Abstract
Bronchoscopy examination is essential for diagnosing and managing respiratory diseases. While Multimodality Large Language Models (MLLMs) can enhance the efficiency and accuracy of medical report writing, existing datasets lack descriptive and comprehensive annotations for complex cases, hindering their ability to facilitate adequate learning of image-report relationships. To address this problem, we introduce BERD, a Bronchoscopy Examination Report Dataset, which includes 3,692 bronchoscopy examination reports. Among these reports, 6,330 representative images are annotated with single-image text descriptions and classification labels. BERD emphasizes the provision of versatile and detailed descriptions of findings. All these reports and annotations were performed by experienced clinicians specializing in bronchoscopy. Furthermore, experimental results show that fine-tuning state-of-the-art MLLMs on BERD significantly improves their ability to generate accurate and comprehensive reports, advancing AI applications in bronchoscopy.
Similar content being viewed by others
Background & Summary
Bronchoscopy examination is a vital diagnostic and therapeutic tool in respiratory medicine1. It allows direct visualization of the tracheobronchial tree, enabling clinicians to identify abnormalities such as inflammations, infections, tumors, or structural changes2. In addition to its diagnostic utility, bronchoscopy examination is widely used for therapeutic interventions, such as foreign body removal, airway stenting, or lavage for microbiological analysis3. The findings from bronchoscopy are typically documented in detailed reports that provide crucial information for diagnosis, treatment planning, and follow-up care.
However, generating these reports is a labor-intensive task that relies heavily on the experience and expertise of clinicians4. Each report must not only accurately document the observed findings but also provide cohesive and structured descriptions to ensure effective communication among clinicians. The increasing demand for bronchoscopy examination writing in clinical workflows has amplified the need for efficient and accurate report generation methods, highlighting the potential role of artificial intelligence in automating and enhancing this process.
With recent advancements in artificial intelligence, Multimodality Large Language Models (MLLMs) have shown great promise in medical applications, especially in tasks requiring the integration of visual and textual data5. These models6,7,8,9,10, trained on paired image-text datasets, can analyze medical images and generate descriptive reports, offering a solution to the time and expertise constraints faced in clinical settings. For bronchoscopy examination reports, MLLMs can potentially automate the generation of structured, accurate, and comprehensive reports, reduce the workload of clinicians and improve reporting quality.
Despite these advancements, the training of MLLMs for generating bronchoscopy examination reports is hindered by the limitations of existing datasets. Most publicly available datasets for bronchoscopy focus on narrow tasks, providing only limited support for report generation, as shown in Table 1. For instance, the BroncoLC11 dataset is designed exclusively for tumor localization, offering annotations about tumor presence and its corresponding bronchial location, but neglects other common findings such as sputum, clot, or bleeding that are critical in routine bronchoscopic reports. Similarly, the UAAL12 dataset, primarily developed for bronchoscopy navigation, focuses solely on the position of the bronchoscopic device relative to the airway, without capturing any pathological or descriptive information. The PKDN13 dataset, while notable for its annotated bronchoscopic images, is a proprietary resource and focuses only on binary classification tasks (lesion vs. non-lesion), offering no insights into nuanced findings necessary for comprehensive report generation. The BI2K14 dataset, though broader in scope, divides the data into benign lesions, malignant lesions, and normal conditions, which still falls short of the granularity required to describe routine findings such as sputum, bleeding, edema, or congestion.
The limitations of these datasets highlight their inadequacy in facilitating MLLMs for detailed and comprehensive bronchoscopy examination report generation. Unlike radiology images, such as CT or MRI, where datasets like MIMIC15 and PMC16 provide paired image-text report data for training models capable of generating structured reports, bronchoscopy examination reports remain a shortage. Existing datasets have constrained the field to tasks such as navigation or single-lesion segmentation, leaving the task of comprehensive report generation largely unaddressed. This gap has hindered the ability of AI systems to provide meaningful assistance to clinicians, especially in automating the time-consuming process of detailed report writing.
To address these challenges, our BERD dataset provides a high-quality resource for training and evaluating MLLMs. By including 3,692 bronchoscopy examination reports, with 6,330 images annotated with detailed descriptions, BERD enables MLLMs to learn holistic and nuanced representations of bronchoscopic findings. Unlike existing datasets, BERD emphasizes report-centric annotations, capturing a wide range of findings, including common yet clinically significant observations. This dataset bridges the gap between current MLLM capabilities and the demands of clinical bronchoscopy report generation, paving the way for more accurate, efficient, and clinically relevant AI applications.
Methods
To facilitate the development of AI-powered automatic report generation in the Bronchoscopy field, we collected an image-caption pair dataset with high-quality complete annotations done by two professional clinicians. In the process of data collection, we removed all parts that might contain personal information about patients and clinicians, retaining only bronchoscopy images and objective descriptive reports without any private information. This retrospective study was approved by the Clinical Research and Laboratory Animal Ethics Committee of the First Affiliated Hospital of Sun Yat-sen University (Approval Number: Ethical Review No. [2024]517), permitting data collection, annotation, subsequent research, and publication. Since this study does not involve specimen collection, does not interfere with patient examination procedures, and does not include follow-up or biological samples, an application for exemption from informed consent was submitted and approved within the hospital.
Bronchoscopy examination reports
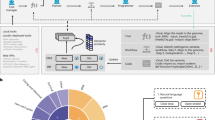
The dataset was collected by the Department of Pulmonary and Critical Care Medicine, First Affiliated Hospital, Sun Yat-sen University, Guangzhou, China. Between 2022 and 2023, a total of 8,477 bronchoscopy examinations were performed by experienced clinicians in the hospital, from which we selected 3,692 representative patient cases with 6,330 images. Each original report was generated with four images that were selected by the clinicians, as shown in Fig. 1. Clinicians can take screenshots at critical moments during bronchoscopy examinations and mark their positions using a bronchoscope made by Olympus. The captured images are usually of representative significance. After the examination, the clinician writes an examination report and selects the four most representative images from all the captured images to include in the final report.
Examples of translated original bronchoscopy reports. A report typically contains four images selected by the clinicians who conducted the examination. The four images are annotated with location. (a) Examples of the abnormal cases, where the lesions are depicted with their positions in the report. (b) Example of normal cases, where one template is used when no lesion is found.
Image-caption pair
For each bronchoscopy report, the process of pairing images with captions was carefully conducted to ensure clinical relevance and accuracy. Clinicians first manually reviewed each report to identify the most relevant descriptions for each image in the bronchoscopy examination report. Then, the clinicians removed the location information from the sentence and retained only the descriptive text. In addition, to ensure the robustness of the description and the accuracy of subsequent model training, text containing specific numbers, such as “3 mm” has been removed from the description. These textual descriptions were extracted directly from the examination reports, capturing details such as abnormalities (e.g., tumors, edema, or exudates) and their associated observations, including color, size, and amount. For images with no visible abnormalities, a standardized caption, such as “The lumen is unobstructed, and the mucosa is free of congestion, edema, or erosion. No neoplasms, foreign bodies, or active bleeding are found.” was given, we simplified this sentence to “It is normal.” and assigned to maintain consistency across the dataset. The caption corresponding to the image might contain lesions of more than one type. This meticulous pairing process ensures that every image is tightly linked to a meaningful and comprehensive description, providing a strong foundation for AI models to learn image-text relationships effectively. The original report and position marks were written in Chinese, after the image-caption pair annotation, we translated them into English using a locally deployed LLM (Large Language Model) Qwen3-32B17 to protect data privacy.
LLM-assisted classification annotation
To streamline the classification of images, we integrated a locally deployed LLM into the annotation workflow. Experienced clinicians first defined a comprehensive list of disease categories from expert consensus and bronchoscopy reporting guidelines, including common terms like congestion, edema, or tumor. Using this reference, LLM was employed to extract relevant keywords and synonyms from the captions, automatically categorizing each image-caption pair into one or more predefined classes. After the initial classification, all LLM-generated labels were reviewed and refined by clinicians to ensure clinical accuracy and alignment with medical standards. This semi-automated approach significantly reduced the manual workload while preserving the overall quality and consistency of the dataset.
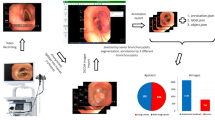
The whole annotation process is illustrated in Fig. 2(a), and the final annotation result is shown in Fig. 2(b).
(a) The process of annotation: we first extract the detailed descriptions from the original report and then distribute corresponding descriptions to each image while removing the unrelated images, and then annotate the labels based on the sentences. (b) Examples of the annotated dataset. The dataset comprises images along with their corresponding locations, descriptions, and labels.
Data Records
The dataset is available from the Science Data Bank at https://doi.org/10.57760/sciencedb.2801818.
The dataset contains two folders, one of which is the annotation folder that contains annotation JSON files. The other folder contains images in PNG format.
The annotation JSON files contain one training annotation file and a testing annotation file. The annotation file includes seven elements:
image_path, image_id, caption, location, width, height, label, patient_id. The image_path is the relative path of the image, image_id is the unique ID of the image, which should be the same as its image name. The caption is the caption annotation. The location is the anatomical location of the image. The height and width are the size of the image. And the label is the classification result of the image. The patient_id is the ID of patients. Different images might come from the same patient. Both training and testing images are in the images folder. The dataset folder structure is shown in Fig. 3.
The structure of the dataset folder.
Technical Validation
Experience of the operators
The department where the bronchoscopy examination reports are collected specializes in minimally invasive diagnosis and treatment of respiratory diseases, with an average annual completion of over 5,000 bronchoscopic procedures, including diagnostic bronchoscopy and complex interventions such as tumor resection, airway stenting, and bronchial fistula occlusion. Therefore, the quality and diversity of the bronchoscopy examination report can be guaranteed.
Experience of the annotators
The annotation for this study was carried out by two bronchoscopists, each with 5 more years of specialized experience in bronchoscopy, supported by two standardized-trained resident clinicians. All annotations were supervised and verified by a senior expert with more than 10 years of bronchoscopic practice, who has performed over 10,000 bronchoscopic examinations and leads technical innovations in navigation-guided biopsies. Referring to clinical atlas standards, bounding boxes and labels for anatomical landmarks and airway lesions were independently annotated by the two experienced bronchoscopists, followed by final review by the senior expert to ensure annotation accuracy.
Analysis of the dataset and annotations
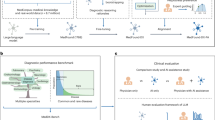
To validate the effectiveness of our dataset across various tasks and demonstrate its superiority over current state-of-the-art (SOTA) closed-source models, we conducted comprehensive experiments. These experiments primarily focused on generating bronchoscopic reports and evaluating the performance of leading closed-source General MLLMs, Medical MLLMs, and those fine-tuned on our dataset. Because the bronchoscopic images contain bloody elements, they cannot pass the image review mechanisms of most closed-source MLLMs. Therefore, closed-source MLLMs were not considered in this evaluation. The goal is to verify the utility of our dataset in this domain and to show that current SOTA models, having no prior exposure to bronchoscopic data, perform poorly on such tasks. The procession of caption generation is shown in Fig. 4. The image passes through a vision encoder and an MLP alignment layer, while the textual input passes through the text input module. The visual input is aligned with textual space. Two inputs are then sent to the LLM to generate the final textual output.
The process of caption generation and report writing. We take the image and default prompt as input to the MLLM, and the output is the corresponding caption of that image. In real practice, the MLLM-generated captions can be revised and utilized by the clinicians while writing the bronchoscopy examination report.
Evaluation metrics
We employed a combination of standard natural language processing (NLP) metrics and expert evaluations to ensure a robust assessment of the generated reports. BLEU: Measures the precision of n-grams in the generated text compared to the reference text. It evaluates how closely the generated reports match the ground truth at the word and phrase levels. BLEU@1 to BLEU@4 represent BLEU scores calculated using 1-gram to 4-gram precision, respectively, where higher n-gram values provide more stringent evaluation of text fluency and coherence. ROUGE-L: Focuses on the recall of sequences between the generated report and the reference, emphasizing the overlap of the longest matching subsequences. METEOR: Considers both precision and recall by aligning words and phrases semantically, using synonyms and stemming to capture meaning. CIDEr: Evaluates the consensus between the generated text and the reference text based on term frequency-inverse document frequency (TF-IDF), ensuring relevance and informativeness in the generated reports. Accuracy: To achieve a more intuitive expression while aligning with the cognition of clinicians when composing bronchoscopy examination reports, we asked clinicians to rate the generated captions. The scoring results were binary, with “1” indicating acceptance, meaning the caption could be directly included as part of the report, and “0” indicating rejection, suggesting that the report contained unreasonable elements. Rejection could arise from various reasons, such as missing content or incorrect descriptions. In such cases, clinicians deemed the results unacceptable and required modifications to be made before they were included in the report.
Experimental results
We randomly extracted 6,014 images for training and 316 images for testing with no overlap in patients between two datasets. First, we evaluated the performance of current general and medical MLLMs on the test set. To align the model outputs more closely with the style of our caption dataset, we utilized prompts and few-shot examples as shown in Fig. 5. The outputs of both general models and medical domain MLLMs were inferior, failing to accurately describe the bronchoscopy images. This highlights that these MLLMs have not undergone pre-training or fine-tuning in the bronchoscopy domain, likely due to the lack of publicly available datasets in this field. To address this, we fine-tuned general models, specifically Qwen2.5VL19 (2B and 7B) and InternVL-320 (3B and 8B) and tested their performance. The results demonstrate that our fine-tuned models achieved significant improvements across all metrics. The best-performing model, InternVL3-8B, achieved BLEU@1 to BLEU@4 scores of 35.06%, 30.50%, 27.70%, and 25.83%, respectively. ROUGE-L reached 36.29%, METEOR reached 38.42%, and CIDEr scored 27.71%. Additionally, clinicians assess the binary classification accuracy of the generated captions. InternVL-3-8B achieved the highest score, with an accuracy of 82.91%, outperforming the second-best model, Qwen2.5VL-7B, by 1.58% as shown in Table 2, and one result example is shown in Fig. 6.
Prompt used in the MLLM report generation. We use this prompt to make generated reports closer to our caption style.
Caption results by different MLLM models. General MLLMs and Medical MLLMs can not recognize the lesion in the image in the test dataset. After fine-tuning on the training dataset, the lesion can be recognized with a concise description as the clinician desired.
Usage Notes
To facilitate the use of the dataset, we offer public access to both the database and all related code. We have provided evaluation metrics for each task and divided the dataset into training and testing sets to ensure a fair comparison. Therefore, we believe this dataset should serve as an excellent benchmark for these relevant tasks and pave the way for report generation tasks. As the details for the data digitization process and codes for pre-processing are provided.
Limitations
Despite the high potential in developing report generation models, the proposed dataset has certain limitations. First, all data was collected from a single hospital using the Olympus bronchoscope. This may limit the generalizability of the dataset to other institutions or equipment types. However, since the Olympus bronchoscope is widely used in clinical practice, the dataset maintains a certain level of standardization and remains broadly applicable to similar clinical settings employing the same bronchoscopic technology. Second, specific numerical values, such as lesion sizes (e.g., “3 mm”), were removed from the reports. This decision was made because such quantitative details cannot be directly observed from the corresponding images and could potentially introduce hallucinations in multimodal large language models during report generation. Third, the reports were written by different clinicians, leading to subtle variations in descriptive style and interpretation.
Data availability
The dataset is available from the Science Data Bank at https://doi.org/10.57760/sciencedb.2801818. The name of the repository is “BERD: Fine-Grained Bronchoscopy Examination Report Dataset”.
Code availability
The code for data processing, dataset statistical analysis, and MLLM validation can be found at https://github.com/lxj22/BERD.
References
Fulkerson, W. Current Concepts - Fiberoptic Bronchoscopy. N. Engl. J. Med. 311, 511–515, https://doi.org/10.1056/nejm198408233110806 (1984).
Ernst, A., Silvestri, G., Johnstone, D. & Diagnost, A. I. C. Interventional pulmonary procedures: Guidelines from the American College of. Chest Physicians. Chest 123, 1693–1717, https://doi.org/10.1378/chest.123.5.1693 (2003).
Criner, G. et al. Interventional Bronchoscopy. American Journal of Respiratory and Critical Care Medicine 202, 29–50, https://doi.org/10.1164/rccm.201907-1292SO (2020).
Zhang, J. et al. AI co-pilot bronchoscope robot. Nature Communications 15, https://doi.org/10.1038/s41467-023-44385-7 (2024).
Xiao, H. et al. A comprehensive survey of large language models and multimodal large models in medicine. Information Fusion 117, https://doi.org/10.1016/j.inffus.2024.102888 (2025).
Bannur, S. et al. MAIRA-2: Grounded Radiology Report Generation. Preprint at https://arxiv.org/abs/2406.04449 (2024).
Chen, J. et al. HuatuoGPT-Vision, Towards Injecting Medical Visual Knowledge into Multimodal LLMs at Scale. Preprint at https://arxiv.org/abs/2406.19280 (2024).
Li, C. et al. LLaVA-med: training a large language-and-vision assistant for biomedicine in one day. Advances in Neural Information Processing Systems 36, 28541–28564, https://dl.acm.org/doi/10.5555/3666122.3667362 (2023).
Wu, C. et al. Towards generalist foundation model for radiology by leveraging web-scale 2D&3D medical data. Nature Communications 16, https://doi.org/10.1038/s41467-025-62385-7 (2025).
Wu, L. et al. UniBiomed: A Universal Foundation Model for Grounded Biomedical Image Interpretation. Preprint at https://arxiv.org/abs/2504.21336 (2025).
Vu, V. et al. BM-BronchoLC - A rich bronchoscopy dataset for anatomical landmarks and lung cancer lesion recognition. Scientific Data 11, https://doi.org/10.1038/s41597-024-03145-y (2024).
Hao, R. et al. Upper Airway Anatomical Landmark Dataset for Automated Bronchoscopy and Intubation. Scientific Data 12, 1907, https://doi.org/10.1038/s41597-025-06169-0 (2025).
Yan, P. et al. PKDN: Prior Knowledge Distillation Network for bronchoscopy diagnosis. Computers in Biology and Medicine 166, https://doi.org/10.1016/j.compbiomed.2023.107486 (2023).
Sun, W. et al. An accurate prediction for respiratory diseases using deep learning on bronchoscopy diagnosis images. Journal of Advanced Research 76, 423–438, https://doi.org/10.1016/j.jare.2024.11.023 (2025).
Johnson, A. et al. MIMIC-III, a freely accessible critical care database. Scientific Data 3, https://doi.org/10.1038/sdata.2016.35 (2016).
Roberts, R. PubMed Central: The GenBank of the published literature. Proceedings of the National Academy of Sciences of the United States of America 98, 381–382, https://doi.org/10.1073/pnas.98.2.381 (2001).
Yang, A. et al. Qwen3 Technical Report. Preprint at https://arxiv.org/abs/2505.09388 (2025).
LUO, X. et al. BERD: Fine-Grained Bronchoscopy Examination Report Dataset. Science Data Bank https://doi.org/10.57760/sciencedb.28018 (2025).
Bai, S. et al. Qwen2.5-VL Technical Report. Preprint at https://arxiv.org/abs/2502.13923 (2025).
Zhu, J. et al. InternVL3: Exploring Advanced Training and Test-Time Recipes for Open-Source Multimodal Models. Preprint at https://arxiv.org/abs/2504.10479 (2025).
Xu, W. et al. Lingshu: A Generalist Foundation Model for Unified Multimodal Medical Understanding and Reasoning. Preprint at https://arxiv.org/abs/2506.07044 (2025).
Acknowledgements
This research was supported by the InnoHK initiative of the Government of the Hong Kong Special Administrative Region.
Author information
Authors and Affiliations
Contributions
Xingjian Luo: Methodology, Experiment, Writing (Draft, Graph and Visualization), Review & Revising. Xinyan Huang: Conceptualization, Supervision, Data Collection, Annotation, Review & Revising. Xusheng Liang: Experiment, Review & Revising. Jiyu Wang: Data Collection, Annotation, Review. Jincui Gu: Data Collection, Annotation, Review. Dong Yi: Conceptualization, Project Administration, Review & Revising. Haohan Zhao: Review & Revising. Haihong Zhang: Conceptualization, Project Administration. Jinlin Wu: Review & Revising. Zhen Lei: Supervision, Review & Revising. Gaofeng Meng: Supervision, Review & Revising. Hongliang Ren: Supervision, Review & Revising. Jiebo Luo: Supervision, Review & Revising. Huai Liao: Conceptualization, Review & Revising, Data Collection, Project Administration. Hongbin Liu: Conceptualization, Supervision, Review & Revising, Project Administration.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, X., Huang, X., Liang, X. et al. Towards Automated Reporting: A Bronchoscopy Report Dataset for Enhancing Multimodality Large Language Models. Sci Data 13, 339 (2026). https://doi.org/10.1038/s41597-026-06692-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06692-8