Abstract
Recent advances in metaviromics have revealed vast viral diversity across aquatic environments, yet coastal marine viromes remain underexplored compared to their open-ocean counterparts. In this study, we analyzed 49 surface water samples from 16 coastal sites around Korea, generating 265 gigabases of metagenomic sequence data. Following quality control, 754 DNA viral contigs of ≥10 kb (medium quality or higher) were recovered, with bacteriophages comprising 95% and nucleocytoplasmic large DNA viruses (NCLDVs) 5% of the total. Among these, Puniceispirillum phage HMO-2011 and Micromonas pusilla virus 12 T exhibited the highest relative abundance within their respective groups. In addition, we provided the dataset of environmental parameters such as water temperature, salinity, etc., as well as viral taxonomic profiling of contig-level metadata. This dataset provides a resource for the investigation of coastal DNA viral communities and supports comparative studies across marine environments.
Similar content being viewed by others
Background & Summary
Viruses represent the most numerous biological entities on Earth, with their total population estimated to surpass 1030 globally1. They lyse hosts, providing various nutrients for the marine environment and contributing to biogeochemical cycles in marine ecosystems2. Recent studies estimate that bacteriophages lyse approximately 20%–40% of bacterial populations daily, highlighting their pivotal role in shaping microbial community structure and function3,4. Their prevalence is strongly associated with the high abundance of their bacterial hosts, including Alphaproteobacteria and Cyanobacteria5,6. Marine viruses are ubiquitous, from the surface to the bottom sediments of the ocean, making them the most abundant biological entities in aquatic ecosystems7. However, the diversity of viral communities in coastal waters and their relationships with environmental factors remain comparatively unknown. Coastal ecosystems are ecologically and economically vital, yet highly susceptible to anthropogenic and climate-induced stressors such as eutrophication, pollution, marine heatwaves, and harmful algal blooms (HABs), all of which can profoundly alter viral community structure and function8. Virome studies in these environments are therefore crucial for elucidating how viruses respond to and shape ecosystem dynamics under such pressures.
Viruses are broadly classified into DNA and RNA types, and their classification has advanced through integration of genomic and replication-based systems. The Baltimore classification, originally proposed in the 1970s, groups viruses into seven categories based on nucleic acid type (DNA or RNA), strandedness, sense, and replication strategy9. While still widely cited as a conceptual model, it has been refined by advances in genomic data. According to the most recent taxonomy from the International Committee on Taxonomy of Viruses10, viruses are now organized hierarchically based on evolutionary relationships inferred from genome sequences. In marine ecosystems, DNA viruses are especially dominant and play critical roles in host mortality, nutrient cycling, and microbial community structure11. The two major groups are bacteriophages and nucleocytoplasmic large DNA viruses (NCLDVs)12,13. Bacteriophages are primarily classified under the class Caudoviricetes, which includes families such as Autographiviridae, Straboviridae, Herelleviridae, and Drexlerviridae—all commonly detected in marine viromes14,15,16. NCLDVs infect a broad range of eukaryotic hosts, from unicellular protists to multicellular algae and metazoans17,18. Notably, NCLDVs predominantly infect autotrophic eukaryotes, such as haptophytes, chlorophytes, and dinoflagellates, playing a significant role in regulating primary production5,19,20. Major families within this group include Mimiviridae (formerly Megaviridae), Phycodnaviridae, Pandoraviridae, Poxviridae, Iridoviridae, Marseilleviridae, Pithoviridae, Ascoviridae, Asfarviridae, and Mininucleoviridae21. While numerous studies have underscored the ecological significance of both bacteriophages and NCLDVs, particularly in structuring microbial food webs, regulating host mortality, and influencing global biogeochemical cycles22,23, their diversity, functional capacities, and host interactions across various marine environments remain insufficiently characterized.
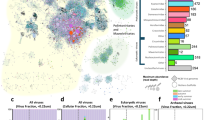
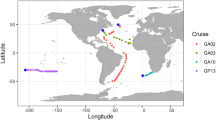
In this study, surface water samples were collected throughout 2021 from 16 coastal sites around the Republic of Korea (Fig. 1). A total of 265 gigabases of raw sequencing reads were generated from 49 samples, which resulted in 4.06 gigabases of assembled contigs after quality filtering (Fig. 2a,b, Table S1). Quality assessment and classification using CheckV categorized the contigs into viral (19.3%) and non-viral (80.7%) groups (Fig. 2c). Contigs of medium or higher quality were filtered based on length thresholds, resulting in 860 contigs ≥3 kb, 840 contigs ≥5 kb, and 754 contigs ≥10 kb. The average length of contigs ≥10 kb was 36,436 base pairs (Fig. 2d,e). Under these stringent thresholds, 19% of the contigs were successfully taxonomically assigned, while the remaining 81% were classified as unassigned, likely due to the lack of significant homology to known viral sequences (Fig. 2f). At the class level, all bacteriophages were classified as Caudoviricetes (Table S2). At the family level, unclassified bacteriophages constituted the largest proportion (66.7%), followed by Zobellviridae (13%) and Stanwilliamsviridae (9.6%) (Fig. 3a). Seven bacteriophages (Puniceispirillum phage HMO-2011, Pelagibacter phage HTVC019P, Lentibacter phage vB_LenP_ICBM1, Pelagibacter phage HTVC011P, Streptomyces phage Gilson, Synechococcus phage S-CBS3, and Lentibacter phage vB_LenP_ICBM2) and Micromonas pusilla virus 12 T and SP1 of Phycodnaviridae (NCLDVs) were predominant at the species level.
Map depicting the 16 sampling sites in the coastal waters of the Republic of Korea in 2021.
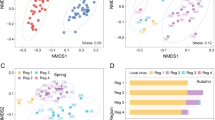
Overview of contig quality check and selection process. (a) Raw metagenomic sequence data. (b) Assembly results of contigs generated from metagenomic sequencing. (c) Classification of assembled contigs into viral and non-viral categories. (d) Contig length distribution after quality check. (e) Distribution of contigs longer than 10 kb. (f) Results of Classified Viruses (e-value: 10−5 or Open Ring Frames associated with viral hallmarker gene) and Unclassified Viral Contigs.
Taxonomic composition of viral contigs based on BLASTn annotation. (a) Family-level distribution of taxonomically assigned viral contigs. (b) Species-level composition of predominant viral taxa (>5% average abundance).
Environmental parameters observed during the sampling period are described on Figshare24. In terms of environmental factors, the water temperature range from11.26–34.03 °C (mean ± Standard deviation: 21.8 ± 5.8 °C). The mean salinity was 30.34, excluding Cheonsuman (St. 13, mean salinity 2.68), a freshwater lake connected to the Cheonsuman coastal area, and Eulsukdo (St. 4, mean salinity 13.08), an estuarine area influenced by freshwater input from the Nakdong River. The concentrations of dissolved inorganic nitrogen (DIN) and dissolved inorganic phosphorus (DIP) were 21.48 ± 22.98 μM and 0.92 ± 0.91 μM, respectively. Chlorophyll-a concentrations ranged from 0.58 to 37.2 μg L−1, with monthly mean exceeding 5 μg L−1 in April - May and November - December. Dissolved organic carbon (DOC) concentrations ranged from 1.00 to 5.73 mg L−1, with higher values observed during periods of elevated chlorophyll-a concentrations.
Methods
Sample collection and environmental measurement
To investigate seasonal variation in coastal viral communities and environmental factors, 49 surface seawater samples were collected from 16 sites in Korean coastal waters in 2021 (Fig. 1). At each sampling site, a total of 60 L seawater samples were collected using a Niskin water sampler (General Oceanics Inc., Miami, FL, USA) and transferred to pre-cleaned polyethylene (PE) bottles. Immediately after collection, samples were maintained at 4 °C and transported to the laboratory for further analysis.
The methodology for assessing environmental parameters and biological factors was based on our previous studies25,26,27. Environmental variables, including temperature and salinity, were measured at each sampling site using a multiparameter water quality sonde (EXO2, YSI Inc., Yellow Springs, OH, USA). To ensure data accuracy and cross-validation, three EXO2 multi-parameters were deployed and operated simultaneously. For dissolved inorganic nutrients, dissolved organic carbon (DOC), and chlorophyll-a (Chl-a) analyses, 2 L of seawater were collected from each site into sterile polyethylene (PE) containers, stored in ice-cooled boxes, and transported to the South Sea Research Institute, KIOST. A 50-mL subsample was filtered through a 47-mm GF/F filter (Whatman, Clifton, NJ, USA) under gravity. The resulting filtrates were transferred into acid-washed PE bottles and either immediately analyzed or stored at –80 °C for no longer than 7 days prior to analysis. Dissolved inorganic nitrogen (NO₂ + NO₃ + NH₄+) and dissolved inorganic phosphorus concentrations were determined using a QuAAtro39 continuous flow analyzer (SEAL Analytical, UK). DOC concentrations were quantified via high-temperature catalytic oxidation using a TOC-VCPH analyzer (Shimadzu, Kyoto, Japan). To determine Chl-a concentrations, 1 L of seawater was filtered through a GF/F membrane filter and extracted in 90% acetone under dark conditions at 4 °C for 24 hours. Chl-a concentrations were then measured using a fluorometer (Trilogy; Turner Designs, Sunnyvale, CA, USA). All measurements were conducted in triplicate to ensure reproducibility.
Virus flocculation, resuspension, DNA extraction, and sequencing
Marine viral collection methods include ultracentrifugation28, filtration using ultrafine membranes (<0.2 µm)29, and aggregation with iron ions (Fig. 4). Among these, Fe-based virus flocculation, filtration, and resuspension (FFR) is highly efficient (>90% recovery), cost-effective, and reliable, making it suitable for studies on marine viral ecology and genomics30. In this study, a modified FFR method was applied to analyse marine DNA virus communities31. To extract viral genomic DNA (gDNA), 20-L of seawater was filtered through a 5-μm membrane (TMTP04700; Merck Millipore, MA, USA) to remove large organic and inorganic particles. Viruses were concentrated via flocculation using Fe3+ ions and collected on a 0.2-μm polycarbonate membrane (111106; Whatman, Buckinghamshire, UK), which was stored at 4 °C. Most experiments were performed in triplicate. Following a previously established protocol27, we prepared an FeCl₃ solution containing 16.53 mg Fe³+ mL−1, and added 1 mL of this solution per 10 L of seawater to induce virus particle flocculation. For DNA extraction, the membrane was cut into small sections and placed in a suspension buffer (10 mL of 0.1 M EDTA, 0.2 M MgCl2, 0.2 M Ascorbate) in a 50-mL conical tube (Fig. 4). Viruses were released by suspending them in the buffer, and the pH was adjusted to 6 with approximately 5 mL of 10 M NaOH solution. Total gDNA was extracted using the Viral Gene-spin Viral DNA/RNA Extraction Kit (iNtRON Biotechnology, Seoul, South Korea). The extracted gDNA was used to construct a metavirome library with the NEBNext Ultra II DNA Library Prep Kit (Illumina, San Diego, CA, USA), involving random DNA fragmentation, 5′ and 3′ adapter ligation, and amplification via polymerase chain reaction. The prepared library was then sequenced using the Illumina HiSeq X platform in paired-end mode.
Workflow for DNA Viral metavirome Analysis. (a) Sample collection to metavirome analysis. (b) Bioinformatics of DNA viral contigs.
Bioinformatic analyses of DNA viruses
Bioinformatics analysis was conducted using a modified protocol32,33 (Fig. 4). Raw sequencing data were processed using CLC Genomics Workbench v20.0.4 (Qiagen, Hilden, Germany), with low-quality reads and sequencing adaptors removed during the preprocessing step. De novo assembly of viral contigs was performed using metaSPAdes v3.13.034 with the following command: ‘metaspades.py -k 21,33,55,77,99,127–pe1-1 R1.fastq–pe1-2 R2.fastq -t 32 -o output_dir’. Assembly quality was assessed using CheckV v1.435 (‘checkv end_to_end input_dir output_dir -t 32’), applying a minimum contig length threshold of 10 kb and included only medium-quality or higher sequences (Table S1, S3). Contig length distributions (≥3 kb, ≥5 kb, and ≥10 kb) are summarized in Table S1. Contigs were dereplicated and clustered at ≥95% average nucleotide identity (ANI) using VSEARCH v2.28.136,37 with ‘vsearch–cluster_size Sample.fa–id 0.95–strand both–sizein–sizeout–fasta_width 0–uc output.uc–centroids output.fa’. Mapping was performed with BBMap v38.5138 (‘bbmap.sh in1 = R1.fastq in2 = R2.fastq covstats = output.txt’) at 95% identity, followed by removal of sequencing adapter sequences and PhiX174 control phage contamination. Taxonomic annotation was conducted using BLASTn v2.13.039 against the NCBI Viral RefSeq nucleotide database, with thresholds of e-value ≤ 1e−540, identity ≥ 70%, and bitscore ≥ 50 (Table S4). The BLASTn command used was: ‘blastn -query input.fasta -db RefSeqViral_nucl -evalue 1e-5 -max_target_seqs. 10 -outfmt “6 qseqid sseqid pident length mismatch gapopen qstart qend sstart send evalue bitscore stitle qcovs” -num_threads 16> output.outfmt6’. To complement nucleotide-based classification, BLASTp was also performed against the Viral RefSeq protein database to verify the presence of viral hallmark genes: ‘blastp -query input.fasta -db RefSeqViral_prot -evalue 1e-5 -max_target_seqs. 1 -outfmt “6 qseqid sseqid pident length mismatch gapopen qstart qend sstart send evalue bitscore stitle scomnames” -num_threads 16> output.outfmt6’. Based on the annotation results, contigs were categorized as known viruses (containing annotated viral genes) or unknown viruses (lacking identifiable viral protein matches). In total, 68 viral species were identified under these parameters.
Data Records
The Illumina sequencing data have been deposited in the NCBI Sequence Read Archive (SRA) under BioProject accession PRJNA121880341. Moreover, the assembled fasta files of viral contigs (≥10 kb) used in this study are deposited in GenBank Nucleotide under accession numbers PV702959 to PV703754. A total of 796 nucleotide sequences (accession numbers PV702959–PV703754) have been uploaded to BioProject PRJNA121880341. In addition, the environmental metadata are publicly available on Figshare (https://doi.org/10.6084/m9.figshare.29167460.v1)42, and the assembled viral contigs (.fasta) are deposited in a separate Figshare repository (https://doi.org/10.6084/m9.figshare.29603600)24.
Technical Validation
Library quality control
The raw metavirome reads were assessed for quality using the CLC Genomics Workbench v20.0.4 to ensure sequencing data integrity. The calculated Q scores showed that an average of 63.04% to 93.58% of the reads across all sampling sites had a Q30 or higher. These findings suggest that the sequencing data were of high quality, making them appropriate for metagenomic analysis.
Taxonomic profiling validation
Taxonomic profiling validation confirmed that species-level assignments were consistent and robust. BLASTn classifications were supported by BLASTp results, and contigs containing hallmark or replication-associated genes were reliably classified as known viruses, whereas contigs lacking such genes were categorized as unknown viruses. These results validate the robustness of the taxonomic assignments.
Replication
To ensure accurate interpretation and reproducibility, three replicate (each 20 L) seawater samples were collected at each sampling site. Independent sequencing libraries were prepared for each replicate and subjected to separate quality control (QC), with only those passing QC included in the analyses. For downstream analyses, the triplicate datasets were processed to calculate the mean and standard deviation, which were then used for interpretation.
Data availability
The raw sequencing reads generated in this study are deposited in the NCBI Sequence Read Archive under BioProject accession PRJNA1218803 (SRA Study SRP565150)41, comprising 18 BioSamples and 117 SRA runs. The assembled viral contigs (≥10 kb, n = 796) are available in GenBank under accession numbers PV702959–PV703754, all linked to BioProject PRJNA121880341. The environmental metadata and assembled viral contigs are deposited in a Figshare repositories24,42.
Code availability
The newly developed code was not used in this study.
References
Suttle, C. A. Viruses in the sea. Nature 437, 356–361, https://doi.org/10.1038/nature04160 (2005).
Rastrojo, A. & Alcamí, A. Aquatic viral metagenomics: Lights and shadows. Virus Res. 239, 87–96, https://doi.org/10.1016/j.virusres.2016.11.021 (2017).
Weinbauer, M. G. Ecology of prokaryotic viruses. FEMS Microbiol. Rev. 28, 127–181, https://doi.org/10.1016/j.femsre.2003.08.001 (2004).
Weinbauer, M. G. & Rassoulzadegan, F. Are viruses driving microbial diversification and diversity? Environ. Microbiol. 6, 1–11, https://doi.org/10.1046/j.1462-2920.2003.00539.x (2004).
Suttle, C. A. Marine viruses — major players in the global ecosystem. Nat. Rev. Microbiol. 5, 801–812, https://doi.org/10.1038/nrmicro1750 (2007).
Suttle, C. A. in Ecological, evolutionary, and geochemical consequences of viral infection of cyanobacteria and eukaryotic algae. in Viral ecology (ed. Hurst, C. J.) 247–296 (Academic Press, 2000).
Zhang, Q.-Y., Ke, F., Gui, L. & Zhao, Z. Recent insights into aquatic viruses: Emerging and reemerging pathogens, molecular features, biological effects, and novel investigative approaches. Water Biol. Secur. 1, 100062, https://doi.org/10.1016/j.watbs.2022.100062 (2022).
Du, X. et al. Virome reveals effect of Ulva prolifera green tide on the structural and functional profiles of virus communities in coastal environments. Sci. Total Environ. 883, 163609, https://doi.org/10.1016/j.scitotenv.2023.163609 (2023).
Baltimore, D. Expression of animal virus genomes. Bacteriol. Rev. 35, 235–241, https://doi.org/10.1128/br.35.3.235-241.1971 (1971).
Simmonds, P. et al. Changes to virus taxonomy and the ICTV Statutes ratified by the International Committee on Taxonomy of Viruses (2024). Arch Virol. 169, 236, https://doi.org/10.1007/s00705-024-06143-y (2024).
Mayers, K. M. J. et al. Grazing on Marine Viruses and Its Biogeochemical Implications. mBio 14, e01921–01921, https://doi.org/10.1128/mbio.01921-21 (2023).
Middelboe, M. & Brussaard, C. P. D. Marine Viruses: Key Players in Marine Ecosystems. Viruses 9, 302, https://doi.org/10.3390/v9100302 (2017).
Minch, B. et al. Phylogenetic diversity and functional potential of large and cell-associated viruses in the Bay of Bengal. mSphere 8, e00407–00423, https://doi.org/10.1128/msphere.00407-23 (2023).
Jung, S. W., Kim, K. E., Kim, H.-J. & Lee, T.-K. Metavirome Profiling and Dynamics of the DNA Viral Community in Seawater in Chuuk State, Federated States of Micronesia. Viruses 15, 1293, https://doi.org/10.3390/v15061293 (2023).
Kim, K. E. et al. Ecological Interaction between Bacteriophages and Bacteria in Sub-Arctic Kongsfjorden Bay, Svalbard, Norway. Microorganisms 12, 276, https://doi.org/10.3390/microorganisms12020276 (2024).
Zhu, Y., Shang, J., Peng, C. & Sun, Y. Phage family classification under Caudoviricetes: A review of current tools using the latest ICTV classification framework. Front. Microbiol. 13, https://doi.org/10.3389/fmicb.2022.1032186 (2022).
Colson, P. et al. “Megavirales”, a proposed new order for eukaryotic nucleocytoplasmic large DNA viruses. Arch. Virol. 158, 2517–2521, https://doi.org/10.1007/s00705-013-1768-6 (2013).
Monier, A., Claverie, J.-M. & Ogata, H. Taxonomic distribution of large DNA viruses in the sea. Genome Biol. 9, R106, https://doi.org/10.1186/gb-2008-9-7-r106 (2008).
Gregory, A. C. et al. Marine DNA Viral Macro- and Microdiversity from Pole to Pole. Cell 177, 1109–1123.e1114, https://doi.org/10.1016/j.cell.2019.03.040 (2019).
Endo, H. et al. Biogeography of marine giant viruses reveals their interplay with eukaryotes and ecological functions. Nat. Ecol. Evol. 4, 1639–1649, https://doi.org/10.1038/s41559-020-01288-w (2020).
Mönttinen, H. A. M., Bicep, C., Williams, T. A. & Hirt, R. P. The genomes of nucleocytoplasmic large DNA viruses: viral evolution writ large. Microbial Genomics 7, https://doi.org/10.1099/mgen.0.000649 (2021).
Brum, J. R. & Sullivan, M. B. Rising to the challenge: accelerated pace of discovery transforms marine virology. Nat. Rev. Microbiol. 13, 147–159, https://doi.org/10.1038/nrmicro3404 (2015).
Breitbart, M. Marine viruses: truth or dare. Annu. Rev. Mar. Sci. 4, 425–448, https://doi.org/10.1146/annurev-marine-120709-142805 (2012).
Kim, Y. J. et al. Dynamics of the DNA Viral Community in Korean Coastal Waters. figshare. https://doi.org/10.6084/m9.figshare.29603600 (2025).
Kim, Y. J. et al. Determining ecological interactions of key dinoflagellate species using an intensive metabarcoding approach in a semi-closed coastal ecosystem of South Korea. Harmful Algae 138, 102698, https://doi.org/10.1016/j.hal.2024.102698 (2024).
Kim, H.-J. et al. Co-occurrence between key HAB species and particle-attached bacteria and substrate specificity of attached bacteria in the coastal ecosystem. Harmful Algae 138, 102700, https://doi.org/10.1016/j.hal.2024.102700 (2024).
Kim, M.-J. et al. Co-occurrence patterns between Chlorophyta and nucleocytoplasmic large DNA virus in coastal ecosystem, South Korea. Mar. Environ. Res. 204, 106944, https://doi.org/10.1016/j.marenvres.2025.106944 (2025).
Colombet, J. et al. Virioplankton ‘pegylation’: Use of PEG (polyethylene glycol) to concentrate and purify viruses in pelagic ecosystems. J. Microbiol. Meth. 71, 212–219, https://doi.org/10.1016/j.mimet.2007.08.012 (2007).
Steward, G. F. & Culley, A. I. Extraction and purification of nucleic acids from viruses. in Manual of Aquatic Viral Ecology (eds Wilhelm, S. W., Weinbauer, M. G. & Suttle, C. A.) 154-165 (American Society of Limnology and Oceanography, 2010).
John, S. G. et al. A simple and efficient method for concentration of ocean viruses by chemical flocculation. Environ. Microbiol. Rep. 3, 195–202, https://doi.org/10.1111/j.1758-2229.2010.00208.x (2011).
Kang, J. et al. Zooming on dynamics of marine microbial communities in the phycosphere of Akashiwo sanguinea (Dinophyta) blooms. Mol. Ecol. 30, 207–221, https://doi.org/10.1111/mec.15714 (2021).
Kim, K. E. et al. Optimized Metavirome Analysis of Marine DNA Virus Communities for Taxonomic Profiling. Ocean Sci. J. 57, 259–268, https://doi.org/10.1007/s12601-022-00064-0 (2022).
Kim, K. E. et al. Covariance of Marine Nucleocytoplasmic Large DNA Viruses with Eukaryotic Plankton Communities in the Sub-Arctic Kongsfjorden Ecosystem: A Metagenomic Analysis of Marine Microbial Ecosystems. Microorganisms 11, 169, https://doi.org/10.3390/microorganisms11010169 (2023).
Nurk, S., Meleshko, D., Korobeynikov, A. & Pevzner, P. A. metaSPAdes: a new versatile metagenomic assembler. Genome Res. 27, 824–834 (2017).
Nayfach, S. et al. CheckV assesses the quality and completeness of metagenome-assembled viral genomes. Nat. Biotechnol. 39, 578–585, https://doi.org/10.1038/s41587-020-00774-7 (2021).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584, https://doi.org/10.7717/peerj.2584 (2016).
Roux, S. et al. Minimum Information about an Uncultivated Virus Genome (MIUViG). Nat. Biotechnol. 37, 29–37, https://doi.org/10.1038/nbt.4306 (2019).
Bushnell, B. BBMap: a fast, accurate, splice-aware aligner. (2014).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410, https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
Roux, S. et al. Ecogenomics of virophages and their giant virus hosts assessed through time series metagenomics. Nat Commun 8, 858, https://doi.org/10.1038/s41467-017-01086-2 (2017).
Kim, Y. J. et al. Assembled viral contigs (≥10 kb) from South Korea coastal waters. NCBI GenBank https://identifiers.org/ncbi/bioproject:PRJNA1218803 (2025).
Kim, Y. J. et al. Environmental Monitoring Metadata from Korean Coastal Waters. figshare. https://doi.org/10.6084/m9.figshare.29167460.v1 (2021).
Acknowledgements
The genomic DNA samples were stored in the Library of Marine Samples at the Korea Institute of Ocean Science & Technology (KIOST), Republic of Korea. This research was supported by the Korea Institute of Marine Science & Technology Promotion (KIMST), funded by the Ministry of Oceans and Fisheries of Korea (RS-2021-KS211475) and by National Marine Biodiversity Institute of Korea (project name: 2025 Establishment and Operation of an Information System for the Registration, Evaluation, Collection, Preservation, and Management of Bioresources from the High Seas; PG55180).
Author information
Authors and Affiliations
Contributions
Yu Jin Kim and Seung Won Jung conceived and designed the study; Yu Jin Kim, and Kang Eun Kim collected and tabulated the data; Yu Jin Kim, Kang Eun Kim, Hyun-Jung Kim, Taehee Lee, and Seung Won Jung constructed the database and analysed the data; Yu Jin Kim, Kang Eun Kim, Min-Jeong Kim, Seon Min Kim, and Seung Won Jung contributed materials and analysis tools; Yu Jin Kim, Joon Sang Park, Min-Jeong Kim, Seon Min Kim, and Seung Won Jung wrote the original draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kim, Y.J., Kim, K.E., Kim, HJ. et al. Dynamics of the DNA Viral Community in Korean Coastal Waters. Sci Data 12, 1782 (2025). https://doi.org/10.1038/s41597-025-06062-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-025-06062-w