Abstract
The Ontology of Adverse Events (OAE) was launched in 2011 to define, standardize and integrate various adverse events (AEs) arising after medical interventions. The terminological framework of OAE has undergone consistent expansion since its inception, driven by its successful implementation in numerous AE investigations. In this paper, we document substantial ontological extensions addressing patient anatomic regions and clinical manifestations, encompassing symptoms, physical signs, and pathological processes. Current statistical analysis reveals that OAE has 10,829 formally defined terms with unique identifiers. Compared to the 3,088 ontology terms included in the last OAE publication in 2014, 7,741 new terms have been added to OAE, which represents significant progress of the ontology in clinical granularity and domain coverage. The OAE framework enables structured representation of critical determinants influencing clinical outcomes, including but not limited to administration routes, dosage parameters, and demographic variables such as patient age. Through its standardized semantic architecture, OAE provides an integrative platform for cross-disciplinary analysis of AE patterns, etiological factors, and outcome trajectories in clinical interventions.
Similar content being viewed by others
Background & Summary
The Ontology of Adverse Events (OAE) is a community-based biomedical ontology that is designed to be interoperable and logically well-formed and to incorporate accurate representations of adverse events (AEs) associated with medical interventions (e.g., drug administration and vaccination). As a comprehensive resource of AEs, OAE enables computational inference and sophisticated algorithms that allow AE data to be integrated with other data (e.g., phenotypic data and genomic data) and then support combined AE profile and pathogenetic mechanism analyses. Nowadays, OAE is available without restriction for public view at various ontology linked data resources including OntoBee1 (default OAE linked server), BioPortal2, OLS3, and AberOWL4.
To improve representation and organization of AE information, some vocabulary resources have been developed and widely used, including the Medical Dictionary for Regulatory Activities (MedDRA)5, the Common Terminology Criteria for Adverse Events (CTCAE)6, and the World Health Organization (WHO)’s Adverse Reaction Terminology (WHO-ART)7. The OAE differs from these available AE coding vocabularies in several crucial ways. First, the OAE has provided more logically well-formed definitions and a more structured classification of AEs than any other coding vocabulary. Second, the OAE is not a simple terminology, but rather a full community-based biomedical ontology that follows the Web Ontology Language (OWL) and the Open Biological and Biomedical Ontologies (OBO) Foundry principles8, thus serving as a computational resource that supports computer-assisted reasoning and analysis. Third, OAE represents AEs based on patient anatomic regions and clinical outcomes, as well as taking into account other elements (e.g., patient age, patient gender, and current illness) of the process that leads from initial medical intervention to subsequent outcomes, all of which assist in realizing precision medicine.
Generally speaking, OAE has experienced the following three stages of development. In the first stage, OAE was initially developed as an ongoing realism-based effort that aimed to reduce the confusion in existing AE terminologies and was subsequently published at the 2nd International Conference on Biomedical Ontology (ICBO-2011)9. In the second stage, OAE redefined the key term ‘adverse event’ (OAE_0000001) as a process subsequent to a medical intervention while not assuming any causal association between them. The term ‘causal adverse event’ (OAE_0000003) was newly created to represent the existence of data of a certain sort, namely data establishing a causal association between adverse events and medical interventions10. Within the OAE framework, a clear distinction is made between an adverse event and a causal adverse event, where the latter is defined as a specific subtype of the former. This change made the OAE more robust as a representation of the domain of AE reporting. In the third stage, OAE-based methods are widely used in the field of medical informatics and play an increasingly important role in representing, classifying, and analyzing the AE information. The OAE continuously evolves and improves through the integration of new terms derived from its practical applications.
This study systematically examines the structural and functional evolution of the OAE over the past decade, with a dual focus on delineating major ontological advancements and conducting a comprehensive review and summary of existing OAE-based research. Furthermore, we also report on recent efforts to translate the OAE into Chinese and introduce it into the monitoring and analysis for adverse events following immunization (AEFI) in China.
Extend the breadth and scope of OAE
The OAE is a community-driven biomedical ontology that ontologically organizes knowledge domains related to adverse medical occurrences. Its primary objective is to systematically represent AE manifestations across diverse anatomical locations and to logically classify adverse outcomes into distinct categories such as symptoms, clinical signs, and pathological processes (Fig. 1). As of 31 December 2024, the OAE has contained 10,589 AE-related terms (Fig. 2 and Table 1), reflecting an annual growth rate of 13.9% since the last Journal of Biomedical Semantics article published in 201410.
General OAE design pattern.
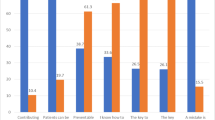
Evolution of OAE terminology. This visualization shows the distribution of top-level terms within the OAE. The dark-blue portion represents new terms introduced following the initial publication10, corresponding to the period from 20 April 2014 to 31 December 2024.
AE data acquisition
The newly added terms were derived from mining the existing knowledge bases of the corresponding entries, including:
-
1)
Clinical AE reports in the Vaccine Adverse Events Reporting System (VAERS) and the Food and Drug Administration (FDA) Adverse Events Reporting System (FAERS). VAERS is a national early warning system to detect possible safety problems in U.S.-licensed vaccines. It accepts and analyzes reports of AEs (possible side effects) after a person has received a vaccination. Anyone can report an AE to VAERS11. The FAERS database contains AE reports, medication error reports and product quality complaints resulting in AEs that were submitted to FDA. The database is designed to support the FDA’s post-marketing safety surveillance program for drug and therapeutic biologic products12.
-
2)
Symptoms, signs, and adverse drug reactions that are cataloged within the MedDRA terminology system. MedDRA is a rich and highly specific standardized medical terminology developed by the International Conference on Harmonisation (ICH) to facilitate sharing of regulatory information internationally for medical products (e.g., pharmaceuticals, vaccines, and drug-device combination products) used by humans5. We have referenced MedDRA in our OAE development by cross-referencing related MedDRA identifiers (Fig. 3). The data models of AE analysis provided by the Clinical Data Interchange Standards Consortium (CDISC) were also referenced13.
Fig. 3 Demonstration of standardized annotation for a new term. The annotation for ‘acute abdomen AE’ (OAE_0004600) is presented as an example, following the defined annotation schema of the OAE.
-
3)
Adverse events identified from OAE-based scientific researches and peer-reviewed journal articles. Numerous pharmacovigilance investigations have employed OAE methodologies for systematic analysis. Some AEs revealed in these studies but not yet included in the OAE terminology system would be identified when OAE-based hierarchical classification was conducted. For example, in our researches on AEs associated with hepatitis A/B and brucellosis vaccines14,15, we successfully identified and subsequently incorporated more than 70 novel AE terms into OAE. To supplement and validate term definitions, we drew on AE information from journal articles and expert consultations, particularly for rare events, which was then incorporated into the OAE as supporting references.
AE data annotation
The precise annotation of terminology constitutes a critical element in quality control during ontology development. The OAE provides annotations for AE terms defined by two major patterns:
-
1)
Import terms that are available in established ontologies. To ensure alignment with the OBO Foundry Principles and facilitate ontology term reuse and interoperability16, the existing ontological terms are systematically integrated into the OAE framework through direct importation mechanisms (Table 1). For example, the complete ontological architecture of the Basic Formal Ontology (BFO) has been imported into the OAE as its foundational upper-level ontology17, demonstrating methodological consistency with established ontological integration protocols. The annotations and original identifiers (IDs) of imported individual ontology terms are preserved through implementation of the OntoFox software18, while maintaining the semantic relationships between these entity terms and their corresponding property associations.
-
2)
Manual annotation of new AE terms. As mentioned above, the annotation comprehensiveness of AE terms should be maximized following established epistemological principles. Figure 4 outlines the foundational annotation scheme governing newly incorporated AE terms within the OAE framework, including label (required), definition (required), definition source (optional), see also (optional), term editor (required), alternative name (optional), subclass of (required), and occurs in (required). Furthermore, to facilitate communication and utilization of OAE among Chinese science and technology professionals, we have developed a “Chinese Translation” annotation option and translated the newly added AE terms into Chinese (Fig. 3). It is noteworthy that some imported AE terms remain undefined, requiring systematic redefinition of these terms by our research team. The definitions of AE terms within OAE are predominantly sourced from established knowledge bases and scholarly repositories, including but not limited to domain-specific databases (e.g., PubMed) and collaboratively curated knowledge platforms (e.g., Wikipedia). For AE-specific terms in OAE, new identifiers incorporating the “OAE_” prefix followed by a seven-digit auto-incremental numerical sequence were generated. For example, the term ‘acute abdomen AE’ is assigned the identifier OAE_0004600, with its complete metadata accessible through the OntoBee ontology browser via its Internationalized Resource Identifier (IRI)1: http://purl.obolibrary.org/obo/OAE_0004600.
Fig. 4 Annotation schema and content of OAE terminology.
As presented in Table 1, OAE comprises 10,829 formally defined representational units. The OAE-specific terms include 9,397 distinct classes alongside 13 ontological properties, specifically categorized as eight object properties, two datatype properties, and three annotation properties. Comparative analysis reveals substantial terminological expansion, with 7,743 novel terms (representing a 251% increase) having been incorporated since the previous 2014 publication documenting 3,086 ontology terms. This quantitative growth demonstrates marked ontological development, reflecting both conceptual granularity enhancement and domain coverage extension.
Interoperability applications of OAE with other knowledge bases
In conjunction with other standardized terminologies and classification systems, the OAE and its user-friendly annotation framework enable semantic interoperability within the domain of digital medicine. The Disease Specific Ontology of Adverse Events (DSOAE) has been developed to understand and represent disease-intervention specific AEs for a specific domain such as chronic kidney disease (CKD) through the extension and adaptation of general ontologies (e.g., OAE) by incorporating domain-specific information and usage requirements19. With the support of DSOAE, a data-driven knowledge base for CKD-specific AEs has been established, and successfully applied in data analysis and integration.
Drug adverse events (DAEs) represent a critical focus area in pharmacovigilance, and substantial progress has been made in the development of domain-specific knowledge bases to systematically document these phenomena. Notably, ontology-driven frameworks have emerged as pivotal tools in this domain. For example, OAE-based DAE-specific ontologies, such as the Ontology of Drug Neuropathy Adverse Events (ODNAE) and the Ontology of Cardiovascular Drug Adverse Events (OCVDAE)20,21, have been developed to standardize the characterization of drug-associated neuropathy-inducing and cardiovascular-related DAEs, respectively. To logically represent the complex relations among drugs, drug components, mechanisms of action, AEs, age, diseases, and other related factors, an ontology design pattern was developed and implemented in the community-driven open-source Ontology of Drug Adverse Events (ODAE)22. The ODAE expands both the OAE and the National Drug File-Reference Terminology (NDF-RT), providing a generic representation of DAEs across varying conditions, which enables the systematic querying of scientific questions.
Since its establishment in 2011, the annotated terms and hierarchical architecture of OAE have been extensively cross-referenced by diverse domain-specific ontologies and databases23,24,25,26. The development and application of OAE-based ontology extensions, coupled with the sharing and integration of interoperable data with other ontologies or knowledge repositories, have collectively contributed to enhancing the depth, coverage, and sophistication of the OAE system.
Usage
The OAE has been extensively applied in research on adverse events associated with medical interventions. As of December 31, 2024, a total of 148 studies indexed in Google Scholar have cited the OAE methodology. In this section, we systematically categorize and review these studies, with a focus on representative cases to illustrate their applications.
OAE-based AE studies for vaccines
Vaccines are mostly preventive measures administered to healthy individuals for disease prevention, consequently imposing higher safety requirements and eliciting heightened public sensitivity towards associated adverse events. As mentioned above, the VAERS is a pioneering, comprehensive, and widely utilized surveillance system for monitoring vaccine adverse events (VAEs). The first OAE-based VAEs study was performed in 201227, which developed a Combinatorial, Ontology-based Detection of AE (CODAE) method to analyze the AE data associated with killed and live influenza vaccines sourced from the VAERS database. In this study, the statistically enriched VAEs were classified and analyzed using the hierarchical structure of OAE, MedDRA, and SNOMED-CT respectively. The OAE method provided better classification results than the other two methods.
The advantages of utilizing the OAE-based semantic web representation and data analysis approach were demonstrated to be remarkably effective in subsequent VAE studies, which incorporated more sophisticated statistical algorithms. For example, Xie et al. conducted an OAE-based systematic analysis of AE profiles associated with Hepatitis A/B vaccines, employing a logistic regression model accompanied with Markov Chain Monte Carlo (MCMC) sampling, and then identified 13 AEs (e.g., hepatosplenomegaly) attributable to the synergistic effects of vaccine-vaccine interactions (VVIs)14. This study represents the first report to analyze VVIs using clinically reported AE case data based on an ontological approach integrated with statistical modeling. Zi et al. adopted the CODAE method to evaluate AE profiles of three licensed human papillomavirus (HPV) vaccines (i.e., Cervarix, Gardasil, and Gardasil 9) using the reported data from VAERS28. Through application of a gender/age-stratified Gamma-Poisson Shrinker (GPS) model, 46 statistically significant serious adverse events (SAEs) were found to be associated with at least one HPV vaccine. The SAE profile was represented and classified via the OAE to enable mechanistic analysis.
It is noted that the findings derived from these AE surveillance studies should be interpreted with caution regarding causal attribution between vaccines and AEs owing to the methodological constraints inherent in spontaneously reported AE data29. Future research requires rigorous design and implementation of population-based epidemiological studies alongside controlled clinical trials to establish causal relationships between vaccines and specific AEs, with particular emphasis on characterizing the occurrence dynamics of SAEs.
OAE-based AE studies for drugs
In contrast to vaccines, which are typically administered to healthy individuals as prophylactic measures, the majority of drugs are prescribed to patients with pre-existing medical conditions. Responses to drug treatments, particularly with respect to AEs, may be significantly influenced by both the underlying disease pathophysiology and its clinical progression. Consequently, the systematic monitoring and analytic evaluation of drug adverse events (DAEs) present substantially greater methodological challenges compared to vaccine safety surveillance.
In the USA, the DAEs are monitored by the FAERS system, which mandates the application of the MedDRA as a standardized medical terminology system for classification and coding of reported DAEs. The FDA has been actively exploring the application of ontology-based systems pharmacology to enhance mechanistic analysis of DAEs and improve predictive toxicology assessments30,31. There are a number of specific projects and/or activities that have been launched or are currently in progress, such as creation of a standardized ontology for the universe of “molecular toxic targets”32, and development of a multiple-scale network linking both the molecular targets and organ toxicities to clinical drug toxicities using the MedDRA ontology25.
Given that the application of OAE in vaccine research has demonstrated promising interpretable outcomes, our research team has prioritized extending this analytical framework to the more complex domain of DAE analysis. A scalable auto-assistance platform for extraction of potential DAEs from medical case reports has been built by Gurulingappa et al.33. It relies on a high-quality corpus that was manually annotated using the OAE-driven methodology, and consequently has the ability to capture valid, under-reported, and novel potential DAEs not yet present in product leaflets. Sarntivijai et al. established a semantic framework to link biological mechanisms to phenotypes of DAEs by combining OAE with MedDRA in FAERS data analysis34. Through MedDRA-OAE term mapping, they integrated MedDRA-based DAEs associated with tyrosine kinase inhibitors (TKIs) and monoclonal antibodies (mAbs) into the OAE terminology system. This enabled subsequent semantic mapping to established biomedical ontologies, including the Human Phenotype Ontology (HP) for clinical manifestations35, the Uberon Anatomy Ontology (UBERON) for anatomical localization36, and the Gene Ontology (GO) for molecular functional annotations37. This multi-ontology integration approach effectively bridged pharmacological adverse event documentation with fundamental biological understanding. Hur et al. developed an ontology-driven literature mining system to detect DAEs from drug labels and investigate mechanistic pathways of neuropathy-inducing drugs (NIDs)38. Their OAE-based analytical approach further found that the identified 43 benzimidazole-related DAEs exhibited multisystem distribution patterns, with predominant manifestations observed in the behavioral and neurological, digestive, skin, and immune systems. This investigation substantiates that the integration of ontological text mining with network pharmacology constitutes a robust methodological framework for identifying drug-class-specific DAEs and elucidating their multiscale biological interconnections.
Advancing precision medicine implementation
It is widely acknowledged that precision medicine has significantly transformed modern medical practice. This innovative paradigm, originally coined to describe therapeutic strategies (e.g., routes of drug administration and vaccination) tailored to individual patients based on their unique genetic profiles, biomarker signatures, phenotypic characteristics, and psychosocial determinants, fundamentally differs from conventional approaches targeting patients with similar clinical manifestations39. The foremost and most formidable challenge in implementing precision medicine involves establishing a robust multidisciplinary framework and an integrated knowledge base. Such infrastructure must effectively bridge the translational gap between basic research discoveries and clinical applications, necessitating unprecedented collaboration among biomedical researchers, diagnostic laboratories, healthcare providers, and patient communities40.
Arguably, ontologies as formal knowledge representation systems serve as frameworks for integrating and analyzing heterogeneous clinical datasets alongside scientific evidence derived from diverse healthcare stakeholders. These structured vocabularies enable systematic data harmonization across disparate sources, crucially facilitating the standardization and interoperability of domain-specific data resources required for advancing precision medicine initiatives. Haendel et al. conducted a seminal investigation of ontologies, including the OAE, demonstrating their critical role in computational reasoning systems for enabling granular patient stratification41. Their research substantiated the utility of these structured knowledge systems across three key domains: diagnostic precision optimization, clinical care pathway personalization, and translational research methodologies. They conclusively established that ontologies facilitate multidimensional patient characterization through standardized semantic interoperability, thereby enabling phenotype-driven classification at clinically actionable resolutions. Furthermore, Xie et al. performed a systematic review examining precision vaccinology—an emerging discipline applying precision medicine principles to vaccine development—and its potential enhancement by ontological engineering, such as Vaccine Ontology (VO), OAE, and Ontology of Vaccine Adverse Events (OVAE)42. Based on these ontologies, they formulated a novel ontological framework strategy to advance precision vaccinology, proposing its systematic integration into vaccine target identification, mechanistic modeling of immune responses, and AE prediction during a vaccination campaign.
The significance of OAE in the surveillance of AEs associated with medical interventions in China
The current AE reporting systems for medical interventions in China exhibit limitations in terminology classification. Exemplified by the National Adverse Events Following Immunization Information System (CNAEFIS)43, it categorizes adverse events following immunization (AEFI) into seven broad types based on potential causative factors: common AEs, abnormal AEs, vaccine quality events, program errors, coincidental events, psychogenic reactions, and unexplained reactions. In addition, cases that cannot be definitively classified under these predefined categories are arbitrarily designated as “other” AEs. This oversimplified approach has resulted in the accumulation of substantial AEFI data without logical categorization or analytical coherence. In reality, AEFI manifest through diverse clinical presentations and etiological mechanisms. While partial adoption of ICD-9/10 coding standards has been implemented, the existing framework demonstrates insufficient systematic integration of diagnostic terminology, thereby compromising its capacity to support rigorous post-hoc analyses of AEs. This critical infrastructure gap underscores the urgent need to establish a standardized medical terminology classification system to enhance AE surveillance research.
Over the past decade, ontological engineering has evolved into a pivotal methodological paradigm for biomedical big data research and precision medicine applications. Its demonstrated capabilities in semantic integration and knowledge representation have attracted sustained attention from the Chinese scientific community. In 2018, the National Population and Health Science Data Sharing Platform spearheaded the formation of the China Biomedical Ontology Consortium (OntoChina, http://www.ontochina.org/). This initiative aims to construct ontology repositories for Chinese biomedical big data research through the adoption of international best practices in ontological engineering and cross-domain collaboration, while promoting ontological applications in national health informatization. Of particular significance is the foundational work by Dr. Yan Zhu’s research team at the China Academy of Chinese Medical Sciences, which completed the Chinese localization of the BFO44, thereby establishing a standardized upper-level framework for developing Chinese-language biomedical ontologies.
The OAE has been successfully applied in multiple studies analyzing medical intervention-associated AEs. Notably, our research team has methodologically integrated standardized Chinese terminological equivalents during OAE’s iterative development cycles. Building upon recent advancements in the localization of the BFO, the comprehensive sinicization of OAE, encompassing both granular terminological mapping and logical relationship adaptation, has now attained technical viability. In recent years, an increasing number of Chinese researchers have adopted the OAE as a standardized terminological framework to advance related studies. For instance, Li et al. utilized the OAE to perform a comprehensive analysis of DAE case reports associated with domestic anti-infection drugs, as illustrated in Fig. 545. Their study delineated the epidemiological characteristics of these DAEs, including demographic trends, drug administration routes, and severity profiles. These findings offer actionable recommendations to optimize evidence-based clinical practices for anti-infection therapies. By strategically utilizing OAE’s open-source architecture in conjunction with established ontological engineering platforms (e.g., OntoBee for ontology browsing1, OntoFox for term extraction18, and OntoRat for collaborative editing46), this initiative enables semantic interoperability between OAE and existing Chinese biomedical ontology systems. Consequently, the systematic implementation of OAE in China would not only optimize the technical infrastructure for medical intervention safety surveillance but also catalyze paradigm-shifting advancements in causality analysis frameworks for medical intervention-adverse event investigations.
OAE-based analytical framework for anti-infection DAE case reports in China.
Conclusion
In recent years, the terminology system of OAE has undergone substantial updates and refinement through the collaborative efforts of our research team combined with expert contributions from relevant domains. We will broaden our efforts to extend the OAE terminology into multiple specialized subontologies, particularly those addressing drug- and vaccine-related adverse events. Concurrently, we will strengthen interoperability frameworks with established ontologies, with the goal of streamlining the creation of novel domain-specific knowledge systems.
Data availability
The OAE is hosted in the OBO Foundry library (https://obofoundry.org/ontology/oae.html). Public access is provided through both the NCBO BioPortal (https://bioportal.bioontology.org/ontologies/OAE) and OntoBee (http://www.ontobee.org/ontology/OAE) platforms, which support browsing, querying, and downloading of the ontology.
Code availability
The source code of OAE is freely available at https://github.com/OAE-ontology/OAE.
References
Ong, E. et al. Ontobee: A linked ontology data server to support ontology term dereferencing, linkage, query and integration. Nucleic Acids Res 45, D347–D352, https://doi.org/10.1093/nar/gkw918 (2017).
Salvadores, M., Alexander, P. R., Musen, M. A. & Noy, N. F. BioPortal as a Dataset of Linked Biomedical Ontologies and Terminologies in RDF. Semant Web 4, 277–284 (2013).
Côté, R. G., Jones, P., Apweiler, R. & Hermjakob, H. The Ontology Lookup Service, a lightweight cross-platform tool for controlled vocabulary queries. BMC Bioinformatics 7, 97, https://doi.org/10.1186/1471-2105-7-97 (2006).
Hoehndorf, R., Slater, L., Schofield, P. N. & Gkoutos, G. V. Aber-OWL: a framework for ontology-based data access in biology. BMC Bioinformatics 16, 26, https://doi.org/10.1186/s12859-015-0456-9 (2015).
Brown, E. G., Wood, L. & Wood, S. The medical dictionary for regulatory activities (MedDRA). Drug Saf 20, 109–117, https://doi.org/10.2165/00002018-199920020-00002 (1999).
Atkinson, T. M. et al. The association between clinician-based common terminology criteria for adverse events (CTCAE) and patient-reported outcomes (PRO): a systematic review. Supportive care in cancer: official journal of the Multinational Association of Supportive Care in Cancer 24, 3669–3676, https://doi.org/10.1007/s00520-016-3297-9 (2016).
Alecu, I., Bousquet, C., Mougin, F. & Jaulent, M. C. Mapping of the WHO-ART terminology on Snomed CT to improve grouping of related adverse drug reactions. Stud Health Technol Inform 124, 833–838 (2006).
Smith, B. et al. The OBO Foundry: coordinated evolution of ontologies to support biomedical data integration. Nat Biotechnol 25, 1251–1255, https://doi.org/10.1038/nbt1346 (2007).
He, Y., Xiang, Z., Sarntivijai, S., Toldo, L. & Ceusters, W. AEO: A Realism-Based Biomedical Ontology for the Representation of Adverse Events. ICBO 2011 (2011).
He, Y. et al. OAE: The Ontology of Adverse Events. J Biomed Semantics 5, 29, https://doi.org/10.1186/2041-1480-5-29 (2014).
Shimabukuro, T. T., Nguyen, M., Martin, D. & DeStefano, F. Safety monitoring in the Vaccine Adverse Event Reporting System (VAERS). Vaccine 33, 4398–4405, https://doi.org/10.1016/j.vaccine.2015.07.035 (2015).
Giunchi, V., Fusaroli, M., Hauben, M., Raschi, E. & Poluzzi, E. Challenges and Opportunities in Accessing and Analysing FAERS Data: A Call Towards a Collaborative Approach. Drug Saf 46, 921–926, https://doi.org/10.1007/s40264-023-01345-w (2023).
Burnstead, B. & Furlan, G. Unifying drug safety and clinical databases. Curr Drug Saf 8, 56–62, https://doi.org/10.2174/1574886311308010008 (2013).
Xie, J., Zhao, L., Zhou, S. & He, Y. Statistical and Ontological Analysis of Adverse Events Associated with Monovalent and Combination Vaccines against Hepatitis A and B Diseases. Sci Rep 6, 34318, https://doi.org/10.1038/srep34318 (2016).
Xie, J. et al. Ontology-Based Meta-Analysis of Animal and Human Adverse Events Associated With Licensed Brucellosis Vaccines. Front Pharmacol 9, 503, https://doi.org/10.3389/fphar.2018.00503 (2018).
Jackson, R. et al. OBO Foundry in 2021: operationalizing open data principles to evaluate ontologies. Database (Oxford) 2021, https://doi.org/10.1093/database/baab069 (2021).
Smith, B. & Ceusters, W. Ontological realism: A methodology for coordinated evolution of scientific ontologies. Appl Ontol 5, 139–188, https://doi.org/10.3233/AO-2010-0079 (2010).
Xiang, Z., Courtot, M., Brinkman, R. R., Ruttenberg, A. & He, Y. OntoFox: web-based support for ontology reuse. BMC Res Notes 3, 175, https://doi.org/10.1186/1756-0500-3-175 (2010).
Kang, Y., Fink, J. C., Doerfler, R. & Zhou, L. Disease Specific Ontology of Adverse Events: Ontology extension and adaptation for Chronic Kidney Disease. Comput Biol Med 101, 210–217, https://doi.org/10.1016/j.compbiomed.2018.08.024 (2018).
Guo, A. et al. Ontology-based collection, representation and analysis of drug-associated neuropathy adverse events. J Biomed Semantics 7, 29, https://doi.org/10.1186/s13326-016-0069-x (2016).
Wang, L. et al. Ontology-based systematical representation and drug class effect analysis of package insert-reported adverse events associated with cardiovascular drugs used in China. Sci Rep 7, 13819, https://doi.org/10.1038/s41598-017-12580-4 (2017).
Yu, H. et al. ODAE: Ontology-based systematic representation and analysis of drug adverse events and its usage in study of adverse events given different patient age and disease conditions. BMC Bioinformatics 20, 199, https://doi.org/10.1186/s12859-019-2729-1 (2019).
Xiao, Y. et al. CIDO-COVID-19: An Ontology for COVID-19 Based on CIDO. Annu Int Conf IEEE Eng Med Biol Soc 2021, 2119–2122, https://doi.org/10.1109/EMBC46164.2021.9629555 (2021).
Herrero-Zazo, M., Segura-Bedmar, I., Hastings, J. & Martinez, P. DINTO: Using OWL Ontologies and SWRL Rules to Infer Drug-Drug Interactions and Their Mechanisms. J Chem Inf Model 55, 1698–1707, https://doi.org/10.1021/acs.jcim.5b00119 (2015).
Zaman, S., Sarntivijai, S. & Abernethy, D. R. Use of Biomedical Ontologies for Integration of Biological Knowledge for Learning and Prediction of Adverse Drug Reactions. Gene Regul Syst Bio 11, 1177625017696075, https://doi.org/10.1177/1177625017696075 (2017).
Hogan, W. R. et al. Therapeutic indications and other use-case-driven updates in the drug ontology: anti-malarials, anti-hypertensives, opioid analgesics, and a large term request. J Biomed Semantics 8, 10, https://doi.org/10.1186/s13326-017-0121-5 (2017).
Sarntivijai, S. et al. Ontology-based combinatorial comparative analysis of adverse events associated with killed and live influenza vaccines. PLoS One 7, e49941, https://doi.org/10.1371/journal.pone.0049941 (2012).
Zi, W., Yang, Q., Su, J., He, Y. & Xie, J. OAE-based data mining and modeling analysis of adverse events associated with three licensed HPV vaccines. Heliyon 8, e11515, https://doi.org/10.1016/j.heliyon.2022.e11515 (2022).
Varricchio, F. et al. Understanding vaccine safety information from the Vaccine Adverse Event Reporting System. Pediatr Infect Dis J 23, 287–294, https://doi.org/10.1097/00006454-200404000-00002 (2004).
Abernethy, D. R., Woodcock, J. & Lesko, L. J. Pharmacological mechanism-based drug safety assessment and prediction. Clin Pharmacol Ther 89, 793–797, https://doi.org/10.1038/clpt.2011.55 (2011).
Bai, J. P. & Abernethy, D. R. Systems pharmacology to predict drug toxicity: integration across levels of biological organization. Annu Rev Pharmacol Toxicol 53, 451–473, https://doi.org/10.1146/annurev-pharmtox-011112-140248 (2013).
Sorger, P. K. et al. Quantitative and systems pharmacology in the post-genomic era: new approaches to discovering drugs and understanding therapeutic mechanisms. An NIH white paper by the QSP workshop group (2011).
Gurulingappa, H., Mateen-Rajput, A. & Toldo, L. Extraction of potential adverse drug events from medical case reports. J Biomed Semantics 3, 15, https://doi.org/10.1186/2041-1480-3-15 (2012).
Sarntivijai, S. et al. Linking MedDRA(®)-Coded Clinical Phenotypes to Biological Mechanisms by the Ontology of Adverse Events: A Pilot Study on Tyrosine Kinase Inhibitors. Drug Saf 39, 697–707, https://doi.org/10.1007/s40264-016-0414-0 (2016).
Köhler, S. et al. The Human Phenotype Ontology in 2021. Nucleic Acids Res 49, D1207–D1217, https://doi.org/10.1093/nar/gkaa1043 (2021).
Mungall, C. J., Torniai, C., Gkoutos, G. V., Lewis, S. E. & Haendel, M. A. Uberon, an integrative multi-species anatomy ontology. Genome Biol 13, R5, https://doi.org/10.1186/gb-2012-13-1-r5 (2012).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 25, 25–29, https://doi.org/10.1038/75556 (2000).
Hur, J., Özgür, A. & He, Y. Ontology-based literature mining and class effect analysis of adverse drug reactions associated with neuropathy-inducing drugs. J Biomed Semantics 9, 17, https://doi.org/10.1186/s13326-018-0185-x (2018).
Jameson, J. L. & Longo, D. L. Precision medicine–personalized, problematic, and promising. N Engl J Med 372, 2229–2234, https://doi.org/10.1056/NEJMsb1503104 (2015).
Aronson, S. J. & Rehm, H. L. Building the foundation for genomics in precision medicine. Nature 526, 336–342, https://doi.org/10.1038/nature15816 (2015).
Haendel, M. A., Chute, C. G. & Robinson, P. N. Classification, Ontology, and Precision Medicine. N Engl J Med 379, 1452–1462, https://doi.org/10.1056/NEJMra1615014 (2018).
Xie, J., Zi, W., Li, Z. & He, Y. Ontology-based Precision Vaccinology for Deep Mechanism Understanding and Precision Vaccine Development. Curr Pharm Des 27, 900–910, https://doi.org/10.2174/1381612826666201125112131 (2021).
Liu, D. et al. Surveillance of adverse events following immunization in China: Past, present, and future. Vaccine 33, 4041–4046, https://doi.org/10.1016/j.vaccine.2015.04.060 (2015).
Zhu, Y., Zheng, J., Li, X., Yang, X. & He, Y. Introduction to basic formal ontology and its Chinese version. J Med Inform 42, 24–28 (2021).
Li, M., Cao, Y., Zhang, H., He, Y. & Wang, L. Ontology-based analysis of adverse drug reactions reports associated with domestic anti-infection drugs. Chinese Journal of Pharmacoepidemiology 26, 5 (2017).
Xiang, Z., Zheng, J., Lin, Y. & He, Y. Ontorat: automatic generation of new ontology terms, annotations, and axioms based on ontology design patterns. J Biomed Semantics 6, 4, https://doi.org/10.1186/2041-1480-6-4 (2015).
Acknowledgements
We would like to thank Barry Smith, Mathias Brochhausen, Darrell R. Abernethy, Elizabeth Blair, Izabela Birsanescu, Keith Burkhart, Luca Toldo, Sydni Joubran, Yu Lin, Abra Guo, Bingjian Yang, Desikan Jagannathan, Edison Ong, Kelly Yang, Kevin Mo, Liwei Wang, Meiu Wong, Noemi Garg, Qingping Liu, Rebecca Racz, Shelley Zhang, Sirarat Sarntivijai, Zuoshuang Xiang, Ling Wan, David Ameriguian, and Jessica DeGuise for their valuable guidance and contributions. JX received support from the STI 2030-Major Projects under Grant (Grant No. 2022ZD0211400) and the Natural Science Foundation of Chongqing of China (Grant No. CSTC2018JCYJAX0243). SL received support from the Natural Science Foundation of Chongqing of China (Grant No. CSTB2022NSCQ-MSX1234).
Author information
Authors and Affiliations
Contributions
Conception, design, manuscript preparation: J.X., Y.H., C.P., Q.Y.; Participant recruitment: J.X., Y.H.; Data acquisition: C.P., Q.Y., X.Z., S.L., J.X.; Processing pipeline: C.P., Q.Y., X.Z., S.L., J.X.; Data processing: C.P., Q.Y., X.Z., S.L., J.X.; Quality control: J.X., S.L., Y.H.; All authors provided feedback and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Pan, C., Yang, Q., Zhang, X. et al. The Ontology of Adverse Events in 2025. Sci Data 13, 259 (2026). https://doi.org/10.1038/s41597-026-06584-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06584-x