Abstract
Machine learning classification algorithms have emerged as promising tools to support the early detection of skin cancers. Existing algorithms typically assess malignancy of skin lesions based on a single skin image. This is in contrast with how clinicians integrate information from their physical examination, comparing multiple skin lesions of an individual and changes in lesions over time. Including contextual information could greatly enhance machine learning algorithms. However, contextual information in skin image datasets is predominantly scarce and inconsistent. Additionally, a dataset containing images of the same lesion across multiple time points and varying resolutions is also lacking. To address these gaps, we present a comprehensive dataset derived from skin monitoring of 480 study participants recruited from a general population sample (n = 196) and a high-risk for melanoma cohort (n = 284). This dataset includes images of 250,162 skin lesions obtained from three-dimensional total body imaging (tile images), along with corresponding dermoscopic images of 9,389 lesions. For 340 of the participants, longitudinal tile and dermoscopic images (ranging from 2 to 7) are provided.
Similar content being viewed by others
Background & Summary
Skin cancers emerge when there is a genetic mutation of skin cells resulting in uncontrolled proliferation, often in response to ultraviolet radiation1,2. Cutaneous melanoma (called melanoma from hereon) is the deadliest form of skin cancer responsible for 80% of skin cancer-related deaths. Basal cell carcinomas (BCCs) and squamous cell carcinomas (SCCs) are very common skin cancers, but less deadly than melanoma3. Although melanoma is relatively rare, its global incidence has increased over the past 50 years4,5. Some of this growth in incidence may have resulted from increased diagnosis scrutiny6. While late-stage skin cancers have a higher risk of mortality and require complex and costly treatment, early-stage diagnosis is associated with excellent survival and lower treatment costs7,8. Due to commonly being located on the skin surface, skin cancers are often visible and potentially detectable by looking at the skin9. Because of this visibility, various skin imaging technologies have been proposed to enhance early detection10. Machine learning algorithms, particularly neural networks, have shown potential in classifying skin images into benign and malignant lesions11. Most algorithms developed to date have only been trained on labelled dermoscopic (magnified) images11,12,13,14.
Dermoscopic images are high-resolution, magnified images of skin lesions that allow clinicians to view deeper skin structures by reducing skin surface reflectivity15. These images are valuable in the differential diagnosis of melanoma and other skin cancers; however, their interpretation, even by trained clinicians is time-consuming, and results are highly dependent on the clinician’s experience16. Although machine learning has great potential in classifying these skin images, training algorithms require a large number of accurately annotated images17. The underlying training dataset plays a crucial role in the accuracy, generalisability, and clinical usefulness of algorithms18. Several dermoscopic datasets have been compiled for the training and evaluation of neural networks19. Dermatological atlases intended for educational purposes, have also been used as an input for algorithm development19.
Despite the large number of images available in currently existing dermoscopic datasets, they are likely biased toward more notable lesions that led to the dermoscopic image being taken20. Additionally, these datasets are mostly limited to isolated skin images, leading algorithms trained on them to base their decision-making about the malignancy of a lesion solely on a single skin image. This contrasts with how clinicians make decisions by integrating information from anamnesis, physical examination, evaluating whether the lesion in question resembles other lesions on the same patient, and sometimes changes in the lesion over time.
One limitation of dermoscopic datasets has been the lack of information regarding the overall lesion phenotype of an individual20. To overcome this, the International Skin Imaging Collaboration (ISIC) 2020 provided a dataset that includes dermoscopic images from multiple lesions of the same person21. This dataset was also used to support the development of algorithms based on the “ugly duckling” concept. This concept suggests that benign moles of a person often share similarities in pattern, shape, colour, and size, while melanoma is more likely to stand out. Although this dataset enabled comparison of multiple lesions from the same individual, it remained biased toward more atypical lesions selected for dermoscopic imaging20. To minimize selection bias and provide a more comprehensive representation of lesion phenotypes, the ISIC 2024 offered skin images with smartphone-compatible resolution obtained from three-dimensional total body photographs (3D-TBP) of participants20.
Despite recent advancements in ISIC and other datasets, over-representation of atypical lesions in dermoscopic datasets remains a limitation. Moreover, datasets that contain images of the same lesion in both dermoscopic and smart-phone resolution remain limited. Skin image datasets also have limitations with regards to the available metadata. For example, despite the clinical importance of ethnicity and Fitzpatrick skin type, “Patient ethnicity data were available for 1415 images (1.3% of all images), and Fitzpatrick skin type data for 2236 (2.1%)” page 6919.
Evidence has shown that including such metadata can significantly increase the accuracy of machine learning algorithms22,23. Metadata also provides valuable information about the characteristics of the populations used to train and validate algorithms. This information is important because, while machine learning algorithms typically perform at a satisfactory level when tested on skin images from the same population used for training, they often underperform when evaluated on data from different populations24,25,26. Metadata information provides transparency to assess the robustness of results beyond the original population.
Lack of metadata and inconsistencies in collection and reporting may stem from a lack of consensus regarding which metadata is essential19. Having a dataset with comprehensive metadata provides an opportunity to identify the minimal metadata that is critical to collect in order to increase algorithm accuracy and aide generalisability evaluation.
One of the key indicators of a potential melanoma is the change in size, colour, shape, or elevation of a lesion over time27. Advancements in imaging systems and machine learning algorithms now make it possible to detect and monitor almost all skin lesions in individuals over time. These longitudinal skin images are particularly valuable for detecting early signs of malignant transformation and developing algorithms for tracking changes based on longitudinal data.
To overcome the limitations of previous datasets and provide a resource that includes more clinical information, we present a dataset from skin monitoring of 480 participants across two longitudinal studies on general (n = 196) and high-risk populations (n = 284). The dataset includes low-resolution tile images of detected pigmented lesions extracted from 3D-TBP (on average, 521 lesions per person), and corresponding high-resolution dermoscopic images for lesions that were larger than 5 mm or were of interest by either the participant or clinician (on average, 20 lesions per person). This dataset overall includes tile images for 250,162 skin lesions (including 28 melanomas) along with corresponding dermoscopic images for 9,389 of these skin lesions (including 19 melanomas). Longitudinal tile and dermoscopic images (ranging from 2 to 7 time points) are available for 340 participants. This skin image dataset is accompanied by comprehensive individual-level metadata on lesion anatomic location, individuals’ number of naevi, demographic information, skin cancer history, freckling, skin colour, as well as sun exposure and sun protection behaviour data.
Methods
General
The data presented in this manuscript were derived from two longitudinal studies conducted by the Dermatology Research Centre at the University of Queensland. The studies are titled “Mind your Moles”, and “Health Outcome Program Study”.
Mind your moles (MYM)
The first study, “Mind Your Moles” (MYM study), enrolled people from the general population of adults living in Southeast Queensland, Australia, with details of the study reported previously28. Participants were recruited from the Australian Electoral Roll. Eligibility criteria included having at least one naevus and being willing to attend 3D-TBP every six months for a period of three years. This study received ethics approval from the Human Research Ethics Committee of Metro South Health (HREC/16/QPAH/816), the University of Queensland (2016000554), and the Queensland University of Technology (1600000515). A total of 196 participants consented to sharing their images for future research.
Health outcomes program study (HOPS)
The second study, “Health Outcomes Program Study” (HOPS study), was a randomised controlled trial (RCT) and enrolled adults at high risk of melanoma living in Southeast Queensland, Australia, with the study protocol reported previously29. Participants were recruited by referral from dermatologists and medical practitioners or through the University of Queensland Dermatology Research Centre’s registry of research volunteers. A total of 284 participants filled out the data-sharing consent form. Eligibility criteria included being diagnosed with at least one melanoma before the age of 40 years, or two or more melanomas before the age of 65 years, or having a strong family history, or dysplastic naevus phenotype. Eligible participants were randomly assigned to one of two groups: the intervention group, which continued their usual follow-up with their regular doctor and underwent longitudinal 3D-TBP along with longitudinal dermoscopy imaging every six months for two years, or the control group, which continued their usual follow up with their regular doctor and received 3D-TBP and dermoscopy imaging only once at their last study visit. This study received ethics approval from the Human Research Ethics Committee of Metro South Health (HREC/17/QPAH/816) and The University of Queensland (2018000074).
Data collection through sequential visits
Participants of the MYM study and the intervention group of the HOPS study were followed up with imaging sessions every six months for three or two years, respectively (Fig. 1). Visits included 3D-TBP, dermoscopy imaging, clinical skin examination, and questionnaire completion. The control group of the HOPS study was followed with 6-monthly questionnaires only and had one complementary 3D-TBP and dermoscopy imaging at the 24-month timepoint after completing their last study questionnaire. An overview of the most important information collected and reported for each visit is presented in Figs. 1, 2.
Follow-up schedule for MYM and HOPS participants.
Summary of the data collected and presented in the dataset. * Naevus counts were obtained using the naevus detection algorithm inbuilt into the 3D imaging software30.
Skin monitoring
Sequential 3D-TBP
TBP was performed using a VECTRA Whole Body 360 (Canfield Scientific Inc., Parsippany-Troy Hills, NJ, USA). The VECTRA consists of a framework of 92 cameras that collect images simultaneously from different angles and uses software to combine them into a 3D avatar. The VECTRA software includes a Convolutional Neural Network (CNN) that detects pigmented skin lesions30. For each pigmented lesion identified through 3D-TBP, corresponding lesion images (tile images) were extracted and included in the dataset. Not all the tile images underwent manual validation. Further details regarding the accuracy of the lesion detection algorithm are provided in the Technical Validation section.
An overview of the available skin image data including images extracted from 3D avatar and their corresponding dermoscopic image between different demographics, clinical groups, and anatomical locations, is provided in Table 1.
Sequential dermoscopic images
Dermoscopic images were taken of pigmented lesions with a diameter of 5 mm or greater and other lesions that were either of concern for the participant or the clinician/melanographer. Dermoscopic images were captured using either the VEOS SLR Dermoscopic Camera or the Canon EOS Rebel T6i. Details about the specific camera used for each image are included in their metadata. The number of dermoscopic images across different population groups are presented in Table 1.
Questionnaire data
At each study visit, a clinical research assistant administered the questionnaire. The baseline questionnaire included questions on demographics, socioeconomic status, sun behaviour, and skin cancer history. Questions about sun behaviour were repeated during subsequent visits, as shown in Figs. 1, 2.
Many additional questions, particularly from the high-risk population were collected. These questions were mainly about the frequency of skin checks, quality of life, opinion about melanoma fatality, and attitude towards using 3D imaging. Since this data was not relevant for algorithm development and to minimize confusion, they were excluded from the shared dataset. However, these data are available upon request from the corresponding author or research committee. A complete list of all questions asked through the questionnaire can be seen in the Questionnaires_and_clinical_assessment_data.pdf file within the dataset.
Clinical data
A clinical skin examination was performed by a medical professional or trained melanographer and documented on a standard form. The information collected included eye colour, hair colour, innate skin colour, facultative skin colour, freckling score, and spectrophotometry of skin colour.
Data Records
The dataset has been made permanently accessible for public download through UQ eSpace at https://doi.org/10.48610/a13deaf31. It includes tile images extracted from 3D-TBP images for 250,162 skin lesions over the study period, with an average of 521 skin lesions per participant. Dermoscopic images are available for 9,389 of these lesions, corresponding to an average of 20 lesions with dermoscopic imaging per participant. Additionally, longitudinal dermoscopic images are available for 7,038 of these lesions, totalling 35,909 dermoscopic images in the dataset. Histopathologic results are provided for 1,267 of these lesions, including 30 melanomas, 80 basal cell carcinomas, and 48 squamous cell carcinomas. Lesions without histopathology results can be considered benign as clinically they were not identified as needing further examination or excision.
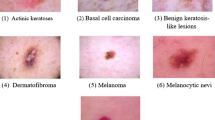
Metadata includes anatomical location of lesions and participant’s characteristics including age group, gender, eye colour, hair colour, skin colour, freckling score, sun exposure, sunburn history, ancestry, number of naevi, skin cancer history, and family history of melanoma. An overview of the skin image dataset, including number of clinical images and their corresponding dermoscopic image across different demographic and clinical groups, and anatomical locations, is presented in Table 1. Similarly, Fig. 3 provides an overview of the distribution of skin lesions across various diagnosis categories.
The number of skin lesions available across diagnosis categories, with one example for each category for illustration. * For 11 melanomas only the tile images are available.
Dataset format
Clinical and dermoscopic images are in Portable Network Graphics (PNG) format and the link between tile and dermoscopic images along with their metadata is provided in a linked comma-separated values (CSV) file.
Dermoscopic images are stored in a folder and each of them has a unique ID. Information about their anatomical location, diagnosis, and camera type used for image capture is provided in a CSV file, as shown in Table 2.
Tile images of all lesions detected by lesion detection algorithms for each participant are stored in folders labelled according to the participant and visit number. Information on the diagnosis category, anatomical location of the lesion, corresponding dermoscopic image, and tile image ID for each lesion for future visits is provided in a CSV file, as shown in Table 3.
Other participant characteristics including demographics and risk factors are presented in a separate CSV file, as shown in Table 4. All participant’s data has been deidentified with a random ID assigned to each case.
Technical Validation
Tile images were extracted from all lesions detected by the inbuilt VECTRA CNN on the participant’s 3D-TBP. This CNN had been developed and tested by our team, and demonstrated a sensitivity of 79% and a specificity of 91% in detecting naevi larger than 2 mm when assessed prospectively30. From the lesions that were thought to be suspicious, dermoscopic images were taken and when deemed necessary, lesions were referred for excision. A histopathology report was collected for excised lesions. Overall, histopathologic results were obtained for 1,267 lesions. Non-biopsied lesions were followed up in subsequent visits and deemed benign if they did not show any malignant changes.
Usage Note
The main capabilities of the presented dataset, along with important considerations, are summarized below.
-
Data on benign lesions: Although the number of histopathologically verified melanomas in our dataset is limited, our dataset includes a substantial number of tile and dermoscopic images of benign lesions from both general and high-risk population participants. This can supplement existing dermoscopic datasets to overcome the overrepresentation of suspicious lesions and help develop more accurate melanoma detection algorithms.
-
Information on overall lesion phenotype: We have provided data on multiple skin lesions for the same participant, including tile images of all lesions of participants and dermoscopic images from multiple lesions of the same individual. This data can be used to develop algorithms based on comparing multiple lesions from the same person allowing to understand what a typical pigmented lesion for a certain person looks like.
-
Metadata: Our dataset includes comprehensive metadata alongside skin images, enabling the identification of key metadata items that enhance the accuracy of machine learning algorithms. Moreover, given that machine learning algorithms generally perform better on populations similar to those used for training, the availability of detailed metadata allows for an evaluation of the model’s generalizability and its potential reliability for each individual based on their characteristics.
-
Tile images and corresponding dermoscopic image: Our dataset includes images of the same lesion in both dermoscopic and clinical quality.
-
Longitudinal skin images: The time series data can be used to develop algorithms on longitudinal data and check the practicality of skin cancer early detection.
-
Overlap: 9.9% of our tile images and 29.7% of our dermoscopic images overlap with the images in ISIC 2024 and ISIC 2020 datasets. However, it is important to retain these images because in this dataset they now form part of sequencing imaging and are linked to our dermoscopic and clinical data.
Limitations and further study
Numerous datasets are available for training and evaluating machine learning algorithms, but a dataset that provide longitudinal data is lacking. Additionally, existing datasets, particularly dermoscopic images, have limitations such as overrepresentation of suspicious lesions and a lack of lesion phenotype information. Similarly, datasets that include comprehensive metadata or images of lesions at various resolutions is also scare. To fill these major gaps and to better reflect clinical reality, here we provide a new dataset. Even though this dataset has several strengths, it also has some limitations, particularly that it contains only a small number of histopathologically verified melanomas, and also lacks diversity in ethnicity among study participants who predominantly had white skin colour with Northern European ancestry. Further studies that provide longitudinal skin image datasets with a larger number of melanoma cases and greater ethnic diversity are strongly recommended to improve generalizability and diagnostic accuracy of algorithms.
Code availability
Custom codes generated for extracting lesion images are available at https://github.com/Nimaghahari/Longitudinal_skin_images.
References
Matthews, N. H., Li, W.-Q., Qureshi, A. A., Weinstock, M. A. & Cho, E. Epidemiology of melanoma. Exon Publications, 3–22 (2017).
Narayanan, D. L., Saladi, R. N. & Fox, J. L. Ultraviolet radiation and skin cancer. International journal of dermatology. 49, 978–986 (2010).
Urban, K., Mehrmal, S., Uppal, P., Giesey, R. L. & Delost, G. R. The global burden of skin cancer: A longitudinal analysis from the Global Burden of Disease Study, 1990–2017. JAAD international. 2, 98–108 (2021).
Erdmann, F. et al. International trends in the incidence of malignant melanoma 1953–2008—are recent generations at higher or lower risk? International journal of cancer. 132, 385–400 (2013).
Garbe, C. & Leiter, U. Melanoma epidemiology and trends. Clinics in dermatology. 27, 3–9 (2009).
De Gruijl, F. R. & Armstrong, B. K. Cutaneous Melanoma: Sheep in Wolves Clothing? Anticancer Research. 42, 5021–5025 (2022).
Rigel, D. S. & Carucci, J. A. Malignant melanoma: prevention, early detection, and treatment in the 21st century. CA: a cancer journal for clinicians. 50, 215–236 (2000).
Morton, D. L., Davtyan, D. G., Wanek, L. A., Foshag, L. J. & Cochran, A. J. Multivariate analysis of the relationship between survival and the microstage of primary melanoma by Clark level and Breslow thickness. Cancer. 71, 3737–3743 (1993).
Rigel, D. S. et al. Importance of complete cutaneous examination for the detection of malignant melanoma. Journal of the American Academy of Dermatology. 14, 857–860 (1986).
Kudrin, K. et al. Early diagnosis of skin melanoma using several imaging systems. Optics and Spectroscopy. 128, 824–834 (2020).
Esteva, A. et al. Dermatologist-level classification of skin cancer with deep neural networks. nature. 542, 115–118 (2017).
Tschandl, P., Rosendahl, C. & Kittler, H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Scientific data. 5, 1–9. Harvard Dataverse https://dataverse.harvard.edu/dataset.xhtml?persistentId=doi:10.7910/DVN/DBW86T (2018).
Brinker, T. J. et al. Deep learning outperformed 136 of 157 dermatologists in a head-to-head dermoscopic melanoma image classification task. European Journal of Cancer. 113, 47–54 (2019).
Haenssle, H. A. et al. Man against machine: diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Annals of oncology. 29, 1836–1842 (2018).
Braun, R. P., Rabinovitz, H. S., Oliviero, M., Kopf, A. W. & Saurat, J.-H. Dermoscopy of pigmented skin lesions. Journal of the American Academy of Dermatology. 52, 109–121 (2005).
Eltayef, K., Li, Y. & Liu, X. Detection of Melanoma Skin Cancer in Dermoscopy Images. Journal of Physics: Conference Series. 12034 (2016).
Deng, J., et al. ImageNet: A Large-Scale Hierarchical Image Database. IEEE conference on computer vision and pattern recognition. 248–255 (2009).
Ayan, E. & Ünver, H. M. Skin cancer diagnosis using convolutional neural networks for smartphone images: A comparative study. Journal of Radiation Research and Applied Sciences. 15, 262–267 (2018).
Wen, D. et al. Characteristics of publicly available skin cancer image datasets: a systematic review. The Lancet Digital Health. 4, e64–e74 (2022).
Kurtansky, N. R. et al. The SLICE-3D dataset: 400,000 skin lesion image crops extracted from 3D TBP for skin cancer detection. Scientific Data. 11, 884. Kaggle https://www.kaggle.com/competitions/isic-2024-challenge (2024).
Rotemberg, V. et al. A patient-centric dataset of images and metadata for identifying melanomas using clinical context. Scientific data. 8, 34. Kaggle https://www.kaggle.com/competitions/siim-isic-melanoma-classification (2021).
Yap, J., Yolland, W. & Tschandl, P. Multimodal skin lesion classification using deep learning. Experimental dermatology. 27, 1261–1267 (2018).
Pacheco, A. G. & Krohling, R. A. The impact of patient clinical information on automated skin cancer detection. Computers in biology and medicine. 116, 103545 (2020).
Codella, N. et al. Skin lesion analysis toward melanoma detection 2018: A challenge hosted by the international skin imaging collaboration (isic). arXiv preprint arXiv:1902.03368 (2019).
Navarrete-Dechent, C. et al. Automated dermatological diagnosis: hype or reality? The Journal of investigative dermatology. 138, 2277–2279 (2018).
Dick, V., Sinz, C., Mittlböck, M., Kittler, H. & Tschandl, P. Accuracy of computer-aided diagnosis of melanoma: a meta-analysis. JAMA dermatology. 155, 1291–1299 (2019).
Balch, C. M. et al. Final version of the American Joint Committee on Cancer staging system for cutaneous melanoma. Journal of Clinical Oncology. 19, 3635–3648 (2001).
Koh, U. et al. ‘Mind your Moles’ study: protocol of a prospective cohort study of melanocytic naevi. BMJ open. 8, e025857 (2018).
Primiero, C. A. et al. Evaluation of the efficacy of 3D total-body photography with sequential digital dermoscopy in a high-risk melanoma cohort: protocol for a randomised controlled trial. BMJ open. 9, e032969 (2019).
Betz-Stablein, B. et al. Reproducible naevus counts using 3D total body photography and convolutional neural networks. Dermatology. 238, 4–11 (2022).
Ghahari, N. et al. A longitudinal dataset of tile and corresponding dermoscopic images with metadata for identifying skin cancers, UQ eSpace, https://doi.org/10.48610/a13deaf (2025).
Acknowledgements
The studies from which our data were obtained were supported by the National Health and Medical Research Council (NHMRC 2006551, 2009923, 2034422, 1153046). We acknowledge the clinical staff for their assistance in photographing patients using the VECTRA 360 and capturing dermoscopic images, as well as the patients who contributed images to the dataset.
Author information
Authors and Affiliations
Contributions
All authors contributed to the manuscript preparation and provided feedback during the revision process. N.G. co-wrote the manuscript with M.J.; N.G. and A.M. extracted images from the dataset and linked them longitudinally. L.C., B.B.S., A.M., D.J., C.P., S.S.C., J.T., H.P.S. Contributed to the design of manuscript and dataset and all authors approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
H.P.S is shareholder of e-derm consult GmbH and MoleMap by Dermatologists Pty Ltd. He provides teledermatological reports regularly for both companies. H.P.S also consults for Canfield Scientific Inc and is an adviser of First Derm.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ghahari, N., Caffery, L., Betz-Stablein, B. et al. A longitudinal dataset of tile and corresponding dermoscopic images with metadata for identifying skin cancers. Sci Data 12, 1602 (2025). https://doi.org/10.1038/s41597-025-05880-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-05880-2