Abstract
Apple (Malus domestica Borkh.) is a major fruit crop with a rich genetic history shaped by whole-genome duplication, domestication, and selective breeding. Discovering apple genetic diversity through genome sequencing provides new opportunities to improve disease resistance, environmental adaptation, and fruit quality. Here, we present 19 haplotype-resolved genome assemblies of apple, sequenced using PacBio HiFi reads with approximately 30 × coverage. Each haplome assembly has a mean length of 675.3 Mb and contains on average 47,445 annotated protein-coding genes. These haplome assemblies have a high completeness, with mean complete BUSCO scores of 98.8%. We identified 578 previously uncharacterized orthogroups shared across all 38 haplomes, indicating that these assemblies capture novel genetic diversity. Many of the assemblies are also highly contiguous, with on average three to four phase switches per chromosome. These data will accelerate genome-wide association studies, helping researchers to find and use genetic diversity for the improvement of key traits. Additionally, these data can offer insights into evolutionary history, domestication, and genetic diversity, supporting apple breeding and the broader Rosaceae research community.
Similar content being viewed by others
Background & Summary
Apple (Malus domestica Borkh.) is the third most valuable fruit crop grown globally1. The development of genomic resources, such as high-quality genome assemblies, can enable the dissection of key traits and lay the foundation for genomics-assisted breeding. Initial attempts at assembling apple genomes were, for a long-time, limited by the high degree of heterozygosity present in apple genotypes. The first whole genome sequence (WGS) for apple was reported for the cultivar ‘Golden Delicious’ and was generated using Sanger and 454 pyrosequencing technology2. An additional genome assembly of the cultivar ‘Golden Delicious’ was later published, using over 100-fold coverage Illumina short read combined with 29-fold coverage PacBio long read sequencing3. These assemblies were highly fragmented in part due to sequencing limitations and apple genome heterozygosity, as evidenced by a maximum N50 size of 111,619 bp3.
To bypass the technical challenges in assembling heterozygous genomes, a homozygous doubled haploid genotype of the apple cultivar ‘Golden Delicious’ was sequenced and assembled using a combination of Illumina short read, PacBio long read, and Bionano optical genome mapping technologies4. The resulting GDDH13v1.1 assembly had a genome contiguity (N50) that was an order of magnitude greater (5.5 Mb) than the former ‘Golden Delicious’ assembly (0.11 Mb) and served as the reference genome for apple. Consequently, researchers continued to exploit haploid accessions for subsequent genome assembly. In 2019, another homozygous line was sequenced, an anther-derived trihaploid ‘Hanfu’ (HFTH1) line, using Illumina short read, PacBio long read, Bionano optical mapping data, and Hi-C data for assembly5. In 2022, a homozygous tetra-haploid ‘Royal Gala’ plant was sequenced using Illumina short read, PacBio long read, and Hi-C library6. However, haploid assemblies of homozygous genotypes derived from anther culture only capture one of the two haplotypes from their heterozygous donors. With the rapid advances in long read sequencing technologies and assembly methods7, it is feasible to sequence heterozygous apple genotypes and generate haplotype–resolved assemblies containing both haplotypes. Haplotype-resolved genome assemblies distinguish both parental haplotypes, providing an improved representation of diploid species genomes, such as apple8.
The first attempt at a haplotype-resolved genome assembly was for the cultivar ‘Gala Galaxy’ using PacBio long reads for assembly, Illumina short reads for polishing, together with Bionano optical mapping for scaffolding9. Using the same methodology, along with a 10x Genomics library, the genomes of ‘Gala’ and two wild progenitors, Malus sieversii Ldb.and Malus sylvestris Mill., were assembled and phased10. This was followed by additional phased genome assemblies of the apple cultivars ‘Honeycrisp’, ‘Antonovka’, ‘Red Fuji’, and ‘WA 38'11,12,13,14. Recently, a haplotype-resolved assembly of the dwarfing apple interstock hybrid ‘SH6’ (Malus honanensis Rehder × Malus domestica Borkh.) and a telomere-to-telomere phased genome of assembly of ‘Golden Delicious’ were also published15,16. Furthermore, near-gapless haplotype-resolved assemblies for the dwarf rootstock ‘M9’, semi-rigorous rootstock ‘MM106’ and popular cultivar ‘Fuji’ were released17. These developments show that haplotype-resolved genome assemblies are a valuable resource for accurately characterizing highly heterozygous, diploid species, such as apple.
In this study, we sequenced 19 apple accessions (Fig. 1) using Pacific Bioscience’s high-fidelity sequencing technology. This resulted in 19 chromosome-level, haplotype-resolved genome assemblies, corresponding to 38 haplotype assemblies (haplomes). The haplome sizes ranged between 601.8 Mb to 767.1 Mb, with a mean of 675.3 Mb. The haplomes were highly complete with total complete Benchmarking Universal Single-Copy Orthologs (BUSCO) scores ranging between 93.1% to 99.3% (\(\bar{x}\) = 98.8%). When evaluated with pedigree-phased high-quality single nucleotide polymorphism (SNP) array data, on average three to four phase switches were found per pseudo-chromosome, suggesting highly contiguous phasing. Protein-coding genes were annotated, and the number of gene models per haplome ranged between 43,278 to 49,666 (\(\bar{x}\) = 47,445). Orthologous clustering of these proteins, with proteins from reference genome assemblies from GDDH13 v1.1, HFTH1 v1.0 and Honeycrisp v1.1.a1 (n = 1,988,899), resulted in defining 60,012 orthogroups4,5,11. Among the identified orthogroups, 13,985 were found to be shared across all haplotype-resolved chromosome-level genomes. These publicly available genomes are a resource for advancing genomic studies in apple with broader applications across Rosaceae species.
Photographs of apples from the cultivars that were sequenced in this study.
Methods
Samples collection, library construction and sequencing
Nineteen apple accessions were selected for whole-genome sequencing (WGS), of which fourteen were drawn from the REFPOP18. Photographs of the apples collected from each accession are shown in Fig. 1. These accessions were chosen based on the following criteria: (i) phenotypic extremes: to represent the range of stomatal density, with accessions selected for high and low stomatal density as described by Zuffa et al.19 (ii) genetic diversity: to capture a broad spectrum of genetic diversity within the REFPOP, as illustrated by the inclusion of cultivars such as ‘Giambun’, ‘Prima’, ‘Rouget’, and ‘Tropical Beauty’ (Fig. 2) (iii) historical and commercial importance: to include cultivars of historical or commercial relevance, such as the cultivars ‘Granny Smith’ and ‘McIntosh’. The remaining five accessions were selected by breeders at Agroscope based on additional breeding and selection priorities. For WGS, approximately two grams of fresh growing leaves were harvested from the same tree of each accession in 2023. The sampled fresh leaves were flash frozen in liquid nitrogen, within one hour of collection and stored at −80 °C. The DNA extraction, library construction, and WGS were done by the Arizona Genomics Institute, USA, aiming for thirty-fold coverage per genome with the Revio system (Pacific Biosciences, Menlo Park, California, USA) to generate HiFi long reads. A workflow of the bioinformatic pipeline is shown in Fig. 3.
Principal component analysis (PCA) of REFPOP accessions based on 480 K SNP chip data. The original 480 K SNP chip dataset was converted to numeric format (0–1), with duplicated markers removed prior to analysis. PCA was performed using the ‘prcomp’ function in R. Green points represent all REFPOP accessions, while those labeled in black correspond to the accessions selected for WGS. The x- and y-axes represent principal components (PCs), with the variance explained indicated in parentheses. The four panels display the first eight PCs, which together account for 31% of the total variance.
Bioinformatic workflow of the sequencing and assembly pipeline.
Genome assembly
Raw data was converted from ‘bam’ to ‘fastq’ format using Samtools v1.19.220. These data were then assembled using Hifiasm v0.19.8 and the resulting ‘asm’ haplome assemblies were extracted in ‘fasta’ format using Awk21. These assemblies were then organized into chromosomes based on the HFTH1 v1.0 reference genome using RagTag v2.1.05,22, which incorporated Minimap2 v2.26 for alignments with the long assembly to reference mapping preset “-x asm5” and “-f 0.02”23.
Mitochondrial contamination errors were detected after uploading the haplome assemblies to NCBI (https://www.ncbi.nlm.nih.gov/home/genomes/) for quality assessment. The mitochondrial contaminants were removed from the genome using BEDTools v2.27.124. The remaining sequences were extracted using SeqKit v2.4.025, and concatenated together using Biopython v1.8426.
Genome annotation
To annotate transposable elements (TEs) in the haplome assemblies, a pan-pome fruit transposable element library (pomeFruitTElib) was constructed based on six Malus spp. (GDDH13 v1.14, HFTH1 v1.05, Gala diploid Genome v1.010, Royal Gala v1.0, M. sieversii v2, and M. sylvestris v26), six Pyrus spp. (P. communis Bartlett DH v227, P. bretschneideri v1.128, P. pyrifolia Nijisseiki r.1.0.pmol29, P. pyrifolia Cuiguan v1.130, P. betuleafolia v1.031, and P. ussuriensis × P. communis, Zhongai v1.032) and one Gillenia genome assembly33. The haplome assemblies were processed in parallel using the ‘genePal’ pipeline developed at The New Zealand Institute for Plant and Food Research Limited (https://github.com/Plant-Food-Research-Open/genepal), which includes repeat masking with pomeFruitTElib, ab inito gene prediction with BRAKER334, liftoff gene models from ‘Viridiplantae’ in OrthoDB35 and published Malus assemblies (Honeycrisp v1.1.a111, HFTH1 v1.05, GDDH13 v1.14), followed by functional annotation with EggNOG-mapper (https://github.com/eggnogdb/eggnog-mapper) on EggNOG v5.0 database36.
Genome completeness
To assess the comprehensiveness of the assemblies, we used the BUSCO v5.1.2 tool, leveraging orthologous genes from the ‘embryophyta_odb10’ data37.
Orthologous clustering
Orthologous relationships for genes were calculated using OrthoFinder v2.5.438,39. For genes predicted in each haplome, the translated amino acid sequence of the primary transcript was selected as the representative protein. In addition, protein sequences from GDDH13 v1.14, HFTH1 v1.05, and Honeycrisp v1.1.a111 were included in the orthologous analysis. An all-vs-all multiple sequence alignments of proteins from these genomes were performed with Mafft v7.30740,41,42. The OrthoFinder result was plotted using R v4.3.343.
Genome phasing quality check
The phasing quality of each haplome assembly was evaluated using the R-script developed by Vanderzande et al.44. Briefly, this reference SNP array data was obtained from the 20K SNP array45 and through other studies46,47,48 and was error-cleaned and phased according to Vanderzande et al.49. First, each haplome was compared to the iGL genetic map46, which is based on SNP array data45. Probe sequences for each SNP in this iGL map were aligned to each haplome using BLAST50 and the SNPs’ likely positions were determined according to Vanderzande et al.44. Then, per 2 cM interval of the iGL map, the proportion of SNPs not having a unique location in the haplome and the proportion of SNPs having a location inconsistent with the genetic map were recorded to indicate problematic regions where a haplome may have issues. Second, uniquely aligned SNPs that showed consistency with the iGL map were extracted from each haplome assembly and compared to the reference SNP array data. The proportion of genotypic inconsistencies between the SNP array data and alleles extracted from the assembly was determined to ensure the correct individual was sequenced. Furthermore, the phasing of the assembly was evaluated by comparing the SNP alleles from each haplome in the assembly to the phased SNP array data. Supplementary Table 1 provides information for which individuals reference SNP array data and accurate reference phasing information was available. For cultivars ‘Giambun’, ‘Kitchkova’, ‘Rouget’ and ‘Scilly Pearl’, SNP array data could not be accurately phased because only a few direct relatives were present. For ‘Ipador’ no reference SNP array data was available. For these reasons, results from the cultivars ‘Giambun’, ‘Kitchkova’, ‘Rouget’, ‘Scilly Pearl’ and ‘Ipador’ were excluded.
Data Records
The genome assemblies have been deposited at GenBank under the accessions: ‘Braeburn’ haplotype 1, GCA_052939155.151; ‘Braeburn’ haplotype 2, GCA_052939175.152; ‘Coxs Orange Pippin’ haplotype 1, GCA_052938675.153; ‘Coxs Orange Pippin’ haplotype 2, J GCA_052938685.154; ‘Edward VII’ haplotype 1, GCA_052938595.155; ‘Edward VII’ haplotype 2, J GCA_052938615.156; ‘Enterprise’ haplotype 1, GCA_052939425.157; ‘Enterprise’ haplotype 2, GCA_052939435.158; ‘ Esopus Sptizenburg’ haplotype 1, GCA_052939395.159; ‘ Esopus Sptizenburg’ haplotype 2, GCA_052939415.160; ‘Fiessers Erstling’ haplotype 1, GCA_052938715.161; ‘Fiessers Erstling’ haplotype 2, GCA_052938735.162; ‘Giambun’ haplotype 1, GCA_052939355.163; ‘Giambun’ haplotype 2, GCA_052939365.164; ‘Giga ‘ haplotype 1, GCA_052939315.165; ‘Giga ‘ haplotype 2, GCA_052939335.166; ‘Granny smith’ haplotype 1, GCA_052939275.167; ‘Granny smith’ haplotype 2, GCA_052939295.168; ‘Jonathan’ haplotype 1, GCA_052939245.169; ‘Jonathan’ haplotype 2, GCA_052939235.170; ‘Kitchkova’ haplotype 1, GCA_052939185.171; ‘Kitchkova’ haplotype 2, GCA_052939215.172; ‘McIntosh’ haplotype 1, GCA_052939115.173; ‘McIntosh’ haplotype 2, GCA_052939125.174; ‘Milwa’ haplotype 1, GCA_052939075.175; ‘Milwa’ haplotype 2, GCA_052939095.176; ‘Prima’ haplotype 1, GCA_052939015.177; ‘Prima’ haplotype 2, GCA_052938985.178; ‘Priscilla’ haplotype 1, GCA_052938555.179; ‘Priscilla’ haplotype 2, GCA_052938575.180; ‘Rouget’ haplotype 1, GCA_052938935.181; ‘Rouget’ haplotype 2, GCA_052938975.182; ‘Scilly Pearl’ haplotype 1, GCA_052938915.183; ‘Scilly Pearl’ haplotype 2, GCA_052938925.184; ‘SQ59’ haplotype 1, GCA_052939035.185; ‘SQ59’ haplotype 2, GCA_052939045.186; ‘Tropical Beauty’ haplotype 1, GCA_052938535.187; ‘Tropical Beauty’ haplotype 2, GCA_052938515.188.
All raw data and assemblies have been deposited in NCBI under BioProject PRJNA116848589 and Sequence Read Archive (SRA) SRP56069090. The BioProject accessions for each haplome are shown in Supplementary Table 1.
Technical Validation
Genome contiguity
The final haplome sizes, ranged between 592,406,660 to 676,548,961 bp, with a mean of 648,968,431 bp (Fig. 4, Supplementary Table 1). For all assemblies, 90% (N90) of the bases were allotted to 16–17 contigs, the only exception being haplotype 1 of ‘Esopus Spitzenbug’ (N90 = 365) (Fig. 4, Supplementary Table 1). These results indicate that most assemblies represent chromosome level assemblies.
Barplot of the assembly statistics for each assembled haplome, showing total scaffold length, scaffold count, L90, BUSCO completeness, and predicted gene number.
The haplome assemblies, evaluated for completeness using BUSCO with the embryophyta_odb10 data, yielded complete BUSCO scores ranging between 93.1% to 99.3% with a mean of 98.8% (Fig. 4, Supplementary Table 1). While most of the genes were single copy (\(\bar{x}\) = 62.0%) the remaining were mostly duplicated copies (\(\bar{x}\) = 31.0%), which is expected in a species whose genome underwent a recent whole genome duplication91. However, the haplotype 2 assembly of cv. ‘Braeburn’ was relatively less complete, with fragmented copies and missing copies accounting for 1.3% and 5.6%, respectively. This was also confirmed in the comparison with the genetic map where only 81% of SNPs could be located (compared to approximately 94% for other assemblies). Given the high BUSCO scores we conclude that the assembled haplomes are of high completeness.
Gene and protein content
The number of protein coding genes per haplome ranged between 43,278 and 49,666, with a mean of 47,445. This is greater than the doubled haploid GDDH13 v1.14 genome (n = 42,140) and the triple haploid HFTH1 v1.05 genome (n = 39,617) but more in line with that of the phased genomes of Gala v1.0 and Malus sieversii v1.0 and Malus sylvestris v1.010 (n = 45,199-45,352).
Orthogroups
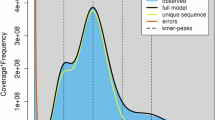
Protein sequences were identified for 1,988,899 genes among 42 haplome assemblies (two consensus haploid assemblies and 20 cultivars with haplotype-resolved assemblies). In total 1,978,214 (99.46%) of the genes were assigned to 60,012 orthogroups and only 10,685 (0.54%) were singleton genes. There were 13,985 orthogroups where all assemblies were present, while there were 8,647 single-copy orthogroups. These data indicate 2,669 orthogroups are not found in HFTH1 v1.0, 1,010 orthogroups are not found in GDDH13 v1.1, and 578 orthogroups are not found in both HFTH1 v1.0 and GDDH13 v1.1, but are present in all others. An overall visualization of the results is illustrated in Supplementary Fig. 1. These data demonstrate the completeness (based on overlap) and novel diversity (based on abundance of single-copy orthogroups) of the genomes sequenced.
Genome phasing quality check
To assess the quality of the phasing of the haplome assemblies, we compared the haplomes to SNP array data from individuals in an accession’s pedigree, for fourteen cultivars with complementary SNP array data. We found that, in general, phasing was mostly contiguous at a local level with phase switches occurring on average three to four times per chromosome (Fig. 5). For the cultivar ‘Braeburn’, the higher number of phase switches indicates a lower quality of phasing and is likely due to low sequence coverage. Therefore, this assembly could be improved in the future using more sequence data (Fig. 5). The phasing analysis indicates that the majority of haplomes provided here represent high quality phased genome assemblies. The challenges of phasing certain cultivars, for example due to lack of phased reference SNP data from direct relatives, indicates we have captured accessions that possess novel genetic diversity compared to popular breeding material that has already been sequenced. Overall, the genome assemblies presented here represent a resource for small scale local imputation. However, full chromosome scale imputation still requires further improvement in phasing.
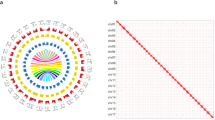
Visual representation of the phasing of 14 genome assemblies, with each circle representing a diploid genome assembly. Haplome 1 (inner) and Haplome 2 (outer) show parental origins (green/yellow), with intra-chromosomal color switches marking phase switches. Chromosomes are separated by black lines; homozygous segments >5 cM and genotype inconsistencies are shown as blue and pink bars, respectively.
Data availability
All raw data and assemblies are available on NCBI under BioProject PRJNA116848589.
Code availability
The bioinformatic tools and software used for the analyses were executed according to their respective published manuals. The versions and parameters of each bioinformatic tool and software that was used are listed within the methods section.
References
FAOSTAT. Food and Agriculture Organization of the United Nations. http://www.fao.org/faostat/en (2021).
Velasco, R. et al. The genome of the domesticated apple (Malus × domestica Borkh.). Nat. Publ. Gr. 42, 3–3 (2010).
Li, X. et al. Improved hybrid de novo genome assembly of domesticated apple (Malus x domestica). Gigascience 5 (2016).
Daccord, N. et al. High-quality de novo assembly of the apple genome and methylome dynamics of early fruit development. Nat. Genet. 49, 1099–1106 (2017).
Zhang, L. et al. A high-quality apple genome assembly reveals the association of a retrotransposon and red fruit colour. Nat. Commun. 10, 1–13 (2019).
Tian, Y. et al. Transposon insertions regulate genome-wide allele-specific expression and underpin flower colour variations in apple (Malus spp.). Plant Biotechnol. J. 20, 1285–1297 (2022).
Harvey, W. T. et al. Whole-genome long-read sequencing downsampling and its effect on variant-calling precision and recall. Genome Res 33, 2029–2040 (2023).
Li, H. & Durbin, R. Genome assembly in the telomere-to-telomere era. Nat. Rev. Genet. 25, 658–670 (2024). 2024 259.
Broggini, G. A. L. et al. Chromosome-scale de novo diploid assembly of the apple cultivar ‘Gala Galaxy’. bioRxiv 2020.04.25.058891, https://doi.org/10.1101/2020.04.25.058891 (2020).
Sun, X. et al. Phased diploid genome assemblies and pan-genomes provide insights into the genetic history of apple domestication. Nat. Genet. 52, 1423–1432 (2020).
Khan, A. et al. A phased, chromosome-scale genome of ‘Honeycrisp’ apple (Malus domestica). GigaByte 2022, gigabyte69 (2022).
Švara, A., Sun, H., Fei, Z. & Khan, A. Chromosome-level phased genome assembly of “Antonovka” identified candidate apple scab-resistance genes highly homologous to HcrVf2 and HcrVf1 on linkage group 1. G3 Genes|Genomes|Genetics 14 (2023).
Peng, H. et al. A haplotype-resolved genome assembly of Malus domestica ‘Red Fuji’. Sci. Data 11, 1–9 (2024).
Zhang, H. et al. A haplotype-resolved, chromosome-scale genome for Malus domestica Borkh. ‘WA 38’. G3 Genes|Genomes|Genetics 14 (2024).
Li, J. et al. The chromosome-level genome assembly of the dwarfing apple interstock Malus hybrid ‘SH6’, https://doi.org/10.1038/s41597-024-03405-x.
Su, Y. et al. Phased telomere-to-telomere reference genome and pangenome reveal an expansion of resistance genes during apple domestication. Plant Physiol 195, 2799–2814 (2024).
Li, W. et al. Near-gapless and haplotype-resolved apple genomes provide insights into the genetic basis of rootstock-induced dwarfing. Nat. Genet. 56, 505–516 (2024).
Jung, M. et al. The apple REFPOP—a reference population for genomics-assisted breeding in apple. Hortic. Res. 7, 189 (2020).
Zuffa, F. et al. Interannual Variation of Stomatal Traits Impacts the Environmental Responses of Apple Trees. Plant. Cell Environ. 48, 2478–2491 (2025).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. Gigascience 10, 1–4 (2021).
Cheng, H. et al. Haplotype-resolved assembly of diploid genomes without parental data. Nat. Biotechnol. 40, 1332–1335 (2022).
Alonge, M. et al. Automated assembly scaffolding using RagTag elevates a new tomato system for high-throughput genome editing. Genome Biol 23, 1–19 (2022).
Li, H. New strategies to improve minimap2 alignment accuracy. Bioinformatics 37, 4572–4574 (2021).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: A Cross-Platform and Ultrafast Toolkit for FASTA/Q File Manipulation. PLoS One 11, e0163962 (2016).
Cock, P. J. A. et al. Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423 (2009).
Linsmith, G. et al. Pseudo-chromosome–length genome assembly of a double haploid “Bartlett” pear (Pyrus communis L.). Gigascience 8, 1–17 (2019).
Xue, H. et al. Chromosome level high-density integrated genetic maps improve the Pyrus bretschneideri ‘DangshanSuli’ v1.0 genome. BMC Genomics 19, 1–13 (2018).
Shirasawa, K., Itai, A. & Isobe, S. Chromosome-scale genome assembly of Japanese pear (Pyrus pyrifolia) variety ‘Nijisseiki’. DNA Res. 28 (2021).
Gao, Y. et al. High-quality genome assembly of ‘Cuiguan’ pear (Pyrus pyrifolia) as a reference genome for identifying regulatory genes and epigenetic modifications responsible for bud dormancy. Hortic. Res. 8, 1–16 (2021).
Dong, X. et al. De novo assembly of a wild pear (Pyrus betuleafolia) genome. Plant Biotechnol. J. 18, 581–595 (2020).
Ou, C. et al. A de novo genome assembly of the dwarfing pear rootstock Zhongai 1. Sci. Data 6, 1–8 (2019).
Ireland, H. S. et al. The Gillenia trifoliata genome reveals dynamics correlated with growth and reproduction in Rosaceae. Hortic. Res. 8, 1–14 (2021).
Gabriel, L. et al. BRAKER3: Fully automated genome annotation using RNA-seq and protein evidence with GeneMark-ETP, AUGUSTUS and TSEBRA. bioRxiv Prepr. Serv. Biol., https://doi.org/10.1101/2023.06.10.544449 (2024).
Kuznetsov, D. et al. OrthoDB v11: annotation of orthologs in the widest sampling of organismal diversity. Nucleic Acids Res 51, D445–D451 (2023).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Res 47, D309–D314 (2019).
Manni, M., Berkeley, M. R., Seppey, M., Simão, F. A. & Zdobnov, E. M. BUSCO Update: Novel and Streamlined Workflows along with Broader and Deeper Phylogenetic Coverage for Scoring of Eukaryotic, Prokaryotic, and Viral Genomes. Mol. Biol. Evol. 38, 4647–4654 (2021).
Emms, D. M. & Kelly, S. OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biol 16, 1–14 (2015).
Emms, D. M. & Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol 20, 1–14 (2019).
Katoh, K., Misawa, K., Kuma, K. I. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30, 3059–3066 (2002).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Nakamura, T., Yamada, K. D., Tomii, K. & Katoh, K. Parallelization of MAFFT for large-scale multiple sequence alignments. Bioinformatics 34, 2490–2492 (2018).
R Core Team. R: A Language and Environment for Statistical Computing. https://www.r-project.org/ (2023).
Vanderzande, S., Peace, C. & Weg, E. van de. Whole genome sequence improvement with pedigree information and reference genotypic profiles, demonstrated in outcrossing apple. bioRxiv 2024.08.08.607141, https://doi.org/10.1101/2024.08.08.607141 (2024).
Bianco, L. et al. Development and validation of a 20K single nucleotide polymorphism (SNP) whole genome genotyping array for apple (Malus × domestica Borkh). PLoS One 9 (2014).
Di Pierro, E. A. et al. A high-density, multi-parental SNP genetic map on apple validates a new mapping approach for outcrossing species. Hortic. Res. 3 (2016).
Howard, N. et al. Collaborative project to identify direct and distant pedigree relationships in apple, https://doi.org/10.34894/VQ1DJA (2018).
Howard, N. P. et al. The use of shared haplotype length information for pedigree reconstruction in asexually propagated outbreeding crops, demonstrated for apple and sweet cherry. Hortic. Res. 8, 1–13 (2021).
Vanderzande, S. et al. High-quality, genome-wide SNP genotypic data for pedigreed germplasm of the diploid outbreeding species apple, peach, and sweet cherry through a common workflow. PLoS One 14, e0210928 (2019).
Camacho, C. et al. BLAST+: Architecture and applications. BMC Bioinformatics 10, 1–9 (2009).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939155.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939175.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938675.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938685.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938595.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938615.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939425.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939435.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939395.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939415.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938715.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938735.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939355.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939365.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939315.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939335.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939275.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939295.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939245.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939235.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939185.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939215.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939115.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939125.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939075.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939095.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939015.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938985.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938555.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938575.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938935.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938975.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938915.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938925.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939035.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052939045.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938535.1 (2025).
Yates, S. A. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_052938515.1 (2025).
NCBI Bioproject https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1168485 (2025).
NCBI Sequence Read Archive https://www.ncbi.nlm.nih.gov/sra/SRP560690 (2025).
Sanzol, J. Dating and functional characterization of duplicated genes in the apple (Malus domestica Borkh.) by analyzing EST data. BMC Plant Biol 10, 1–22 (2010).
Acknowledgements
We acknowledge the UK National Fruit Collection, Better3fruit (BE), Fresh Forward (NL) and Varicom (CH) for granting access to the plant material investigated in this study. Further we acknowledge the support of the REFPOP community, especially Dr. Michaela Jung and Dr. Andrea Patocchi for support in the genotype selection, Andrea Knauf and Luzia Lussi for support in sample collection, the HEST Informatic Support Group at ETH Zurich for computational resource and technical support. We thank Dr. Nick Howard for the assistance in providing SNP array data. This work was supported by ETH Research Grant (ETH-32 21-1) (FZ, GD), the Engage ETH Joint-Initative (SW), and the Pipfruit Technology Development Programme at The New Zealand Institute for Plant and Food Research Limited (CD).
Funding
Open access funding provided by Swiss Federal Institute of Technology Zurich.
Author information
Authors and Affiliations
Contributions
B.S., S.Y., F.Z., G.D. and G.A.L.B. conceived the study. F.Z. prepared the samples. G.A.L.B., S.Y., Y.C., S.V., C.D. analyzed the data. S.W., S.Y. and G.A.L.B. wrote the manuscript with assistance from C.D., F.Z., S.V. and B.S. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Watts, S., Yates, S., Vanderzande, S. et al. Haplotype-resolved chromosome-level genome assemblies of nineteen apple (Malus domestica Borkh.) cultivars. Sci Data 13, 258 (2026). https://doi.org/10.1038/s41597-026-06583-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06583-y