Abstract
Sleep architecture and integrity significantly influence neural recovery and cognitive restoration. These are particularly relevant in ischemic stroke survivors where sleep-disordered breathing (SDB) is a common comorbidity. To address the lack of stroke-specific sleep data, we present the Polysomnography Dataset for Sleep Analysis in Ischemic Stroke Patients (iSLEEPS), the first Asian and one of the largest stroke-specific sleep databases. Data collection was carried out between September-2018 and December-2021 at NIMHANS, India. iSLEEPS comprises 100 overnight PSG recordings with comprehensive expert annotations. Each recording includes sleep stages manually scored at 30-second epochs, detailed respiratory events, periodic limb movements, oxygen desaturation episodes, and clinical metrics, as per AASM (2017) guidelines. Our cohort demonstrates a high prevalence of SDB, enabling the investigation of stroke-sleep pathophysiology interactions. To illustrate dataset utility, we implemented automated sleep stage classification using deep learning methods. The Long Short-Term Memory model achieved the highest accuracy (74.70%), followed by Transformer (67.44%) and Convolutional Neural Network (61.65%). This dataset addresses crucial gap in stroke sleep research, supporting comprehensive analysis of post-stroke sleep disturbances.
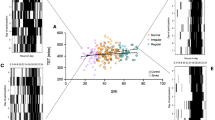
Similar content being viewed by others
Background & Summary
Sleep is a fundamental physiological process essential for cognitive performance, synaptic plasticity, and overall neurological health1. Sleep disorders, particularly insomnia and sleep-disordered breathing (SDB), affect a significant proportion of the global population. The prevalence of insomnia is estimated at approximately 37.5% worldwide2, while SDB affects between 9% and 38% of adults3. In India, community-based studies suggest insomnia prevalence is around 18-20%4,5. Recent systematic reviews estimate that over 100 million working-age Indians live with obstructive sleep apnea (OSA), with 37% meeting criteria for moderate-to-severe disease6. These figures underscore the clinical and public health burden of sleep disorders in both global and Indian contexts.
Stroke remains one of the leading causes of disability and mortality worldwide. Sleep apnea is an independent risk factor that doubles the likelihood of stroke recurrence. Between 70-80% of stroke patients exhibit some form of SDB, which significantly impacts recovery and outcomes7,8,9. Indian cohorts report similar findings, with approximately 75-82% of ischemic stroke patients demonstrating SDB on PSG10. Stroke survivors with moderate-to-severe SDB typically have higher modified Rankin Scale (mRS) and NIH Stroke Scale (NIHSS) scores, indicating greater disability and dependency. The most common type of SDB is OSA, affecting nearly two-thirds of stroke patients, while central sleep apnea (CSA) occurs in 12-17%, and Cheyne-Stokes respiration (CSR) is less common but associated with severe neurological deficits11.
The gold standard for diagnosing SDB is polysomnography (PSG), which monitors EEG, EOG, EMG, ECG, respiratory effort, oxygen saturation, and airflow overnight. However, PSG is resource-intensive and underemployed in acute stroke care, particularly in low-resource settings. Despite its diagnostic value, access to large PSG datasets of stroke patients remains limited, especially in non-Western populations12,13.
Several sleep-related datasets exist, but each has notable limitations that restrict their utility in stroke research. The Apnea-ECG database14, hosted on PhysioNet, consists of 70 overnight ECG recordings with apnea annotations. While this dataset has been widely used for developing automated apnea detection algorithms, its reliance on a single physiological channel (ECG) prevents comprehensive multimodal analysis of sleep stages, arousals, or neurological signals. The OSASUD dataset (Italy)15 includes 30 patients admitted to a stroke unit and provides ECG and photoplethysmography (PPG) signals for obstructive sleep apnea detection. Despite its relevance to stroke, the dataset is limited by its small sample size, lack of full PSG channels, and absence of manual scoring across sleep stages. The SVUH/UCD Sleep Apnea database16 contains PSG data from 25 patients, primarily drawn from a non-stroke population, making it unsuitable for investigating stroke-specific sleep pathophysiology. Similarly, the HuGCDN2014 collection from China17 provides recordings for 77 patients, but it does not include detailed event annotations such as respiratory arousals or hypnograms, which are essential for benchmarking automated models.
Large-scale clinical trials have also provided valuable epidemiological insights into sleep-disordered breathing after stroke, but their raw data remain inaccessible. For example, the SAS Care 1 study (Europe)18 prospectively followed 166 patients with acute stroke and transient ischemic attack (TIA), reporting a very high prevalence of SDB at baseline (85.6%) and persistent elevation at three months (82.7%). Although these findings underscore the clinical importance of SDB in stroke, the absence of open raw PSG data limits reproducibility and secondary analyses by the wider community.
Taken together, the existing databases either (i) focus on single-channel physiological recordings, (ii) involve small or non-stroke populations, (iii) lack comprehensive annotations, or (iv) are not openly available for reuse. As a result, no current open-access dataset offers large-scale, multimodal PSG with expert sleep staging and respiratory event annotations specifically in stroke patients, particularly from non-Western populations.
To address this gap, we curated the iSLEEPS dataset, comprising 100 overnight PSG recordings from ischemic stroke patients at the National Institute of Mental Health and Neurosciences (NIMHANS), Bengaluru, India. Each study includes multimodal PSG signals, manual annotations of sleep stages and respiratory events following AASM 2017 criteria, and associated demographic and clinical metadata. By offering a stroke-specific, non-Western cohort with high-quality, anonymized PSG data and rich annotations, iSLEEPS represents a unique open resource for advancing research on sleep and stroke. It provides opportunities for clinical investigation, benchmarking of machine learning algorithms, and comparative studies across populations, supporting the broader goals of open science in sleep medicine.
Methods
Study Subjects
The iSLEEPS cohort comprises 100 patients recruited between September 2018 and December 2021 at the Neurology Department of the National Institute of Mental Health and Neurosciences (NIMHANS)10, Bengaluru, India. All patients were diagnosed with ischemic stroke within one month of onset. Exclusion criteria were applied to ensure compliance and safety: individuals unable to tolerate polysomnography, those with severe aphasia precluding consent, or those at risk of withdrawal syndromes were not included. Common vascular comorbidities such as diabetes, atrial fibrillation, heart disease, and obesity were not considered exclusions, thereby reflecting the clinical diversity of real-world stroke populations. The resulting sample (mean age 50.5 ± 12.0 years) comprised 23 females and 77 males.
The iSLEEPS dataset includes 100 subjects with a mean age of 50.52 years (SD = 12.00), comprising 23 females (mean age 47.82 ± 12.00 years) and 77 males (mean age 47.83 ± 12.01 years).
Ethical consideration
The study was carried out at the National Institute of Mental Health and Neurosciences (NIMHANS), Bengaluru, India, a leading tertiary care referral teaching hospital. Approval was obtained from the NIMHANS Institutional Ethics Committee [No. NIMHANS/34th IEC (BS&NS DIV.)/2022 dated 05.02.2022]. Ethical considerations were strictly followed throughout the study. All participating patients provided written informed consent, ensuring they were fully aware of the study procedures and objectives. Consent included permission for anonymized data to be deposited in publicly available repositories for future research, in keeping with open science guidelines.
Overnight polysomnography
Overnight PSG was performed according to AASM 2017 guidelines using the SOMNOscreen plus system (SOMNOmedics GmbH, Germany). Recordings included six EEG, two EOG, and three EMG channels, with reference electrodes at bilateral mastoids. Additional signals comprised airflow, thoracoabdominal effort, SpO2, snore, ECG, and body position. Sampling frequencies were 128 Hz for EEG/EOG/respiratory channels, 256 Hz for EMG/ECG/snore, and 4 Hz for SpO2. The SOMNOmedics amplifier incorporates integrated hardware filters designed to optimize physiological signal acquisition while minimizing environmental and physiological noise. For EEG, EOG, and EMG channels, the bandpass hardware filters were set between 0.2–35 Hz, with a 50 Hz notch filter applied to suppress mains interference. ECG signals were acquired with a low-frequency cutoff of 0.3 Hz and a high-frequency cutoff of 70 Hz, also with a notch filter enabled. Respiratory channels, including thoracoabdominal belts, nasal pressure, and thermistor signals, were filtered with a bandpass range of 0.1–15 Hz. Pulse oximetry (SpO2) was recorded at 4 Hz, with integrated manufacturer filters reducing motion and light-related artifacts. Prior to data acquisition, electrode impedances for EEG and EOG were maintained below 10 kΩ (ideally within 5–10 kΩ) to ensure high-quality recordings. All data captured by the SOMNOmedics system were stored in EDF+ format, preserving raw signals for subsequent re-analysis.
Medical data annotation process and Data anonymization
After the completion of polysomnography, the recorded files were converted to European Data Format (EDF) using Dominos software. The manual sleep score was then performed using Polyman software by independent researchers who followed the 2017 American Academy of Sleep Medicine guidelines. This process was supervised by a neurologist with extensive experience in sleep medicine, ensuring that the scoring was accurate and consistent. The resulting hypnogram data, which provides a detailed record of the patient’s sleep stages and events, was exported in EDF format from the Polyman software for further analysis.
The detection of apnea/hypopnea events during the recorded sleeping hours was performed manually by simultaneous observation of all channels of the PSG system, according to the general criteria for apnea episode scoring. The final diagnosis concerning the categorization of the patient into one of the reported apnea severity cases: “Severe”, “Moderate”, “Mild Apnea”, and “Normal” was extracted through the Apnea/Hypopnea Index (AHI). The AHI is defined as the ratio of the total count of apneic episodes in the entire sleep study over the total sleep time (TST) in hours, which results in the mean count of apneic events per sleeping hour. Up to 5 apnea/hypopnea episodes per hour classify the subject into the class of “Normal breathing” during sleep, while higher values indicate a gradually increasing severity of SAS (“Mild Apnea”: 5 episodes/h ≤ AHI < 15 episodes/h; “Moderate Apnea”: 15 episodes/h ≤ AHI < 30 episodes/h; “Severe Apnea”: 30 episodes/h ≥ AHI).
To ensure patient confidentiality and data security, the polysomnographic data and scored EDF files were anonymized using custom scripts developed in Jupyter, a Python-based platform. These scripts were designed to remove any identifying information from the data, ensuring that patient privacy was maintained while allowing for comprehensive analysis. The anonymized data were then securely stored in the cloud, making them accessible for subsequent research and analysis while maintaining the highest standards of data protection and privacy.
Data preprocessing
We processed the raw recordings stored in EDF files by segmenting them into 30-second epochs at a sampling frequency of 100 Hz. We downsampled the EEG, EOG, and EMG signals from 128 Hz to 100 Hz to ensure consistency with commonly used AI-based sleep algorithms and publicly available sleep datasets. Each sleep epoch was then matched with the corresponding sleep stage annotations provided in .xlsx files. These annotated epochs were subsequently saved in. npz format for each of the 100 subjects. All preprocessing and baseline model code is available in the GitHub repository (Website: https://github.com/suvadeepmaiti/iSLEEPS). Automated sleep stage classification using deep learning methods was chosen as an example application to demonstrate the utility of the dataset. For training and testing deep learning models, EEG data from an identified channel was extracted and provided as input to the models. Data extraction typically used either a fixed epoch size of 30 seconds or a sliding-window approach. For the ResNet-based model, one epoch (30 seconds) was used at a time. In contrast, Transformer and LSTM-based models used windowed inputs of five consecutive epochs (150 seconds) to capture temporal dependencies.
Baseline models
This section outlines the popular baseline models employed for sleep stage classification tasks on our dataset using supervised learning techniques. Automated sleep stage classification has gained significant attention in sleep research due to its potential to enhance our understanding of sleep patterns and disorders. Primarily, EEG and EOG channels have been utilized for this purpose. State-of-the-art architectures based on CNN, LSTM, and Transformer models have been employed for automated sleep stage classification baselines19.
We benchmarked our dataset using ResNet-18, SE-ResNet-LSTM, and SE-ResNet Transformer-based model architectures on raw, single-channel EEG (C4-A1 or C4-M1) and EOG (O2-A1 or EOG1-M1) signals. For the ResNet-based models, the input was fixed epoch of 30 seconds duration presented at a time. In contrast, for the Transformer and LSTM-based models, we created windowed inputs by considering 5 consecutive epochs (5 × 30 = 150 seconds). This approach ensures that the models can learn from both the past and future epochs, capturing the temporal dependencies within the sequential time series signals.
-
CNN-Based Model20: It utilizes 1D ResNet-18 model architecture to extract features from each sleep epoch. It then employs a single linear layer to classify the epochs into five distinct sleep stages.
-
Transformer-Based Model21: The model architecture comprises a SE-ResNet-Transformer framework, which includes a Squeeze-and-Excitation (SE) block in each residual block to dynamically recalibrate feature maps. The SE-ResNet is followed by a Transformer encoder that leverages self-attention mechanisms to capture intricate temporal dependencies within the sequential time series signals.
-
LSTM-Based Model22: It employs a feature extractor and temporal encoder Bi-LSTM layers. The feature extractor captures intricate time series signal features, while stacked Bi-LSTM layers are used as a temporal context encoder to model the sequential dependencies in the data, resulting in improved sleep stage classification performance.
Experimentation Setup
We utilized official implementations of deep learning models wherever available and carefully replicated them to ensure consistency. A comprehensive evaluation was performed using several state-of-the-art deep learning approaches to benchmark our dataset. The data were split into training (80%), validation (10%), and testing (10%) sets on a patient-wise basis to prevent data leakage, ensuring that all sleep epochs from a given patient were assigned exclusively to one set. Additionally, we employed 10-fold cross-validation for each experimental setup to enhance the robustness of our results.
Training for models was conducted for a minimum of 50 epochs and a maximum of 100 epochs. The Adam optimizer was used with a batch size of 1024 for the ResNet Model and 128 for the other two models, with a fixed learning rate of 0.001 during the optimization process. All experiments were performed using Python 3.10.10, Pytorch 2.0.0+cu117 on RTX 3080 Ti 12GB GPU. The version details of all libraries are listed in the requirement.txt in the GitHub repository.
Data Records
The dataset is hosted on both Zenodo23 and Data Foundation, IHub-Data located at the International Institute of Information Technology, Hyderabad (IIITH), ensuring its availability to the global research community. It can be directly accessed via (Website: iSLEEPS) datasets under the Healthcare Dataset category (iSLEEPS) after user registration. This process facilitates easy download and utilization of the dataset by researchers and clinicians interested in sleep-stroke research. The data organization is as follows:
-
README.pdf: contains a detailed overview of the dataset structure, file formats, and naming conventions to guide users in effectively navigating and utilizing the iSLEEPS dataset.
-
iSLEEPS.zip: The folder contains 100 PSG recordings in EDF (European Data Format) and corresponding annotation files in .xlsx format. Each .edf file contains raw PSG data, while the associated .xlsx file provides detailed sleep and other relevant annotations for that recording.
-
subject_description.xlsx: A comprehensive spreadsheet containing patient demographic data and clinical data.
EDF and annotation files are labeled using the convention “SNxx,” where ‘xx’ represents an anonymized and incrementally assigned patient or case number (e.g., SN1, SN2, ..., SN100). This dataset includes 100 clinically acquired PSG recordings in European Data Format (EDF), collected at the National Institute of Mental Health and Neurosciences (NIMHANS). Of the 100 recordings, 28-channel configurations appear in 80 recordings. The remaining include 21 channels (6 recordings), 22 channels (5 recordings), 23 channels (5 recordings), and 20, 25, 26, or 27 channels (1 recording each). The most frequently used channel names are listed in Table 1 (See Table 1). To ensure participant confidentiality, all EDF headers have been replaced with standardized, de-identified versions in accordance with the dataset’s anonymization protocol. The total duration of recordings amounts to 795.95 hours, with individual studies ranging from 3.93 to 9.82 hours (mean: 7.96 hours). Approximately 90% of the files contain between 7 and 10 hours of data, although the actual sleep time within each recording may vary. Most signals were recorded at a sampling frequency of 128 Hz. Corresponding annotation files (.xlsx) include a total of 95,305 sleep stage annotations, with the number of annotated 30-second epochs per study ranging from 470 to 1177 (mean: 953.05 epochs).
Each recording is accompanied by an annotation file (.xlsx), containing a total of 95,305 annotations, including sleep stage labels assigned at 30-second intervals. These annotations use standardized labels such as “Sleep stage W” (wake), “Sleep stage N1”, “N2”, and “N3” (non-REM stages), “Sleep stage R” (REM), and “Sleep stage?” or “Artifact” to denote ambiguous or unscorable epochs-”typically occurring after “Lights On” or due to technical issues. Such segments usually indicate the end of a study and are not considered for analysis. Of all annotations, 79.48% are related to sleep staging, corresponding to the full 795.95 hours of data: 26.20% (24,970) are “Sleep stage W”, 9.94% (9,471) are “Sleep stage N1”, 41.27% (39,329) are “Sleep stage N2”, 8.74% (8,337) are “Sleep stage N3”, and 11.78% (11,225) are “Sleep stage R”. An additional 2.08% (1,982) are categorized as “Others” (see Fig. 1). In addition, each annotation file includes a Flow Events sheet that documents apnea-related episodes such as Obstructive Apnea, Central Apnea, Mixed Apnea, Hypopnea, Obstructive Hypopnea, and Respiratory Effort-Related Arousal (RERA), along with oxygen desaturation events and sleep-related arousals. Complementary demographic and clinical metadata—such as age, biological sex, medication use, and stroke-specific factors—are consolidated in a separate file, subject_description.xlsx, which provides structured information for all patients in the dataset.The comparison of annotations across datasets in the literature with the iSLEEPS dataset is presented in the Table 2 (See Table 2).
Sleep stage distribution for all 100 subjects. The chart shows the frequency of each sleep stage (Wake, N1, N2, N3, REM) for each subject. The sleep stages are color-coded as follows: Wake (blue), N1 (green), N2 (red), N3 (purple), and REM (orange). The height of each bar represents the total frequency of each sleep stage observed for each subject.
Technical Validation
The annotated apnea-related episodes
The diversity among participants regarding biological sex, age, and the final diagnosis of sleep apnea syndrome (SAS) severity can significantly influence the characteristics of recorded breathing and snoring sounds, as well as the properties of each episode, such as its duration. PSG is typically recommended for patients who report excessive daytime sleepiness or loud snoring at night, symptoms strongly associated with SAS. As expected, a substantial portion of the participants fall into the “Severe Apnea” (38%) and “Moderate Apnea” (23%) categories, whereas “normal” cases make up only 15% of all diagnosed individuals (Fig. 2a). Figure 3 Given the higher risk of SAS in males, there is a pronounced biological sex imbalance in the dataset, with men comprising 76% of the subjects (Fig. 4a).
(a) Distribution of patients among AHI severity classes. The pie chart shows the percentage of patients categorized as Normal (15.0%), Mild Apnea (24.0%), Moderate Apnea (23.0%), and Severe Apnea (38.0%). (b) Distribution of AHI values by apnea severity. The bar chart represents the number of patients with different AHI values, categorized by apnea severity: Normal, Mild Apnea, Moderate Apnea, and Severe Apnea.
Comparison of Actual and Predicted Hypnograms. (a) displays the actual hypnogram for subject-1 of SN1, as annotated by a sleep expert. (b) shows the hypnogram predicted by the LSTM-based model using raw EEG signal input.
(a) Biological sex distribution of the study participants, showing 77.0% male (77 participants) and 23.0% female (23 participants). (b) Age distribution by biological sex, illustrating the number of male and female patients across different age ranges. (c) AHI distribution by biological sex, depicting the number of male and female patients with various AHI values (average number of apneic episodes per sleeping hour).
Interestingly, the distribution of the Apnea-Hypopnea Index (AHI) spans a similar range for both male and female participants (Fig. 4c). This suggests that despite the significant imbalance between different groups, the dataset provides comprehensive information on apnea/hypopnea episodes across a broad spectrum of AHI values, encompassing all SAS severity classes and a wide age range. It is important to note that the statistical measures presented here should not be interpreted as epidemiological data but rather as indicative features of the dataset’s balance. The labeled apnea/hypopnea events were further classified into specific types: “Obstructive Apnea,” “Central Apnea,” “Mixed Apnea,” and “Hypopnea,” among others. The majority (80.8%) of the annotated respiratory events (totaling 15,462 episodes) were either Apnea (13,412 events) or Hypopnea (13,601 episodes). Obstructive sleep apnea episodes were the most frequent among the apneic episodes, accounting for 11.6% of the total, while central apnea episodes comprised 5.6% of the labeled episodes (See Fig. 5a).
(a) Percentage distribution of various respiratory events. The pie chart illustrates the proportion of Hypopnea (80.8%), Obstructive Apnea (11.6%), Central Apnea (5.6%), Mixed Apnea (0.8%), RERA, Body Event, Obstructive Hypopnea, and Central Hypopnea within the dataset. (b) Distribution of the average count of apnea episodes per sleeping hour across different apnea severity categories (Normal, Mild Apnea, Moderate Apnea, Severe Apnea). The bar chart shows the number of patients for each category, indicating the distribution of apnea rates among the study participants. (c) Distribution of the average count of hypopnea episodes per sleeping hour across different apnea severity categories (Normal, Mild Apnea, Moderate Apnea, Severe Apnea). The bar chart shows the number of patients for each category, highlighting the variability in hypopnea rates.
The criteria for labeling various apnea episodes are well-defined in the sleep apnea scoring protocol. These criteria are based on airflow signals, measured through pressure drops and air thermal changes near the patient’s nose, as well as thoracoabdominal movements indicating breathing effort. Additional factors, such as oxygen desaturation and potential arousal indicated by neurological signals, are also used to accurately identify apneic episodes. These criteria are consistently applied to all patients. (See Fig. 5(b,c)) shows the distribution of the four main categories of apneic events (“Obstructive,” “Central,” “Mixed,” and “Hypopnea”) per hour of sleep, in relation to the patient’s overall diagnosis (SAS severity class: “Severe,” “Moderate,” “Mild,” and “Normal”)24. As expected with the prevalence of central apnea syndrome, the frequency of central and mixed apnea episodes falls within the 0-5 and 0-20 apneas per hour range, respectively, regardless of the patient’s severity classification. Hypopnea events are particularly frequent in moderately apneic individuals, though this observation is based on a limited number of moderate apneic subjects in the dataset and requires further investigation. (See Fig. 5b) and (See Fig. 5c) compare the Apnea Index (AI) and Hypopnea Index (HI). The AI is calculated as the sum of all types of apnea events per hour of sleep, while the HI includes only hypopnea episodes per hour of sleep.
Sleep stage classification for PSG data validation
We have used state-of-the-art deep learning models as a baseline sleep stage classifier and included them in the codebase to demonstrate the technical quality as well as the potential utility of the dataset, especially the PSG data. These algorithms predict the sleep stages (W, N1, N2, N3 & REM) based on 30 seconds of raw EEG (C4:M1 or C4:A1) and EOG (E1:M2 or EOG1:A2) channels after they are down-sampled to 100 Hz.
Evaluation Metrics & Results
In our evaluation, we used a robust set of metrics to thoroughly assess our proposed method’s performance in automatically classifying sleep stages. We employed accuracy (ACC), kappa (κ), and F1-scores (F1) to gauge our model’s proficiency. We examined both the individual F1-scores for each sleep stage and the overall categorization effectiveness using the macro-averaged F1-score (MF1), providing a balanced evaluation across all sleep stages.
Table 3 (See Table 3) reports the 10-fold stratified cross-validation results on 100 subject sleep studies using single-channel EEG and EOG signals. To complement the quantitative metrics, Fig. 3 (See Fig. 3) presents side-by-side hypnograms of an expert annotation and the model’s prediction, visually illustrating the temporal agreement between ground truth and automatic classification. Deep learning models perform well on healthy datasets22 but show reduced performance in stroke patients, underscoring the need for algorithms adapted to altered sleep physiology.
Data availability
All the 100 PSGs included in this dataset have been saved in standard EDF format. At the same time, the annotation data has been provided in a spreadsheet file and is available through Zenodo23 the Data Foundation portal (website: https://india-data.org/dataset-details/0b801dfa-4e42-4ec6-9c56-c6892b907ed2). This dataset can be combined with other datasets to be used for training machine learning models and educational purposes. Despite its valuable applications, the dataset has limitations. There is a class imbalance that can affect the training and generalization of machine learning models.
Code availability
The code for pre-processing and deep learning models is publicly available on GitHub at: (Website: https://github.com/suvadeepmaiti/iSLEEPS)
References
Tononi, G. & Cirelli, C. Sleep and synaptic homeostasis: a hypothesis. Brain Research Bulletin 62(2), 143–150, https://doi.org/10.1016/j.brainresbull.2003.09.004 (2003).
Morin, C. M. et al. Insomnia disorder. Nature Reviews Disease Primers 1, 15026, https://doi.org/10.1038/nrdp.2015.26 (2015).
Jennum, P. C. & Riha, R. L. Epidemiology of sleep apnoea/hypopnoea syndrome and sleep-disordered breathing. European Respiratory Journal 33(4), 907–914, https://doi.org/10.1183/09031936.00180108 (2009).
Pengpid, S. & Peltzer, K. Prevalence and correlates of insomnia symptoms among older adults in India: Results of a national survey in 2017-2018. Archives of Mental Health 22(2), 139–147, https://doi.org/10.4103/amh.amh_19_21 (2021).
Datta, K., Bhutambare, A. & Mallick, H. N. Wake-up Call for India on Sleep Health: Systematic Review Findings. Indian Journal of Public Health 69(4), 614-621, https://doi.org/10.4103/ijph.ijph_504_24 (2025).
Reddy, P. et al. Epidemiology of obstructive sleep apnea in India: systematic review and meta-analysis. Sleep Medicine 100, 25–34, https://doi.org/10.1016/j.smrv.2023.101829 (2023).
Plomaritis, P. et al. Sleep-disordered breathing in acute stroke: a single-center, prospective, longitudinal study. Journal of Clinical Medicine 12(3), 986, https://doi.org/10.3390/jcm12030986 (2023).
Tanayapong, P. & Kuna, S. T. Sleep disordered breathing as a cause and consequence of stroke: a review of pathophysiological and clinical relationships. Sleep Medicine Reviews 59, 101499, https://doi.org/10.1016/j.smrv.2021.101499 (2021).
Baglioni, C. et al. Polysomnographic characteristics of sleep in stroke: a systematic review and meta-analysis. PLoS One 11(3), e0148496, https://doi.org/10.1371/journal.pone.0148496 (2016).
Srijithesh, P. R. et al. Evolution of Sleep Apnea and Arousals in Patients with Ischemic Stroke: A Longitudinal Follow-Up Study. Annals of Indian Academy of Neurology 27(6), 651–656, https://doi.org/10.4103/aian.aian_439_24 (2024).
Mythirayee, S. et al. Utility of Sleep Questionnaires for Detecting Sleep Apnea in Ischemic Stroke Patients. Annals of Indian Academy of Neurology 28(2), 241–246, https://doi.org/10.4103/aian.aian_730_24 (2025).
Hermann, D. M. & Bassetti, C. L. Sleep-related breathing and sleep-wake disturbances in ischemic stroke. Neurology 73(16), 1313–1322, https://doi.org/10.1212/wnl.0b013e3181bd137c (2009).
Parra, O. et al. Time course of sleep-related breathing disorders in first-ever stroke or transient ischemic attack. American Journal of Respiratory and Critical Care Medicine 161(2), 375–380, https://doi.org/10.1164/ajrccm.161.2.9903139 (2000).
Penzel, T., Moody, G. B., Mark, R. G., Goldberger, A. L. & Peter, J. H. The apnea-ECG database. Computers in Cardiology 2000. Vol. 27. IEEE; 255–258. https://doi.org/10.1109/CIC.2000.898505 (2000).
Bernardini, A., Brunello, A., Gigli, G. L., Montanari, A. & Saccomanno, N. OSASUD: A dataset of stroke unit recordings for the detection of Obstructive Sleep Apnea Syndrome. Scientific Data 9(1), 177, https://doi.org/10.1038/s41597-022-01272-y (2022).
St. Vincent’s University Hospital; University College Dublin. St. Vincent’s University Hospital / University College Dublin Sleep Apnea Database (Version 1.0.0). PhysioNet https://doi.org/10.13026/C26C7D (2007).
Juliá-Serdá, G., Navarro-Esteva, J. & Ravelo-García, A. G. HuGCDN2014–OXI Sleep Apnea Database (Version 1.0). Mendeley Data https://doi.org/10.17632/cdxs63gdzc.1 (2023).
Ott, S. R. et al. SAS Care 1: sleep-disordered breathing in acute stroke and transient ischaemic attack-“prevalence, evolution and association with functional outcome at 3 months, a prospective observational polysomnography study. ERJ Open Research. 6(2). https://doi.org/10.1183/23120541.00334-2019 (2020).
Maiti, S. Toward Wearable Sleep Quality Assessment: Supervised Deep Learning for Sleep Stage Classification (Master’s Thesis). International Institute of Information Technology - Hyderabad (IIIT-H), India. Available at: https://is.gd/b9YbNq (2024).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition. 770–778, https://doi.org/10.48550/arXiv.1512.03385 (2016)
Maiti, S., Sharma, S. K. & Bapi, R. S. Enhancing healthcare with EOG: A novel approach to sleep stage classification. In: ICASSP 2024-IEEE International Conference on Acoustics, Speech and Signal Processing. p. 2305–2309, https://doi.org/10.1109/ICASSP48485.2024.10446703 (IEEE; 2024).
Sharma, S., Maiti, S., Mythirayee, S., Rajendran, S. & Bapi, R. S. Transparency in sleep staging: Deep learning method for EEG sleep stage classification with model interpretability. arXiv preprint arXiv:2309.07156. https://doi.org/10.48550/arXiv.2309.07156 (2023).
Maiti, S., Mythirayee, S., Bapi, R. S., Srijithesh, P. R. & Sharma, S. iSLEEPS: Polysomnography Dataset for Sleep Analysis in Indian Ischemic Stroke Patients. Zenodo. https://doi.org/10.5281/zenodo.14873844 (2025).
Korompili, G. et al. PSG-Audio, a scored polysomnography dataset with simultaneous audio recordings for sleep apnea studies. Scientific Data 8(1), 197, https://doi.org/10.1038/s41597-021-00977-w (2021).
Author information
Authors and Affiliations
Contributions
Conceptualization: S. Rajendran and R.S. Bapi; Methodology and formal analysis: S. Maiti and S.K. Sharma; Investigation: S. Maiti, S.K. Sharma and S. Mythirayee; Data curation: S. Mythirayee; Writing - Original Draft: S. Maiti, S.K. Sharma and S. Mythirayee; Writing - Review & Editing: S. Rajendran and R.S. Bapi; Supervision: S. Rajendran and R.S. Bapi. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Maiti, S., Sharma, S.K., Mythirayee, S. et al. Polysomnography Dataset for Sleep Analysis in Ischemic Stroke Patients. Sci Data 13, 421 (2026). https://doi.org/10.1038/s41597-026-06747-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-06747-w