Abstract
Bone marrow smear cytomorphology serves as the cornerstone for diagnosing hematological malignancies. It is particularly complex in myelodysplastic syndrome (MDS), which brings challenges to automated image analysis approaches. This dataset provides 25,009 images of the bone marrow cells from 125 patients with MDS. Image tags consist of 27 categories, including normal cells and abnormal cells in MDS that are characterized by pathological hematopoiesis, such as micro megakaryocytes. Each label underwent rigorous review by up to three independent experts. Cells were cropped after converting their bounding boxes into squares based on the longest side and expansion by 10% to guarantee the integrity of the cell boundaries and morphological features in the cropped images while minimum surrounding cells were included. Through training using this database, an AI model could attain the ability to differentiate dysplastic cells.
Similar content being viewed by others
Background & Summary
The evaluation of bone marrow smear cell morphology is the basis for the diagnosis of blood diseases1. It is particularly important in the case of MDS2,3, where pathological hematopoiesis causes cell morphology to be variable and complex4,5,6. Moreover, the test requires two pathologists to manually differentiate and count at least 500 nucleated cells on Wright-stained bone marrow smears7, which is labor-intensive and time-consuming8,9.
In the past decade automatic morphology analyzers have been used in laboratory work10,11,12. Although development in machine learning has successfully improved the accuracy and efficiency of the performance of these analyzers13, its improvement largely depends on the availability of large and high-quality datasets14,15,16. However, most studies have focused on peripheral blood cells, and the datasets of bone marrow cells are rare17,18,19,20. Given the variety of morphological features and numerousness of bone marrow cell types, some of which have low occurrence, it is hard to obtain a sufficient number of cells for effective algorithm training21,22. In the process of cell segmentation, bone marrow smears may be more difficult than peripheral blood smears owing to cell overlap23. Some studies have reported the complexity of unsatisfactory performance in real-world data using small datasets24. Therefore, the establishment of disease-specific bone marrow cell datasets will benefit the development of AI for cell morphology applications.
The dataset contains 25,009 bone marrow cell images from 125 patients with MDS. The image labels consisted of 32 categories, including normal cells and pathological hematopoietic cells, such as small megakaryocytes and micromegakaryocytes. Cell labels were reviewed by 3 experts. This dataset provides images under different shooting conditions, making it an ideal dataset for testing the robustness of machine learning models. It is particularly suitable for in-depth analysis of cell morphology evaluation in MDS and similar hematological diseases. The cell images are wider than actual cell margins by 10% to ensure cell integrity and to support random cropping in the data augmentation process before training. The publication of this dataset aims to address the limitations of existing datasets and provide more comprehensive data for the application of machine learning models in the evaluation of bone marrow cell morphology in hematological diseases, especially in the diagnosis of MDS.
Methods
The first-time bone marrow smears of 125 patients diagnosed with MDS from the First Affiliated Hospital of Zhejiang University Medical College were collected. (Institutional review board approval number No. 2024-0889) As the bone marrow smear does not contain identifiable personal information, it is exempted from obtaining written informed consent based on this situation. Among the 125 patients with MDS, 69 were male and 56 were female, aged 23–82 years, with a median age of 64 years. MDS was diagnosed based on the World Health Organization (WHO) 2022 classification criteria for hematopoietic and lymphoid neoplasms. Among them, 44 patients were MDS-LB subtype, 46 were MDS-IB1, 31 were MDS-IB2, 3 were MDS-h and 1 was MDS-f.
Sample preparation
Bone marrow smears were prepared manually by the wedge method. A BaSO BSZ-CM16-B automatic slide staining machine with BaSO Wright-Giemsa staining solution (solution a) and BaSO phosphate buffer (ph6.8) (solution b) was used to complete Wright-Giemsa staining of the smears.
Data acquisition
Slides were processed using MorphoInsight, a versatile and automated slide scanning analysis system developed by Kinsight, Co., Ltd., and successor of MorphoGo25,26,27. The system is equipped with slide loading/unloading mechanics, a moving gripper that carries the slide while scanning, and a high-speed camera. Its optical components include a 100X oil objective lens and an LED light source. For each slide, a whole-slide image (WSI) with a pixel distance of 75 nm was generated. Subsequently, the cell detection algorithm of the system obtained cell locations on the WSI as bounding boxes. Medical technologists manually selected the region of interest (ROI) for each slide. After converting the bounding boxes to squares according to longest side and expanding them by 10%, cells in the ROI were cut from the WSI. This step ensured the integrity of cell borders and morphological features within the cut images while minimum surrounding cells were included.
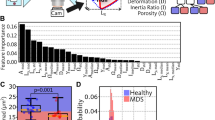
Additional images were collected during routine work using digital cameras (either SmartV550Df, SmartV650Df or SmartV650D) mounted on a microscope with 100X oil objective lens. The light source of the microscope is either a halogen lamp or an LED bulb. These images were processed with similar manner. Cells were located by algorithm, and their locations were confirmed by medical technologists. The bounding boxes were also converted to squares and expanded by 10% before the cells were cut from the images. The entire data-acquisition process is illustrated in Fig. 1.
Data acquisition process diagram.
Data annotations
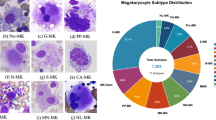
First, a medical technician with approximately eight years of work experience at hospital identified the subtypes of bone marrow nucleated cells. Subsequently, these annotations were reviewed by a medical expert with approximately 20 years of experience. In cases of inconsistency, the cells were rechecked by an expert with 25 years of experience. If no consensus was reached, the image was excluded from the dataset. If the staining quality was considered too poor or the cell segmentation was not suitable for identification, the image was excluded as well. Cell images were classified into 32 categories, including normal bone marrow cells and abnormal cells caused by pathological hematopoiesis in MDS, including blasts, dysplastic erythroblasts, dysplastic granulocytes, small megakaryocytes, and micromegakaryocytes (Fig. 2). These cells are commonly observed and are characteristic of MDS, representing the cell types that pathologists frequently utilize in their routine work for cell classification. Each MDS patient has different dysplastic cell manifestations, and not every type of dysplasia may be present in one patient28. Considering the relatively low occurrence of dysplastic cells, if further subdivided, the number of each cell type will be too small to meet the needs of AI training. And some pathological types are difficult to distinguish, but they can be distinguished from normal cells. Therefore, we consider classifying them by the 3 main myeloid series, grouping pathological anomalies of each series together. We expect classification systems trained with this dataset could inform the operator of abnormalities.
Some annotated bone marrow cells in the database (a–e) blast cells, (f) ring-nucleus neutrophil, (g) binucleated granulocyte, (h) granulocyte with nuclear-cytoplasmic maturation asynchrony, (i,j) neutrophil with Pseudo-Pelger-Huët-like anomaly, (k) erythroblast with nuclear budding, (l) erythroblast with basophilic stippling, (m) trinucleated erythroblast, (n) erythroblast with Howell-Jolly bodies, (o) dysplastic erythroblast, (p–t) micromegakaryocytes, (u–y) small megakaryocytes.
Data Records
The root directory contains two folders. Cell images acquired through MorphoInsight are stored in ‘main’ folder, and additional cell images in ‘add’ folder. In each folder, images are grouped according to cell classification. Image names are formatted using cell classification and an index. Each folder contains an .csv file listing the image paths and corresponding information. For the .csv file in ‘main’ folder, cell classification and diameter are provided. And the .csv file in ‘add’ folder also contains camera resolution setting used to capture the original image.
Technical Validation
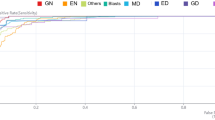
Images acquired through MorphoInsight were randomly split into training, validation, and test sets at ratios of 70%(15987 images), 10%(2352 images), and 20%(4503 images), respectively. Transfer learning using MorphoInsight’s cell classification model, which specializes in normal bone marrow cell classification as the base model, yielded an overall accuracy(number of cells whose model prediction was consistent with expert annotation, divided by total cell number) of 80.13%. During the training, small megakaryocytes, micromegakaryocytes, dysplastic granulocytes, and dysplastic erythroblasts were added as new types. The f1 scores (calculated as harmonic mean of recall and precision) of these four types of dysplastic cells were 50.85%, 64.94%, 31.25%, and 59.31% respectively. Their recall and precision rates are presented in Table 1. Some confusion between the dysplastic cells and their corresponding normal types was observed in the results, which could be explained by the morphological similarities between them. Figure 3. The base model cannot differentiate dysplastic granulocytes and erythroblasts from normal ones. And it predicted small and micro- megakaryocytes as mostly lymphocytes and smudge cells.
Classification results of the trained model by means of the confusion matrix. Rows represent the annotated cell types and the columns the predicted type given by the classifier. Diagonal values (green cells) are the true positive number for each class. Out of diagonal values give the numbers of wrong predictions.
The test demonstrated that an acceptable classification performance of dysplastic cells can be achieved using our dataset. Better results can be expected with more cell images. In fact, the dataset offered here forms only a small portion of the data collected, as each WSI could contain thousands of cells. But the remaining parts have not been thoroughly annotated. It can be used for further learning and optimization of MorphoInsight’s cell classification model.
Usage Notes
The dataset consists of JPEG images and information files in CSV format. No special software is required to view and use this dataset.
As a result of their low occurrence, the number of some cell classes in this dataset is small and may be insufficient to achieve acceptable classification results. If desired, classes such as megaloblastic late erythroblast, histiocyte and dysplastic megakaryocyte could be excluded from training set. Classes similar in morphology and clinical significance could be merged. For example, blast NOC, myeloblast, monoblast and promonocyte could all be treated as blast.
Data availability
The dataset is publicly available on the figshare data repository via https://doi.org/10.6084/m9.figshare.28737170.v129.
Code availability
No custom code was used in this study.
Several developing tools of MorphoInsight were used during dataset curation and technical validation, over which Kinsight holds copyright. Thus these tools are not available for open-access.
References
Khoury, J. D. et al. The 5th edition of the World Health Organization Classification of Haematolymphoid Tumours: Myeloid and Histiocytic/ Dendritic Neoplasms. Leukemia 36, 1703–1719, https://doi.org/10.1038/s41375-022-01613-1 (2022).
Hasserjian, R. P. et al. Diagnosis and classification of myelodysplastic syndromes. Blood 142(26), 2247–2257, https://doi.org/10.1182/blood.2023020078 (2023).
Arber, D. A. et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood 127(20), 2391–405, https://doi.org/10.1182/blood-2016-03-643544 (2016).
Weeks, L. D. et al. Prediction of risk for myeloid malignancy in clonal hematopoiesis. NEJM Evid 2(5). https://doi.org/10.1056/evidoa2200310 (2023).
Babcock, S., Calvo, K. R. & Hasserjian, R. P. Pediatric myelodysplastic syndrome. Semin Diagn Pathol 40(3), 152–171, https://doi.org/10.1053/j.semdp.2023.04.006 (2023).
Joffe, E. et al. Kinetics of pre-myelodysplastic syndromes blood values correlate with disease risk and survival. Hematol Oncol 38(5), 782–791, https://doi.org/10.1002/hon.2777 (2020).
Manuel Salto-Tellez, I. A. Cree, Cancer taxonomy: pathology beyond pathology. European Journal of Cancer 115, 57–60, https://doi.org/10.1016/j.ejca.2019.03.026 (2019).
de Swart, L. et al. Cytomorphology review of 100 newly diagnosed lower-risk MDS patients in the European LeukemiaNet MDS (EUMDS) registry reveals a high inter-observer concordance. Ann Hematol 96, 1105–1112, https://doi.org/10.1007/s00277-017-3009-7 (2017).
Mufti, G. J. et al. Diagnostic algorithm for lower-risk myelodysplastic syndromes. Leukemia 32, 1679–1696, https://doi.org/10.1038/s41375-018-0173-2 (2018).
Chari, P. S. & Prasad, S. Pilot Study on the Performance of a New System for Image Based Analysis of Peripheral Blood Smears on Normal Samples. Indian J Hematol Blood Transfus 34, 125–131, https://doi.org/10.1007/s12288-017-0835-7 (2018).
Njoroge, S. W. & Nichols, J. H. Risk Management in the Clinical Laboratory. Annals of Laboratory Medicine 34, 274–278, https://doi.org/10.3343/alm.2014.34.4.274 (2014).
Nam, M. et al. Digital Morphology Analyzer Sysmex DI-60 vs. Manual Counting for White Blood Cell Differentials in Leukopenic Samples: A Comparative Assessment of Risk and Turnaround Time. Annals of Laboratory Medicine 42, 398–405, https://doi.org/10.3343/alm.2022.42.4.398 (2022).
Yoon, S. et al. Performance of digital morphology analyzer Vision Pro on white blood cell differentials. Clinical Chemistry and Laboratory Medicine (CCLM) 59, 1099–1106, https://doi.org/10.1515/cclm-2020-1701 (2021).
Bodzas, A., Kodytek, P. & Zidek, J. A high-resolution large-scale dataset of pathological and normal white blood cells. Scientific Data 10, 466, https://doi.org/10.1038/s41597-023-02378-7 (2023).
Kouzehkanan, Z. M. et al. A large dataset of white blood cells containing cell locations and types, along with segmented nuclei and cytoplasm. Scientific Reports 12, 1123, https://doi.org/10.1038/s41598-021-04426-x (2022).
Park, S. et al. A large multi-focus dataset for white blood cell classification. Sci Data 11, 1106, https://doi.org/10.1038/s41597-024-03938-1 (2024).
Hegde, R. B. et al. Development of a robust algorithm for detection of nuclei of white blood cells in peripheral blood smear images. Multimed Tools Appl 78, 17879–17898, https://doi.org/10.1007/s11042-018-7107-x (2019).
Ruberto, C. D., Loddo, A. & Putzu, L. Detection of red and white blood cells from microscopic blood images using a region proposal approach. Comput. Biol. Med. 116, 103530, https://doi.org/10.1016/j.compbiomed.2019.103530 (2020).
Çınar, A. & Tuncer, S. A. Classification of lymphocytes, monocytes, eosinophils, and neutrophils on white blood cells using hybrid Alexnet-GoogleNet-SVM. SN Appl. Sci. 3, 503, https://doi.org/10.1007/s42452-021-04485-9 (2021).
Boldú, L., Merino, A., Acevedo, A., Molina, A. & Rodellar, J. A deep learning model (ALNet) for the diagnosis of acute leukaemia lineage using peripheral blood cell images. Comput Methods Programs Biomed. 202, 105999, https://doi.org/10.1016/j.cmpb.2021.105999 (2021).
Xi, Z. et al. Interpretation of Chinese expert consensus on the technical and clinical practice specifications of artificial intelligence assisted morphology examination of blood cells. Journal of Clinical Hematology 37(5), 298–300,305, https://doi.org/10.13201/j.issn.1004-2806.2024.05.003 (2024).
liu, G. et al. The clinical application of artificial intelligence assisted blood cell morphological examination should be promoted and expanded. Chinese Journal of Laboratory Medicine 46(3), 231–237, https://doi.org/10.3760/cma.j.cn114452-20230213-00088 (2023).
Ye, G. & Kaya, M. Automated Cell Foreground-Background Segmentation with Phase-Contrast Microscopy Images: An Alternative to Machine Learning Segmentation Methods with Small-Scale Data. Bioengineering (Basel). 9(2), 81, https://doi.org/10.3390/bioengineering9020081 (2022).
Loddo, A. & Putzu, L. On the Effectiveness of Leukocytes Classification Methods in a Real Application Scenario. AI 2, 394–412, https://doi.org/10.3390/ai2030025 (2021).
Jin, H. et al. Developing and Preliminary Validating an Automatic Cell Classification System for Bone Marrow Smears: a Pilot Study. J Med Syst 7 44(10), 184, https://doi.org/10.1007/s10916-020-01654-y (2020).
Wang, X. et al. The Application of Morphogo in the Detection of Megakaryocytes from Bone Marrow Digital Images with Convolutional Neural Networks. Technol Cancer Res Treat. 22, 15330338221150069, https://doi.org/10.1177/15330338221150069 (2023).
Lv, Z. et al. High-accuracy morphological identification of bone marrow cells using deep learning-based Morphogo system. Sci Rep 13, 13364, https://doi.org/10.1038/s41598-023-40424-x (2023).
Germing, U. et al. Evaluation of dysplasia through detailed cytomorphology in 3156 patients from the Düsseldorf Registry on myelodysplastic syndromes. Leuk Res, 36, 727–734, https://doi.org/10.1016/j.leukres.2012.02.014 (2012).
Dan, S. et al. A large dataset of bone marrow cells in myelodysplastic syndrome for classification systems. figshare. Dataset. https://doi.org/10.6084/m9.figshare.28737170.v1 (2025).
Acknowledgements
We thank Hangzhou Kinsight Biotech Co., Ltd. for providing us with technical and moral support throughout the project.
Author information
Authors and Affiliations
Contributions
The authors confirm contribution to the paper as follows: processing data and generating codes: Junlin Feng and Zhen Huang; design and validate technically: Dan Shen, Xiangli Gao; data collection: Dan Shen, Xiangli Gao, Jixiang Tong, Ting Zhang, Jianhu Li, Shuqi Zhao, Yijing Zhu, Yingqing Xu, Shuchong Yuan and Cheng Jin; manage protocols: Dan Shen, Junlin Feng, and Hongyan Tong. All authors prepared and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
Junlin Feng and Zhen Huang are employees of Kinsight Co., Ltd. There are no other conflicts of interest for other authors.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Shen, D., Gao, X., Tong, J. et al. A large dataset of bone marrow cells in myelodysplastic syndrome for classification systems. Sci Data 12, 1849 (2025). https://doi.org/10.1038/s41597-025-06027-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06027-z