Abstract
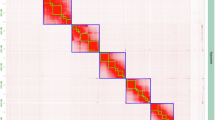
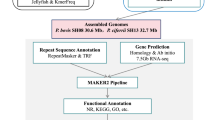
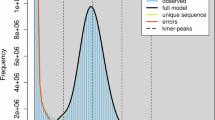
Prototheca wickerhamii is a non-photosynthetic microalgal species that has been implicated in opportunistic human infections. Understanding its genomic features is crucial for both medical applications and symbiosis research. We generated high-quality genome assemblies for two strains of Prototheca wickerhamii, Pw26 and PwS1, using PacBio HiFi reads. The assemblies were evaluated for completeness and accuracy using BUSCO analysis. The assembled genomes for Pw26 and PwS1 were 17.8 MB and 17.4 MB, respectively, with contig N50 values of 1.6 MB. The number of assembled contigs is closely related to the number of chromosomes. The GC content was 63.5% for both genomes. Comparative analysis showed high similarity in genome size and alignment, with Pw26 having slightly more protein-coding genes (46,394) than PwS1 (44,702). Repeat sequences accounted for 6.03% and 4.18% of the genomes in Pw26 and PwS1, respectively. These high-quality genome assemblies provide a valuable resource for comparative genomics and functional exploration of Prototheca wickerhamii. The detailed genomic characterization supports further studies on pathogenic mechanisms.
Similar content being viewed by others
Data availability
All data related to the genome of P. wickerhamii strain PwS1 and Pw26 are available through the following databases or links. Sequence Read Archive (SRA) data was uploaded in NCBI with project ID of PRJNA1314384. The HiFi long reads sequencing data for P. wickerhamii Pw26 and PwS1 have been deposited in the SRA, with accession numbers SRR35231276 and SRR35231275, respectively. The whole-genome shotgun projects for strains Pw26 and PwS1 have been deposited in GenBank under accessions JBTKVZ00000000032 and JBTKVY00000000033, correspondingly. Additionally, SRA data link is https://identifiers.org/ncbi/insdc.sra:SRP616729. The Figshare link: https://doi.org/10.6084/m9.figshare.30030796.v1.
Code availability
This study did not involve the development of any specific code. The data analyses were conducted in accordance with the protocols outlined in the Methods section.
References
Masuda, M. et al. Protothecosis in Dogs and Cats-New Research Directions. Mycopathologia. 186(1), 143–152 (2021).
Kano, R. Emergence of Fungal-Like Organisms: Prototheca. Mycopathologia. 185(5), 747–754 (2020).
Guo, J. et al. Integration of transcriptomics, proteomics, and metabolomics data for the detection of the human pathogenic Prototheca wickerhamii from a One Health perspective. Frontiers in cellular and infection microbiology. 13, 1152198 (2023).
Bakuła, Z. et al. A first insight into the genome of Prototheca wickerhamii, a major causative agent of human protothecosis. BMC genomics. 22(1), 168 (2021).
Guo, J. et al. Genome Sequences of Two Strains of Prototheca wickerhamii Provide Insight Into the Protothecosis Evolution. Frontiers in cellular and infection microbiology. 12, 797017 (2022).
Lass-Flörl, C. & Mayr, A. Human protothecosis. Clinical microbiology reviews. 20(2), 230–242 (2007).
Urban, M. et al. PHI-base: the pathogen-host interactions database. Nucleic acids research. 48(D1), D613–d620 (2020).
Wolff, G., Plante, I., Lang, B. F., Kück, U. & Burger, G. Complete sequence of the mitochondrial DNA of the chlorophyte alga Prototheca wickerhamii. Gene content and genome organization. Journal of molecular biology. 237(1), 75–86 (1994).
Bakuła, Z. et al. Sequencing and Analysis of the Complete Organellar Genomes of Prototheca wickerhamii. Frontiers in plant science. 11, 1296 (2020).
Zhang, Q. Q., Zhu, L. P., Weng, X. H., Li, L. & Wang, J. J. Meningitis due to Prototheca wickerhamii: rare case in China. Medical mycology. 45(1), 85–88 (2007).
Li, J., Huang, Z. & Zhang, R. Unmasking Prototheca wickerhamii: A rare case of cutaneous infection and its implications for clinical practice. The Brazilian journal of infectious diseases: an official publication of the Brazilian Society of Infectious Diseases. 29(3), 104525 (2025).
Etchecopaz A. N., Del Vecchio L., Álvarez C., Mesplet M. & Cuestas M. L. Cytological and microbiological analysis of a Prototheca wickerhamii infection in a cat with cutaneous lesions successfully treated with intralesional amphotericin B. The Journal of small animal practice (2025).
Guo J. et al. Two high-quality genomes of Prototheca bovis strain SH08 and Prototheca ciferrii strain SH13. Scientific data. (2025).
Jagielski, T. et al. Occurrence of Prototheca Microalgae in Aquatic Ecosystems with a Description of Three New Species, Prototheca fontanea, Prototheca lentecrescens, and Prototheca vistulensis. Applied and environmental microbiology. 88(22), e0109222 (2022).
Mareso, C. et al. Optimization of long-range PCR protocol to prepare filaggrin exon 3 libraries for PacBio long-read sequencing. Molecular biology reports. 50(4), 3119–3127 (2023).
Ortigas-Vasquez, A. et al. High-fidelity long-read sequencing of an avian herpesvirus reveals extensive intrapopulation diversity in tandem repeat regions. PLoS pathogens. 21(8), e1013435 (2025).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27(2), 573–580 (1999).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA. 6, 11 (2015).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics. 21(Suppl 1), i351–358 (2005).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenetic and genome research. 110(1-4), 462–467 (2005).
Stanke, M., Schöffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. BMC Bioinformatics. 7, 62 (2006).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics. 12(1), 491 (2011).
Yang, Z. et al. Convergent horizontal gene transfer and cross-talk of mobile nucleic acids in parasitic plants. Nature Plants. 5(9), 991–1001 (2019).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20(1), 278 (2019).
Zhu, M., Wang, X. & Li, X. Genome-wide identification and expression analysis of glutamate receptor-like genes in three Dendrobium species. Biochimica et biophysica acta General subjects. 1869(6), 130789 (2025).
Zuo, W. & Wang, Z. Identification of ulcerative colitis diagnostic markers from differentially expressed genes shared with Hirschsprung disease. Scientific reports. 15(1), 11274 (2025).
Borza, T., Popescu, C. E. & Lee, R. W. Multiple metabolic roles for the nonphotosynthetic plastid of the green alga Prototheca wickerhamii. Eukaryotic cell. 4(2), 253–261 (2005).
Zou, C. et al. Genome and transcriptome wide association study identify candidate genes regulating folate levels in maize. Frontiers in plant science. 16, 1606220 (2025).
Zhu, J. et al. Genome-wide association study and transcriptomic analysis reveal the crucial role of sting1 in resistance to visceral white-nodules disease in Larimichthys polyactis. Frontiers in immunology. 16, 1562307 (2025).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR35231276 (2026).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR35231275 (2026).
Fang, L. et al. High-Quality Genome Assemblies of Two Prototheca wickerhamii Strains, https://www.ncbi.nlm.nih.gov/nuccore/JBTKVZ000000000 (2026).
Fang, L. et al. High-Quality Genome Assemblies of Two Prototheca wickerhamii Strains, https://www.ncbi.nlm.nih.gov/nuccore/JBTKVY000000000 (2026).
Fang, L. et al. High-Quality Genome Assemblies of Two Prototheca wickerhamii Strains. figshare https://doi.org/10.6084/m9.figshare.30030796.v1 (2026).
Zhou, Q. et al. Telomere-to-telomere gapless genome assembly of the giant grouper (Epinephelus lanceolatus). Scientific data. 11(1), 1342 (2024).
Zuo, W. et al. Whole genome sequencing of a multidrug-resistant Bacillus thuringiensis HM-311 obtained from the Radiation and Heavy metal-polluted soil. Journal of global antimicrobial resistance. 21, 275–277 (2020).
Qian, W. et al. Identification of novel single nucleotide variants in the drug resistance mechanism of Mycobacterium tuberculosis isolates by whole-genome analysis. BMC genomics. 25(1), 478 (2024).
Zhou, X., Wang, E., Xu, X. & Zhang, B. Chromosome-level genome assembly of Phytoseiulus persimilis Athias-Henriot. Scientific data. 12(1), 293 (2025).
Sodmann, A. et al. Human dorsal root ganglia are either preserved or completely lost after deafferentation by brachial plexus injury. British journal of anaesthesia. 133(6), 1250–1262 (2024).
Acknowledgements
This work was financially supported by the STU Scientific Research Initiation Grant (NTF25030T), Sanming Project of Shenzhen Longhua Distrcit Central Hospital and Municipal Financial Subsidy of Shenzhen Longhua District Key Medical Discipline Construction.
Author information
Authors and Affiliations
Contributions
J. Jian conceived the study. J. Guo collected the samples, conducted experiments, L. Fang, J. Jian and Y. Luo performed bioinformatics analysis. L. Fang, Q. Ning, J. Jian wrote the manuscript. J. Guo and J. Ning provided suggestion and revised the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Fang, L., Guo, J., Ning, Q. et al. High-Quality Genome Assemblies of Two Prototheca wickerhamii Strains. Sci Data (2026). https://doi.org/10.1038/s41597-026-06916-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-06916-x