figshare.26866807)47,48.
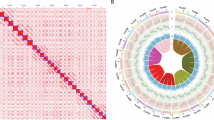
Collinearity between ‘Seolhyang’ and other published F. × ananassa genomes, namely ‘Florida Brilliance’ and ‘Royal Royce’, was confirmed. Translocations on 1D were apparent when the ‘Seolhyang’ genome was compared with the genomes of ‘Florida Brilliance’ (Fig. 2a) and ‘Royal Royce’ (Fig. 2b). Alignments of ‘Seolhyang’ assembly against FaRR1 (‘Royal Royce’) and FaFB1 (‘Florida Brilliance’) also displayed a high degree of collinearity (Figs. 2a and 2b). On the basis of this alignment, we applied the chromosome nomenclature for ‘Seolhyang’ and ‘Royal Royce’, reflecting the putative diploid origins of each respective subgenome (A, B, C, and D)37. Alignments of the ‘Seolhyang’ genome against the diploid F. vesca v4.036 showed a high degree of collinearity except for major translocations on 1 A (Fig. 2c). We confirmed the collinearity and consequently explored the candidate structural variations among ‘Seolhyang’, ‘Florida Brilliance’, and ‘Royal Royce’ by using SYRI41 (Fig. 3). Only ‘Seolhyang’ subgenome A showed higher sequence similarity with diploid F. vesca. Telomeric motifs (5’-TTTAGGG-3’) were explored at the end of each chromosome in the assembly of ‘Seolhyang’. Telomeric motifs enriched in the termini of the pseudo-chromosomes allowed for the identification of 50 telomeres (Table 3). All pseudomolecules contained telomere-rich regions, at least at their ends. Overall, 22 pseudomolecules were potentially telomere-to-telomere, except for Chr 1B, 1 C, 2 A, 3 C, 7 A, and 7B.
Dotplot of ‘Seolhyang’ genome to F. × ananassa cv. Florida Brilliance (a), F. × ananassa cv. Royal Royce (b), and diploid F. vesca ver 4.0 (c). Dot plots were produced using the DGENIE software and alignments with minimap2.
Collinearity analysis between ‘Seolhyang’ genome and other octoploid strawberry genomes, including ‘Florida Brilliance’ (FaFB1) and ‘Royal Royce’ (FaRR1).
Genome annotation
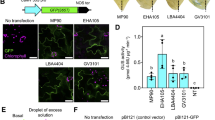
In the ‘Seolhyang’ genome, 346.3 Mb of the repetitive sequence accounted for 43.46% of the genome. Most of this repeat sequence was composed of LTR TEs (25.4%; Table 4). For each chromosome, a genomic region with dense repetitive sequences and a low density of genes, thought to be the centromeres, was identified (Fig. 4). Genome sequences with a long TE (>1 kb) mask were used for gene prediction. De novo prediction of the number of gene-coding proteins in the genome assembly yielded 151,558 transcripts by aligning the RNA-Seq datasets with the assemblies. BUSCO analysis of the transcript assemblies revealed 2,275 complete core eudicot genes (97.8%, 3.5% single-copy, 94.3% duplicated), with 0.5% fragmented and 1.7% missing core eudicot genes. In total, 129,184 genes remained in the ‘Seolhyang’ genome (Table 5).
Distribution of transposable elements and genes in ‘Seolhyang’ genome. (a) length of assembled chromosomes, (b) distribution of DNA transposable elements, and (c) distribution of genes.
Identification of FaMLOs in ‘Seolhyang’ genome assembly
A total of 55 FaMLO genes with MLO domains (cl03887) were identified. According to their homology to FveMLO genes from F. vesca, all FaMLO genes were renamed as FaMLO01C to FaMLO20D (Fig. 5). A maximum of five FaMLO genes were located on chromosome 3 C, while there were no FaMLO genes on chromosome 4 A, 4B, 4 C, and 4D. The characteristics properties of the deduced 55 FaMLO is shown in Table 6. The number of amino acids varied from 171 to 934 aa, most of them (53) were concentrated from 400 to 600 aa. There were only one FaMLO proteins comprising amino acids below 200 aa.
Chromosomal distribution and location of FaMLOs in ‘Seolhyang’ strawberry. Different colors indicate the chromosomes from different subgenomes of cultivated strawberry.
According to phylogenetic analysis for FaMLO genes identified in the present study and previously reported, all the fifty-five FaMLO genes were classified into seven clusters (Fig. 6a). Among them, clade 1 is the largest clade containing 14 members, followed by group 7, which had 11 members of FaMLO genes. To better elucidate the structural characteristics of the FaMLO genes, CDS distributions were analyzed and visualized (Fig. 6b).
Classification and characterization of FaMLO identified in the genome assembly of ‘Seolhyang’. (a) Phylogenetic tree of FaMLOs from diploid and octoploid strawberries. Different branch colors represent the different groups. MLO family members from ‘Seolhyang’ strawberry identified in this study are marked with blue circles. The red stars and black rectangles indicate the previously reported FaMLOs in Fragaria vesca and F. × ananassa var. ‘Camarosa’. (b) Gene structure and conserved domain analysis of FaMLOs. Left part indicated an unroot tree of strawberry FaMLOs, middle part showed the exon–intron distribution of FaMLOs, and the right part displays the distribution of conserved domain on each FaMLO protein.
The collinearity analysis among woodland strawberry (F. vesca), and octoploid strawberry ‘Seolhyang’ was carried out to explore the evolutionary relationship of FaMLOs. According to the result, 55 FaMLOs and 17 FveMLOs were involved to form collinear pairs and were highlighted (Fig. 7).
Collinearity analysis of MLO genes among Fragaria vesca, and Fragaria × ananassa genomes. Grey lines indicate collinear blocks within the two genomes, while the red lines represent collinear MLO gene pairs. The orange and green columns indicate the chromosomes from Fragaria vesca, and Fragaria × ananassa genomes, respectively. Chromosome numbers are displayed at the side of chromosomes.
Code availability
All software and pipelines were executed according to the guidelines and protocols outlined in the respective bioinformatics tools’ manuals. No custom coding or programming was used.
References
Production area and volume of vegetable in South Korea in 2022, https://kostat.go.kr/anse/ (2022).
Production amount and index of agriculture and forestry, http://www.mafra.go.kr (2022).
Kim, D.-R., Gang, G.-h, Cho, H.-j, Yoon, H.-S. & Myoung, I. S. Disease Severity of Angular Leaf Spot Disease by Different Inoculation Method and Eco-Friendly Control Efficacy in Strawberry. The Korean Journal of Pesticide Science 20, 35–40 (2016).
Outlook of agriculture 2024 https://www.krei.re.kr/krei/index.do (2024).
Kim, D.-Y. et al. Changes in growth and yield of strawberry (cv. Maehyang and Seolhyang) in response to defoliation during nursery period. Journal of Bio-Environment Control 20, 283–289 (2011).
Jeong, H. J., Choi, H. G., Moon, B. Y., Cheong, J. W. & Kang, N. J. Comparative analysis of the fruit characteristics of four strawberry cultivars commonly grown in South Korea. Horticultural Science & Technology 34, 396–404 (2016).
Choi, J.-M., Latigui, A. & Yoon, M.-K. Growth and nutrient uptake of ‘Seolhyang’ strawberry (Fragaria× ananassa Duch) responded to elevated nitrogen concentrations in nutrient solution. Horticultural Science & Technology 28, 777–782 (2010).
Je, H.-J. et al. Development of cleaved amplified polymorphic sequence (CAPS) marker for selecting powdery mildew-resistance line in strawberry (Fragaria× ananassa Duchesne). Horticultural Science & Technology 33, 722–729 (2015).
Dae-Young, K. et al. Evaluation of Anthracnose and Fusarium wilt Reistance of Domestic and Foreign Strawberry Germplasms and Selected Lines. Journal of the Korean Society of International Agriculture 32, 423–430 (2020).
Kim, I. et al. Changes in volatile compounds in short-term high CO2-treated ‘Seolhyang’ strawberry (Fragaria× ananassa) fruit during cold storage. Molecules 27, 6599 (2022).
Jee, E. et al. Analysis of volatile organic compounds in Korean-bred strawberries: insights for improving fruit flavor. Frontiers in Plant Science 15, 1360050 (2024).
Saxena, R. K., Edwards, D. & Varshney, R. K. Structural variations in plant genomes. Briefings in functional genomics 13, 296–307 (2014).
Chen, Y. H., Gols, R. & Benrey, B. Crop domestication and its impact on naturally selected trophic interactions. Annual Review of Entomology 60, 35–58 (2015).
Edwards, D. & Batley, J. Plant genome sequencing: applications for crop improvement. Plant biotechnology journal 8, 2–9 (2010).
Lang, D. et al. Comparison of the two up-to-date sequencing technologies for genome assembly: HiFi reads of Pacific Biosciences Sequel II system and ultralong reads of Oxford Nanopore. Gigascience 9, giaa123 (2020).
Loit, K. et al. Relative performance of MinION (Oxford Nanopore Technologies) versus Sequel (Pacific Biosciences) third-generation sequencing instruments in identification of agricultural and forest fungal pathogens. Applied and Environmental Microbiology 85, e01368–01319 (2019).
Hon, T. et al. Highly accurate long-read HiFi sequencing data for five complex genomes. Scientific data 7, 399 (2020).
Li, H. & Durbin, R. Genome assembly in the telomere-to-telomere era. Nature Reviews Genetics, 1-13 (2024).
Han, H. et al. Telomere-to-telomere and haplotype-phased genome assemblies of the heterozygous octoploid ‘Florida Brilliance’strawberry (Fragaria× ananassa). BioRxiv, 2022.2010. 2005.509768 (2022).
Song, Y. et al. Phased gap-free genome assembly of octoploid cultivated strawberry illustrates the genetic and epigenetic divergence among subgenomes. Horticulture research 11, uhad252 (2024).
Zhou, Y. et al. The telomere-to-telomere genome of Fragaria vesca reveals the genomic evolution of Fragaria and the origin of cultivated octoploid strawberry. Horticulture Research 10, uhad027 (2023).
Liu, T., Li, M., Liu, Z., Ai, X. & Li, Y. Reannotation of the cultivated strawberry genome and establishment of a strawberry genome database. Horticulture research 8 (2021).
Mao, J. et al. High-quality haplotype-resolved genome assembly of cultivated octoploid strawberry. Horticulture Research 10, uhad002 (2023).
Cheng, H. et al. Haplotype-resolved assembly of diploid genomes without parental data. Nature Biotechnology 40, 1332–1335 (2022).
Alonge, M. et al. RaGOO: fast and accurate reference-guided scaffolding of draft genomes. Genome biology 20, 1–17 (2019).
Gurevich, A., Saveliev, V., Vyahhi, N. & Tesler, G. QUAST: quality assessment tool for genome assemblies. Bioinformatics 29, 1072–1075 (2013).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome biology 21, 1–27 (2020).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic acids research 46, e126–e126 (2018).
Ou, S. & Jiang, N. LTR_retriever: a highly accurate and sensitive program for identification of long terminal repeat retrotransposons. Plant physiology 176, 1410–1422 (2018).
Ou, S. et al. Benchmarking transposable element annotation methods for creation of a streamlined, comprehensive pipeline. Genome biology 20, 1–18 (2019).
Humann, J. L., Lee, T., Ficklin, S. & Main, D. Structural and functional annotation of eukaryotic genomes with GenSAS. Gene prediction: methods and protocols, 29-51 (2019).
Mapleson, D., Venturini, L., Kaithakottil, G. & Swarbreck, D. Efficient and accurate detection of splice junctions from RNA-seq with Portcullis. GigaScience 7, giy131 (2018).
Camacho, C. et al. BLAST+: architecture and applications. BMC bioinformatics 10, 1–9 (2009).
Boutet, E., Lieberherr, D., Tognolli, M., Schneider, M. & Bairoch, A. in Plant bioinformatics: methods and protocols 89-112 (Springer, 2007).
Shulaev, V. et al. The genome of woodland strawberry (Fragaria vesca). Nature genetics 43, 109–116 (2011).
Hardigan, M. A. et al. Blueprint for phasing and assembling the genomes of heterozygous polyploids: application to the octoploid genome of strawberry. BioRxiv (2021).
Cabanettes, F. & Klopp, C. D-GENIES: dot plot large genomes in an interactive, efficient and simple way. PeerJ 6, e4958 (2018).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Goel, M., Sun, H., Jiao, W.-B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome biology 20, 1–13 (2019).
Bateman, A. et al. The Pfam protein families database. Nucleic acids research 32, D138–D141 (2004).
Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME suite. Nucleic acids research 43, W39–W49 (2015).
Chen, C. et al. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Molecular plant 16, 1733–1742 (2023).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC bioinformatics 5, 1–19 (2004).
Wang, Y. et al. MCScanX: a toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic acids research 40, e49–e49 (2012).
Han, H. D. PacBio HiFi reads for genome assembly of ‘Seolhyang’ strawberry. NCBI Sequence Read Archive. https://identifiers.org/ncbi/insdc.sra:SRP527089 (2024).
Han, H. D. Chromosome-level genome assembly of ‘Seolhyang’ strawberry. NCBI GenBank. https://identifiers.org/ncbi/insdc.gca:JBKFVU000000000.1 (2024).
Han, H. D. Gene annotation and supplementary datasets for the genome assembly of cultivated strawberry ‘Seolhyang’ (Fragaria × ananassa). FigShare https://doi.org/10.6084/m9.figshare.26866807 (2024).
Acknowledgements
This work was supported by the Rural Development Administration of Korea (RS-2023-00225421) and National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (RS-2024-00355164)
Author information
Authors and Affiliations
Contributions
Author Contributions Y.O. and H.H. conceived and designed the experiments. K.H., H.P., and Y.O. prepared the materials. H.H. performed the bioinformatics analysis and prepared the results. Y.O., H.H., Y.J., and S.L. wrote the manuscript. Y.O. and S.L. edited and improved the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Han, H., Jang, Y.J., Han, K. et al. Chromosome-level genome assembly of cultivated strawberry ‘Seolhyang’ (Fragaria × ananassa). Sci Data 12, 1002 (2025). https://doi.org/10.1038/s41597-025-05191-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05191-6