Abstract
Dynamic Contrast-Enhanced Magnetic Resonance Imaging (DCE-MRI) is essential for monitoring breast cancer treatment response, yet deep learning progress is limited by the lack of standardized, multi-center datasets. We present BreastDCEDL, a deep learning-ready dataset comprising pretreatment 3D DCE-MRI scans from 2,070 patients across three cohorts: I-SPY1 (n = 172), I-SPY2 (n = 982), and Duke (n = 916). Raw DICOM files were converted to standardized 3D NIfTI format, preserving signal integrity and spatial resolution. The dataset includes unified tumor annotations—binary masks for I-SPY cohorts and bounding boxes for Duke—and harmonized clinical metadata. Pathologic complete response (pCR) data are available for 1,452 patients (100% of I-SPY1/I-SPY2, 32.5% of Duke). Clinical biomarkers include HR status (64.1% positive) and HER2 status (22.2% positive) for over 99% of patients. Predefined train-validation-test splits (1,532/268/270) maintain balanced distributions. BreastDCEDL fills a critical gap in public imaging resources, supporting development of advanced models for pCR prediction and other breast cancer analyses. The dataset is designed for broad reuse and is accompanied by processing tools to support community-driven AI research in oncology.
Similar content being viewed by others
Background & Summary
Breast cancer is a leading cause of cancer death worldwide. Early detection and accurate treatment monitoring are crucial to better patient outcomes. An important clinical decision after diagnosis is the choice between immediate surgery or neoadjuvant chemotherapy (NAC) before surgery. NAC aims to reduce tumor size, enhancing the potential for surgical success. The optimal outcome is the complete pathologic response (pCR), defined as the absence of residual invasive disease following therapy, confirmed by postsurgical biopsy. Accurate prediction of pCR is critical because it strongly correlates with improved long-term survival and can guide personalized treatment decisions. Pre-treatment pCR probability estimates could support clinical decision-making in several ways: risk stratification for therapy intensification or de-escalation, clinical trial enrichment by identifying patients most likely to benefit from novel agents, and early identification of likely non-responders who may benefit from alternative treatment strategies. Prior work suggests that MRI acquired during or after NAC can outperform pre-treatment imaging for pCR prediction, and that combining imaging with clinicopathologic variables further improves performance1. BreastDCEDL includes harmonized clinical metadata to support multimodal analyses; conversion of mid-treatment I-SPY timepoints is planned for future releases. Dynamic contrast-enhanced magnetic resonance imaging (DCE-MRI) is a principal imaging modality for the evaluation of breast cancer. The technique involves acquiring sequential 3D scans before and after the administration of contrast agents, typically analyzed at three time points: baseline before contrast, early post-contrast (maximum enhancement) and late post-contrast (washout dynamics), as shown in Fig. 1. These acquisitions enable visualization of abnormal tumor vascularity—characterized by increased vessel density, permeability, and disorganized structure—that distinguishes malignant tissues from normal breast parenchyma2,3,4. Deep learning applied to breast DCE-MRI analysis has shown promise, but faces limitations. Multiple systematic reviews5,6,7,8 have identified critical barriers, including insufficient access to large-scale datasets, inconsistent evaluation protocols, and inadequate external validation. These works span from technical deep learning reviews (Houssein6, Wei7) to modality-specific MRI analyses (Gullo5, LoGullo8), collectively highlighting the need for standardized datasets and reproducible benchmarks to advance clinical translation. Although convolutional neural networks have shown promising results through multiphase fusion9, multimodal integration10,11, and multi-institutional validation12, most investigations remain constrained by limited sample sizes and non-standardized data formats. A recent survey of publicly available MRI datasets for AI research13 identified 11 breast MRI collections, of which only the I-SPY1, I-SPY2, and Duke cohorts provide substantial patient numbers with NAC treatment data and pCR outcomes. Other breast MRI datasets are either substantially smaller (e.g., QIN-Breast: 67 patients, Breast-MRI-NACT-Pilot: 64 patients) or focus on diagnosis rather than treatment response prediction. To address these challenges, we present BreastDCEDL, a comprehensive deep learning-ready dataset for breast DCE-MRI. By integrating data from I-SPY214,15,16 (985 patients, 3 excluded for lacking three timepoints; n = 982), Duke17,18 (922 patients, 6 excluded for lacking three timepoints or bounding boxes; n = 916), and I-SPY119,20 (221 patients, of which 172 had at least three scans), we have created a standardized resource comprising 2,070 patients. The dataset transforms complex digital imaging and communications in medicine (DICOM) structures into accessible 3D arrays with preserved signal integrity, unified tumor annotations, and harmonized clinical metadata. Through BreastDCEDL, our objective is to accelerate progress in AI-assisted breast cancer care by providing the research community with the resources needed to develop, validate, and compare deep learning models in a standardized environment.
Dynamic Contrast-Enhanced MRI illustrates tumor segmentation across datasets. Representative DCE-MRI from the Duke, I-SPY1, and I-SPY2 datasets show pre-contrast, early post-contrast, and late post-contrast acquisitions. Columns display individual phases (left), a merged RGB composite (center), and tumor segmentation (right). Tumor localization in the Duke dataset is highlighted by a bounding box.
Methods
BreastDCEDL Dataset Composition
BreastDCEDL integrates three datasets from The Cancer Imaging Archive (TCIA)21: I-SPY2 (Investigation of Serial Studies to Predict Your Therapeutic Response with Imaging And moLecular Analysis 2)14,15,16, I-SPY1 (QuantumLeap Healthcare Collaborative)19,20, and Duke17,18. I-SPY2 contains 985 patients (3 excluded for lacking three timepoints), I-SPY1 has 172 patients with at least three scans, and Duke contains 916 patients with complete DCE data. This integration resulted in a cohort of 2,070 patients (Table 1).
Table 2 summarizes the institutional and scanner distribution across datasets.
DICOM to NIfTI Conversion Pipeline
The transformation from clinical DICOM to machine learning-ready NIfTI required addressing significant heterogeneity across 22+ imaging centers. The original data comprised 8.5 million DICOM slices (4.6TB) organized in hierarchical folders with mixed MRI acquisitions and vendor-specific tags, which we converted to 11,717 standardized NIfTI volumes (183GB) with clear naming convention: <id>_<ds>_acq<N>.nii.gz.
Temporal Reconstruction
DCE-MRI temporal information was encoded differently across vendors. We developed a hierarchical parsing strategy with three levels: (1) Primary identification using clinical trial timepoint tags (0012,0050) for T0 verification; (2) Contrast timing extraction from private tags (0117,1030-1035) including total phases (0117,1030), injection time (0117,1033), and SER timing indices (0117,1035); (3) Fallback parsing using SeriesDescription and ProtocolName when standard tags were absent.
For I-SPY2, tag (0117,1033) provided injection timing for precise alignment of pre-contrast (index 0), peak enhancement (index 2), and washout phases (indices 5–7). Duke data used “Pre”, “Post1”, “Post2” naming conventions in SeriesDescription fields.
Spatial Organization
DICOM slices arrived non-sequentially in 68% of series. Our pipeline: (1) extracted slice location from ImagePositionPatient (0020,0032); (2) computed slice normal vectors from ImageOrientationPatient (0020,0037); (3) sorted slices along anatomical axis using dot product projection; (4) validated against I-SPY analysis masks as spatial ground truth.
This automated pipeline eliminated 20-30 hours of manual preprocessing per researcher compared to raw TCIA data processing.
Tumor Annotation Processing
I-SPY Trials Segmentation
I-SPY trials contained complex processed data including ROI delineations, SER images, enhancement maps, and analysis masks encoding segmentation through byte operations. Segmentation steps were encoded as bit flags: manual VOI selection (1), thresholding (32), with combinations like code 33 (32+1) indicating voxels within VOI above threshold. We decoded these bitmaps into binary 3D masks (1 = malignant, 0 = background), yielding 172 masks for I-SPY1 and 982 for I-SPY2.
Duke Dataset Annotations
Duke sequences included non-fat saturated and fat-saturated scans at 1–2 minute intervals. Expert radiologists annotated bounding boxes for the largest tumor (among up to 9 per patient) based on pre-contrast DCE-MRI, first post-contrast, and their subtraction. We transformed these coordinates to align with standardized DCE-MRI orientation for 916 patients. Temporal resolution is approximately 60 seconds for Duke and 120 seconds for I-SPY trials; to harmonize early post-contrast timing across cohorts, we selected Duke’s second post-contrast acquisition (index 2) to approximate I-SPY early post-contrast timing.
Data Type Preservation and Standardization
Original DICOM used 16-bit integer or unsigned integer representation. We preserved full dynamic range by converting to 64-bit floating point format, maintaining signal integrity for quantitative analysis (Table 3).
Clinicopathologic Data Harmonization
We documented biomarker definition variations across datasets. HR status determination varied: I-SPY2 used greater than or equal to 1% staining, Duke used Allred score greater than 3, and I-SPY1 required greater than or equal to 10% staining. HER2 status assessment differed with Duke using FISH ratio greater than 2.2 versus I-SPY trials using FISH ratio greater than or equal to 2.0.
Original classifications were preserved in metadata files, enabling consistent threshold application. Tumor volumes were calculated by multiplying voxel counts by physical dimensions (I-SPY) or pre-categorized into size classes (Duke).
Benchmark Establishment
Fixed train-validation-test partitions ensure reproducible benchmarking across all 2,070 patients. Training set contains 1,532 patients total (1,099 with pCR data, 29.3% pCR rate). Validation set contains 268 patients total (177 with pCR data, 29.9% pCR rate). Test set contains 270 patients total (176 with pCR data, 30.1% pCR rate).
The 1,452 patients with available pCR data are balanced across splits, while the remaining 618 patients (primarily from Duke without NAC) are included for unsupervised learning and segmentation tasks.
Computational Requirements
Data standardization pipeline: 48 hours on workstation with 64 GB RAM.
Data Records
We developed BreastDCEDL, a comprehensive and standardized DCE-MRI dataset comprising 2,070 patients from three major clinical cohorts: Duke17,18 (n = 916), I-SPY119,20 (n = 172) and I-SPY214,15,16 (n = 982). The processed dataset is deposited on Zenodo22. The dataset includes:
-
I-SPY2: 7,040 DCE-MRI volumes (38.2 GB), 982 segmentation masks (59.1 MB)
-
I-SPY1: 585 DCE-MRI volumes (3.19 GB), 172 segmentation masks (32.1 MB)
-
Duke: 4,105 DCE-MRI volumes (142 GB), 916 bounding box annotations in CSV format
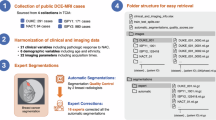
All data have been converted to standardized 3D NIfTI format with preserved DICOM intensity values, spatial resolution, and imaging parameters. Each patient includes 3–12 temporal 3D volumes with corresponding annotations and harmonized clinical metadata. The data organization structure and representative examples from each cohort are presented in Fig. 2.
BreastDCEDL structure and imaging examples. Standardized 3D NIfTI format with clinical annotations facilitates organized data integration. Representative DCE-MRI examples from contributing cohorts illustrate diverse imaging characteristics across institutions.
The dataset includes predefined train-validation-test partitions encompassing all 2,070 patients:
-
Training: 1,532 patients (1,099 with pCR data: 29.3% pCR rate)
-
Validation: 268 patients (177 with pCR data: 29.9% pCR rate)
-
Test: 270 patients (176 with pCR data: 30.1% pCR rate)
Among the subset of 1,452 patients with available pCR outcomes (primarily from I-SPY trials), the response rates are balanced across splits. The remaining 618 patients (mainly from Duke cohort without neoadjuvant treatment) are included in the splits for unsupervised learning, segmentation tasks, and radiomic feature development.
Table 4 summarizes the harmonized metadata fields. Spatial parameters (voxel dimensions, image size) are included to facilitate spatial harmonization. Acquisition-specific parameters such as scanner vendor, field strength, TR, and TE can be extracted from the original TCIA DICOM files using utilities provided in our GitHub repository.
Technical Validation
Data Transformation Validation
We successfully transformed 8,543,482 individual DICOM slices from TCIA into 11,717 analysis-ready 3D NIfTI volumes:
-
100% successful reconstruction of 3D volumes from 2D DICOM slices
-
Preservation verified for original 16-bit intensity values in 64-bit NIfTI format
-
Spatial ordering validated through anatomical landmark checking
-
Temporal sequence alignment confirmed across heterogeneous acquisition protocols
Data Completeness
Table 5 demonstrates the comprehensive nature of the BreastDCEDL dataset across all three cohorts. All 2,070 patients have complete DCE-MRI acquisitions successfully converted to NIfTI format. Tumor annotations are available for every patient, with I-SPY cohorts providing 1,154 volumetric segmentation masks and Duke providing 916 bounding box annotations. Clinical biomarker data show excellent completeness, with HR status available for 99.9% of patients (2,068/2,070) and HER2 status for 99.8% (2,065/2,070). The primary limitation concerns pathologic complete response data, which is available for all I-SPY patients (n = 1,154) but only 32.5% of Duke patients (298/916) due to differences in treatment protocols, resulting in 70.1% overall pCR data availability (1,452/2,070). This distribution reflects the Duke cohort’s inclusion of both neoadjuvant and primary surgery cases, while I-SPY trials exclusively enrolled neoadjuvant chemotherapy patients.
Annotation Validation
Tumor segmentation masks were validated for spatial alignment with DCE-MRI volumes. All 1,154 binary masks correctly overlapped with tumor regions. Tumor volumes ranged from 0.5-187.3 cm3 (median 8.7 cm3), consistent with clinical expectations for breast lesions.
Data Access and Implementation
The dataset’s standardized NIfTI format ensures compatibility with established neuroimaging workflows. Each patient’s temporal DCE series follows a consistent naming convention (<id>_<ds>_acq<N>.nii.gz) with corresponding tumor segmentations (<id>_mask.nii.gz), enabling automated batch processing without manual file matching. Standard libraries including nibabel, SimpleITK, and ITK-SNAP can directly load these volumes without format conversion.
Comprehensive implementation resources are maintained at https://github.com/naomifridman/BreastDCEDL, providing Python utilities for data loading and metadata parsing, temporal alignment verification scripts, Jupyter notebooks demonstrating analysis workflows and data visualization.
Dataset Characteristics and Limitations
BreastDCEDL preserves the inherent heterogeneity of multi-institutional clinical data, which enhances generalizability but requires careful consideration during analysis. The dataset encompasses significant variation in imaging protocols, with temporal resolution ranging from 60 seconds (Duke) to 120 seconds (I-SPY trials), potentially affecting kinetic curve analysis. Biomarker definitions also vary due to institutional standards: HR positivity thresholds range from ≥1% (I-SPY2) to ≥10% (I-SPY1), while HER2 positivity criteria differ between FISH ratio >2.2 (Duke) and ≥2.0 (I-SPY trials). No intensity harmonization or spatial normalization was applied to the converted data; original acquisition characteristics are preserved to allow researchers flexibility in applying harmonization methods appropriate to their specific analyses.
Annotation scope represents an important consideration for segmentation tasks. Only primary tumors are annotated despite multifocal disease being present in numerous cases, with up to nine distinct lesions observed in some Duke patients. This limitation means segmentation models trained exclusively on BreastDCEDL may require additional training data to detect secondary lesions reliably. Furthermore, Duke annotations consist of bounding box coordinates rather than pixel-level masks, necessitating adapted training strategies or conversion procedures for certain applications.
The availability of clinical endpoints varies across the dataset. While all I-SPY trial participants have pCR outcomes due to trial design, only 32.5% of Duke patients have this information, as many underwent primary surgery without neoadjuvant therapy. This distribution provides 1,452 patients with pCR labels suitable for supervised learning, while the remaining 618 patients remain valuable for unsupervised learning, segmentation tasks, and radiomic feature development.
Recommended Research Applications
BreastDCEDL’s standardized format and comprehensive annotations support diverse research directions in computational breast imaging. The predefined train/validation/test splits with balanced pCR rates (29.3%, 29.9%, and 30.1% respectively) ensure reproducible benchmarking across studies. The dataset is particularly well-suited for developing and validating treatment response prediction models, advancing tumor segmentation algorithms across different annotation types, extracting and validating radiomic biomarkers, studying multi-institutional model generalization, and characterizing temporal enhancement patterns across molecular subtypes.
The inclusion of patients without pCR outcomes enables semi-supervised learning approaches that leverage the full dataset, while the preservation of original intensity values supports quantitative perfusion analysis and pharmacokinetic modeling. Researchers can apply consistent biomarker thresholds across cohorts using the preserved original classifications, enabling both harmonized analyses and studies of inter-institutional variability.
Ethics statement
This study used publicly available deidentified datasets from TCIA21, including I-SPY119,20, I-SPY214,15,16, and Duke Breast Cancer MRI17,18 datasets. All original data collection for these datasets was conducted with appropriate institutional review board (IRB) approvals at the respective participating institutions, with informed consent obtained from all participants as described in the original publications. No additional ethical approval was required for this secondary analysis of publicly available anonymized data. All data handling and analysis procedures were in accordance with the relevant data protection regulations and institutional guidelines of Ariel University.
Data availability
The BreastDCEDL dataset is available on Zenodo22. The repository contains standardized NIfTI volumes for 2,070 patients, including:
• I-SPY1: Complete dataset with all DCE-MRI acquisitions and segmentation masks for 172 patients
• I-SPY2: Subset containing 3 timepoints (pre-contrast, early post-contrast, late post-contrast) per patient, cropped to 256 × 256 × Z containing tumor regions
•Duke: Subset with 3 DCE-MRI timepoints per patient, cropped to 256 × 256 × Z tumor-centered volumes for 916 patients. Harmonized clinical metadata and predefined train-validation-test splits are included
Harmonized clinical metadata and predefined train-validation-test splits are included. The complete I-SPY2 pretreatment NIfTI conversion with full-resolution volumes for all 982 patients is also available on TCIA23.
Code availability
The code for data processing is available on GitHub (https://github.com/naomifridman/BreastDCEDL) and archived on Zenodo24. The repository includes data loading utilities, usage examples, Jupyter notebooks demonstrating data access and visualization, and documentation for reproducing the DICOM-to-NIfTI conversion pipeline.
References
Romeo, V. et al. AI-enhanced radiomics for breast cancer diagnosis and prognosis: A systematic review and meta-analysis. European Journal of Radiology 157, 110561, https://doi.org/10.1016/j.ejrad.2022.110561 (2022).
Turnbull, L. W. Dynamic contrast-enhanced MRI in the diagnosis and management of breast cancer. NMR in Biomedicine 22, 28–39, https://doi.org/10.1002/nbm.1273 (2009).
Xiao, J. et al. Dynamic contrast-enhanced breast MRI features correlate with angiogenesis and tumor biology. NPJ Breast Cancer 7, 56, https://doi.org/10.1038/s41523-021-00247-3 (2021).
Li, L. et al. Parameters of dynamic contrast-enhanced MRI as imaging biomarkers for tumor angiogenesis and proliferation in breast cancer. European Journal of Radiology 84, 1224–1232, https://doi.org/10.1016/j.ejrad.2015.02.024 (2015).
Gullo, C., Eskreis-Winkler, S., Morris, E. & Pinker, K. Machine learning and breast MRI: Potential future applications. AJR https://doi.org/10.2214/AJR.18.20469 (2019).
Houssein, M., Emam, M. & Ali, M. & Suganthan, P. A comprehensive review of breast cancer diagnosis using machine learning. Expert Systems with Applications https://doi.org/10.1016/j.eswa.2021.115251 (2021).
Wei, J., Ma, Y. & Liu, F. Deep learning in breast cancer imaging: A systematic review. Engineering in Medicine 4, 100024, https://doi.org/10.1016/j.engmed.2024.100024 (2024).
LoGullo, R., Marcus, J. & Huayanay, L. Challenges in AI-based breast MRI analysis. Radiology: Artificial Intelligence https://doi.org/10.1148/ryai.20242345 (2024).
Liu, Y. et al. Multiphase fusion for breast DCE-MRI using CNNs. Scientific Reports https://doi.org/10.1038/s41598-020-67890-1 (2020).
Dammu, Y., Ren, H. & Duong, M. Multimodal fusion for breast cancer prediction. IEEE Transactions on Medical Imaging https://doi.org/10.1109/TMI.2023.3245678 (2023).
Joo, S. et al. Multimodal deep learning for breast cancer MRI. Computers in Biology and Medicine https://doi.org/10.1016/j.compbiomed.2021.104567 (2021).
Braman, N. et al. Deep learning-based prediction of response to HER2-targeted neoadjuvant chemotherapy from pre-treatment dynamic breast MRI: A multi-institutional validation study. arXiv https://doi.org/10.48550/arxiv.2001.08570 (2020).
Dishner, K. A. et al. A Survey of Publicly Available MRI Datasets for Potential Use in Artificial Intelligence Research. Journal of Magnetic Resonance Imaging 59, 450–480, https://doi.org/10.1002/jmri.29101 (2024).
Li, W. et al. I-SPY2 Breast Dynamic Contrast Enhanced MRI Trial (ISPY2) (Version 1) [Data set]. The Cancer Imaging Archive https://doi.org/10.7937/TCIA.D8Z0-9T85 (2022).
Newitt, D. C. et al. ACRIN 6698/I-SPY2 Breast DWI [Data set]. The Cancer Imaging Archive https://doi.org/10.7937/TCIA.KK02-6D95 (2021).
Li, W. et al. Predicting breast cancer response to neoadjuvant treatment using multi-feature MRI: results from the I-SPY2 TRIAL. npj Breast Cancer 6, 18, https://doi.org/10.1038/s41523-020-00203-7 (2020).
Saha, A. et al. Dynamic contrast-enhanced magnetic resonance images of breast cancer patients with tumor locations [Data set]. The Cancer Imaging Archive https://doi.org/10.7937/TCIA.e3sv-re93 (2021).
Saha, A. et al. A machine learning approach to radiogenomics of breast cancer: a study of 922 subjects and 529 DCE-MRI features. British Journal of Cancer 119, 508–516, https://doi.org/10.1038/s41416-018-0185-8 (2018).
Newitt, D. C. & Hylton, N. M. on behalf of the I-SPY1 Network and ACRIN 6657 Trial Team. Multi-center breast DCE-MRI data and segmentations from patients in the I-SPY1/ACRIN 6657 trials [Data set]. The Cancer Imaging Archive https://doi.org/10.7937/K9/TCIA.2016.HdHpgJLK (2016).
Hylton, N. M. et al. Neoadjuvant Chemotherapy for Breast Cancer: Functional Tumor Volume by MR Imaging Predicts Recurrence-free Survival—Results from the ACRIN 6657/CALGB 150007 I-SPY1 TRIAL. Radiology 279, 44–55, https://doi.org/10.1148/radiol.2015150013 (2016).
Clark, K. et al. The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository. Journal of Digital Imaging, https://doi.org/10.1007/s10278-013-9622-7 (2013).
Fridman, N., Solway, B., Fridman, T. & Barnea, I. BreastDCEDL: A Deep Learning–Ready Breast DCE-MRI Dataset. Zenodo https://doi.org/10.5281/zenodo.17274053 (2025).
Fridman, N. & Goldstein, A. Curated, Segmented, and Deep Learning-Optimized I-SPY 2 MRI Dataset for Prediction of pCR, HR, and HER2 Status (BreastDCEDL-ISPY2) (Version 1) [Data set]. The Cancer Imaging Archive https://doi.org/10.7937/42WQ-TH78 (2025).
Fridman, N., Solway, B., Fridman, T. & Barnea, I. naomifridman/BreastDCEDL: BreastDCEDL v1.0 (v1.0). Zenodo https://doi.org/10.5281/zenodo.15635800 (2025).
Acknowledgements
This research was supported by the Ministry of Innovation, Science & Technology, Israel, under the National Track Research Program (Grant No. 0005463). We gratefully acknowledge The Cancer Imaging Archive (TCIA) for providing access to the I-SPY1, I-SPY2, and Duke Breast Cancer MRI datasets that formed the foundation of this work. The authors thank the investigators and institutions that contributed to the original data collection and made these valuable datasets publicly available for research purposes.
Author information
Authors and Affiliations
Contributions
N.F. led the project conception, methodology design, data standardization, and manuscript preparation. B.S., T.F., and I.B. collaboratively developed the data filtering protocols, processing pipelines, GitHub repository infrastructure, and validation procedures. A.G. supervised the project. All authors participated in manuscript revision and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fridman, N., Solway, B., Fridman, T. et al. BreastDCEDL: A standardized deep learning-ready breast DCE-MRI dataset of 2,070 patients. Sci Data 13, 264 (2026). https://doi.org/10.1038/s41597-026-06589-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06589-6