Abstract
Although several maize genome assemblies are publicly available, those of lines important to European breeding programs are underrepresented. Using PacBio long-read sequencing, we assembled high-quality chromosome-level genomes of 29 key lines of European breeding relevance, encompassing Northern flint and European flint lines used for adaptation to Northern European climate, lines derived from European landraces of tropical origin, and American temperate dent lines adapted to European regions. Genome assembly sizes range from 2.17 to 2.35 gigabases, with scaffold N50s ranging from 219 to 254 megabases. Completeness assessment revealed BUSCO scores ranging from 97.7 to 98.5 and merqury completeness scores ranging from 96.62 to 98.30. Calling structural variants and SNPs relative to the B73 reference sequence revealed the expected separation of inbred groups. Flint lines contribute the highest number of novel variants, thus emphasizing the importance of sequencing flint material to complete the maize pangenome. These high-quality genome assemblies therefore provide new opportunities to understand the dynamics of maize structural variation, and to identify the functional variations underlying maize phenotypic diversity.
Similar content being viewed by others
Background & Summary
Maize (Zea mays ssp. mays) is known for its large genetic diversity, which allowed the species to adapt to a multitude of environments, including tropical and temperate climates. Maize is now grown throughout the world and is the cereal with the highest production worldwide1. Its extensive genetic and phenotypic variation has also been the foundation of modern hybrid breeding. In the U.S., complementary heterotic groups within the dent germplasm - Stiff Stalk Synthetic and non-Stiff Stalk Synthetic, including Lancaster and Iodent lines - have been developed to generate highly productive hybrids, while in Europe, heterotic effects between dent and flint lines have been exploited to develop productive hybrids adapted to cooler climate. In addition to its role as a major food crop, maize is also a model organism in biology, particularly for genome dynamics, due to its large amount of intra-specific structural variation2 and its massive transposable elements content3,4. The discovery that non-coding polymorphisms contribute significantly to a wide range of phenotypic traits5 also led to the establishment of maize as a model for the study of gene expression regulation6,7,8, including the integration of cis-regulatory elements into gene regulatory networks9. Characterizing the genomic diversity of maize is essential for understanding the contribution of structural variants to this diversity, and is a prerequisite to underpinning the functional variation underlying phenotypic variation. Near complete high quality chromosome-scale genome assemblies are critical resources to address these questions.
Despite this wide genetic diversity, for decades, most knowledge about the genomic structure and function of maize has been obtained from a single genotype, B73, an American temperate dent line, therefore representing only a subset of the genetic variability and biology of the species, with a bias towards genetics of the Stiff Stalk Synthetic germplasm. In the past years, efforts have been made to de novo assemble full genome sequences of several other maize lines10,11,12,13,14, including flint material of interest for Europe15,16. While providing first insights into maize structural variation, these studies nevertheless remained limited in characterizing the maize pangenome, as they were generated by different laboratories, using different assembly and annotation strategies. This issue has been overcome by the production of a pangenome analysis of a set of 26 founder inbred lines representing a large fraction of maize diversity, including lines from temperate, subtropical and tropical origin, as well as lines from sweet corn and popcorn germplasm17. The production of high-quality assemblies with high contiguity over repetitive regions revealed large amounts of structural variants. Although most of the variants discovered were in high linkage disequilibrium with SNPs, over 6% of the genomic regions found associated with phenotype were solely detected with structural variants and not with SNPs, indicating their biological relevance and their agronomic value. The cumulative number of pan genes found from this set of 26 lines did not reach a plateau, highlighting the need to explore more extensively genome sequences of the maize germplasm to discover the entire set of maize genes. In particular, the absence of flint material in this dataset hampered a global analysis of the maize germplasm and likely caused an under-appreciation of maize genetic variation. This also limits the use of this pangenome for breeding programs using flint material.
In this study, we expand the current collection of maize whole-genome assemblies by generating high-quality PacBio HiFi-based assemblies for 29 key inbred lines of major relevance to European breeding programs. These include Northern and European flint lines used for adaptation to Northern European climates, inbred lines derived from European landraces of tropical origin, and American dent lines that complete the diversity of the 26 American founder lines (see Table 1).
Methods
Sample collection and genomic DNA extraction
Plants were grown in standard conditions (growth chamber) up to emergence, then moved to obscurity for 2 to 5 days. Young etiolated leaf samples were flash frozen in liquid nitrogen upon collection. Leaf DNA extractions were carried out using three different protocols: EZNA SQ plant kit (Omega, D3095), Mayjonade et al.18 and Nucleobond HMW DNA Kit (Macherey-Nagel, Ref: 740160.20). The protocol used was tracked for each sample and can be found in the DNA samples metadata. DNA was quantified using the Qubit fluorimetry system, with the High Sensitivity kit (Thermo Fisher, Q32854). Fragment size distributions were assessed using the Agilent Fragment Analyzer. Purity measurements were performed using a Thermo Fisher Nanodrop system, thus ensuring absence of contaminants.
Genome sequencing
Generation of HIFI reads using PacBio Sequel II - CCS
Library preparation was performed according to the manufacturer’s instructions “Procedure & Checklist Preparing HiFi SMRTbell Libraries using SMRTbell Express Template Prep Kit 2.0 or 3. 0”. 5 to 10 μg of DNA was purified and sheared to reach 20kb size using the Megaruptor3 system (Diagenode). Size selection with a 10–15 kb cutoff was performed on the BluePippin Size Selection system or the Pippin HT system (Sage Science). Libraries were sequenced on 2 to 4 SMRTcells on a Sequel II instrument with a 2 hours pre-extension and a 30 hours movie, aiming to reach a 25X HIFI reads genome coverage.
Hi-C library preparation and sequencing
Hi-C libraries were prepared from the F2, F4, F252 and MBS847 samples, using isolated nuclei as starting material. The nuclei were obtained from 1g of young leaves, following the method described in Workman et al.19. All nuclei obtained where then fixed in 1.5% formaldehyde and used to perform Hi-C using the Dovetail Hi-C Kit according to the manufacturer’s protocol (Ref: DG-HiC). Briefly, fixed in situ chromatin was digested with DpnII, DNA ends were labeled with Biotin and proximity ligation was performed. After reverse-crosslinking, 1 μg of purified DNA was then sheared to reach a mean fragment size of ~550 bp (Covaris) and used to build a sequencing library using Illumina adapters. Biotin-containing fragments were isolated using M280 streptavidin Dynabeads (Invitrogen) before PCR enrichment of the library (10 PCR cycles). The libraries were sequenced on an Illumina NovaSeq6000 platform to generate 2 × 150 bp pair-end reads, producing a minimum of 48 Gb of Hi-C read data per library.
Genome sequence assembly and validation
Genome sequence assemblies were performed in two consecutive steps, first building contigs from HiFi reads, then organizing these contigs into chromosomes. For a first set of 4 lines, contigs were scaffolded using Hi-C data. These lines were chosen to represent material with various degree of relatedness to B73: two non stiff stalk lines belonging to two different subgroups (F252 and MBS847), and two flint lines representing European flints (F2) and Northern flints (F4). We observed no major rearrangements as compared to B73 for any of the assembled genome sequences (see Supplementary Fig. 1 for a genome comparison illustration using D-GENIES20), and all these were included within contigs. This indicates that our contig length was large enough to ensure good scaffolding using B73 as a reference. We therefore generated reference-guided assemblies for all other inbred lines using B73v5 sequence as reference.
Contig assembly
HiFi reads were assembled in contigs with hifiasm21 version 0.16.1 using default parameters. Contig assembly metrics were generated using the assemblathon_stats.pl script found at https://github.com/KorfLab/Assemblathon.
Contig scaffolding
For F2, F4, F252 and MBS847 lines, Illumina Hi-C reads were aligned onto the contigs with Juicer22, and contigs were scaffolded with 3D-DNA23. Resulting contact maps were manually corrected with Juicebox24. For all three software packages, default parameters were used. Read quantity, read coverage and Hi-C link metrics are presented in Table 6. For all other maize lines, contig sets were scaffolded with ragtag25 version 2.0.1 using default parameters, using the Zm-B73-REFERENCE-NAN-5.0.fa sequence as reference, downloaded from the NCBI website https://www.ncbi.nlm.nih.gov/datasets/genome/GCF_902167145.1/. For each maize line, contigs were organized into 10 pseudo-chromosomes, with unplaced contigs corresponding to only 0.9 to 7.2% of the assembly total length.
Scaffold validation
Scaffold metrics were produced using the assemblathon_stats.pl script26 and the BUSCO (Benchmarking Universal Single-Copy Orthologs)27 metrics with version 5.1.2 using the poales_odb10 lineage. Kmer completeness and sequence quality value of the scaffolds were assessed using Merqury28 version 1.3 with default parameters.
SNPs and structural variants detection
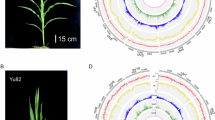
SNPs and structural variants were detected from the raw HiFi reads, aligning the fastq reads from each maize line to the maize reference assembly B73_RefGen_v4 using pbmm2 (https://github.com/PacificBiosciences/pbmm2) with the CSS preset flag. SNPs were detected using DeepVariant (1.3.0) using default parameters (see snp_detection rules in https://github.com/SeqOccin-SV/SeqOccinVariants). Structural variants were detected using the Sniffles29 (https://github.com/fritzsedlazeck/Sniffles) in a two round process. Sniffles was first used to detect variant on an individual basis with the following parameters (–minsupport 12 –minsvlen 100 –max-splits-base 2 –max-splits-kb 0 –min-alignment-length 5000 –minsvlen 20) with default values for the other parameters. The resulting vcf files were filtered to keep only variant with PASS filter and merged using the jasmine software30. BND (breakend) and TRA (translocation) variants were filtered out and the merged SVs were provided as input (–genotype-vcf) to Sniffles along with the BAM files on each individual line, leading to a set of SV genotyped on all the individuals (see Fig. 1).
Quantities of structural variants detected for each inbred line as compared to B73.
Data Records
Reads and assembled genome sequences were deposited in European National Archive under bioproject PRJEB6781231, (see Tables 2–6 for details). SNPs and structural variants data were deposited in the European Variant Archive (Study ID: PRJEB106599)32 and in the ‘Recherche Data Gouv’ repository: https://doi.org/10.57745/7AUTOL33.
Technical Validation
We produced about 2.1 to 6.9 million reads per maize line, with an average read length ranging from 12 kb to 22 kb (Table 2). These high quality HiFi reads were first used to assemble the genomes into contigs, with contig number per maize line ranging from 260 to 3084 (average 1221.1, see Table 3) and N50 contig lengths ranging from 11.8 Mb to 166.0 Mb (average 87.1 Mb, see Table 3). For each maize line, chromosome-scale scaffolds were obtained, with cumulative size of assembled chromosomes ranging from 2.18 Gb to 2.35 Gb (Table 4), in line with the genome sizes expected for maize. As anticipated, tropical lines had larger genome sizes (2.32 Gb) than temperate lines (2.25 Gb). Scaffold N50s range from 219.5 Mb to 253.8 Mb, with L50 from 4 to 5. (Table 4). To ensure the quality, integrity and accuracy of the assembled chromosome sequences generated, we carried our several validation approaches.
Completeness of genome assemblies was evaluated using BUSCO version 5.1.2 with the poales_odb10 containing 4,896 proteins, as well as with Merqury version 1.3. Metrics per genome assembly are presented in Table 5. For all assemblies, >97% of the BUSCO proteins were complete. Merqury results showed genome assemblies quality values >60 and completeness >96.62%.
To further validate the quality of the genome assemblies generated and the genotypes of the DNA sequenced, we investigated the polymorphisms (SNPs, indels and structural variants >50bp) of each line relative to reference line B73. As expected, the number of variants reflected the genetic distances of maize lines from B73 (Fig. 1). Stiff Stalk Synthetic lines showed the lowest amount of variants (7,290,142 SNPs, 829,336 indels and 68,850 SVs, Supplementary Table 2), with the lowest amounts found for lines of the B73 subgroup (Fig. 1 and Table 7). In contrast, flint lines showed the highest number of variants (14,901,375 SNPs, 1,490,896 indels and 119,558 SVs) (Supplementary Table 1). Lancaster and Iodent lines had intermediate values, with Lancaster having slightly more variants (12,365,784 SNPs, 1,282,139 indels and 107,607 SVs) than Iodents lines (11,995,606 SNPs, 1,257,735 indels and 105,935 SVs) (Supplementary Table 2). Lines of tropical origin showed slightly less variants than flint lines. Finally, a PCA based on the SNPs recapitulated the genetic groups and relationships among the lines (Fig. 2). Altogether, these results indicate the high quality of the sequences generated and the reliability of the seedlots sequenced. They also highlight the relevance of our dataset to improve knowledge on maize structural diversity, and the importance of including flint lines in sequencing programs to leverage the maize pangenome.
PCA constructed from the (standardized) genotypes of 1 millions randomly selected SNPs.
Data availability
All raw sequencing data, assembled genomes, and variant data (VCF files) have been deposited in publicly accessible repositories. The PacBio HiFi and Hi-C sequencing reads, as well as the genomes assembled from these data, have been uploaded to the European Nucleotide Archive (ENA) at www.ebi.ac.uk/ena as part of the SeqOccIn project, PRJEB600751634, and are accessible under project PRJEB6781231. Structural Variants and SNPs are available to European Variation Archive (EVA) and accessible under the accession PRJEB10659932. Variant data are linked to the nucleotide data through the sharing of a single BioSample ID. Variant data are also available at data.gouv.fr repository (https://doi.org/10.57745/7AUTOL)33.
Code availability
All the codes used for the analysis can be found on the SeqOccIn project’s GitHub page, following the path Data paper/Zea mays data paper: https://github.com/GeTPlaGe/SeqOccIn/tree/main/Data%20paper/Zeamays. The pipeline used for aligning reads and calling variants is available here: https://github.com/SeqOccin-SV/SeqOccinVariants.
References
Wrigley, C. W. & Nirmal, R. C. The major cereal grains: Corn, rice, and wheat, https://doi.org/10.1002/0471238961.23080501.a01.pub3 (2017).
Wang, Q. & Dooner, H. K. Remarkable variation in maize genome structure inferred from haplotype diversity at the bz locus. Proceedings of the National Academy of Sciences 103, 17644–17649, https://doi.org/10.1073/pnas.0603080103 (2006).
Stitzer, M. C., Anderson, S. N., Springer, N. M. & Ross-Ibarra, J. The genomic ecosystem of transposable elements in maize. PLOS Genetics 17, e1009768, https://doi.org/10.1371/journal.pgen.1009768 (2021).
Ou, S. et al. Differences in activity and stability drive transposable element variation in tropical and temperate maize. Genome Research 34, 1140–1153, https://doi.org/10.1101/gr.278131.123 (2024).
Wallace, J. G. et al. Association mapping across numerous traits reveals patterns of functional variation in maize. PLoS Genetics 10, e1004845, https://doi.org/10.1371/journal.pgen.1004845 (2014).
Zhou, P., Hirsch, C. N., Briggs, S. P. & Springer, N. M. Dynamic patterns of gene expression additivity and regulatory variation throughout maize development. Molecular Plant 12, 410–425, https://doi.org/10.1016/j.molp.2018.12.015 (2019).
Ricci, W. A. et al. Widespread long-range cis-regulatory elements in the maize genome. Nature Plants 5, 1237–1249, https://doi.org/10.1038/s41477-019-0547-0 (2019).
Marand, A. P. et al. The genetic architecture of cell type-specific cis regulation in maize. Science 388, https://doi.org/10.1126/science.ads6601 (2025).
Fagny, M. et al. Identification of key tissue-specific, biological processes by integrating enhancer information in maize gene regulatory networks. Frontiers in Genetics 11, https://doi.org/10.3389/fgene.2020.606285 (2021).
Springer, N. M. et al. The maize w22 genome provides a foundation for functional genomics and transposon biology. Nature Genetics 50, 1282–1288, https://doi.org/10.1038/s41588-018-0158-0 (2018).
Sun, S. et al. Extensive intraspecific gene order and gene structural variations between mo17 and other maize genomes. Nature Genetics 50, 1289–1295, https://doi.org/10.1038/s41588-018-0182-0 (2018).
Yang, N. et al. Genome assembly of a tropical maize inbred line provides insights into structural variation and crop improvement. Nature Genetics 51, 1052–1059, https://doi.org/10.1038/s41588-019-0427-6 (2019).
Lin, T., Song, Y., Lawrence, P., Kheshgi, H. S. & Jain, A. K. Worldwide maize and soybean yield response to environmental and management factors over the 20th and 21st centuries. Journal of Geophysical Research: Biogeosciences 126, https://doi.org/10.1029/2021jg006304 (2021).
Chen, J. et al. A complete telomere-to-telomere assembly of the maize genome. Nature Genetics 55, 1221–1231, https://doi.org/10.1038/s41588-023-01419-6 (2023).
Darracq, A. et al. Sequence analysis of european maize inbred line f2 provides new insights into molecular and chromosomal characteristics of presence/absence variants. BMC Genomics 19, https://doi.org/10.1186/s12864-018-4490-7 (2018).
Haberer, G. et al. European maize genomes highlight intraspecies variation in repeat and gene content. Nature Genetics 52, 950–957, https://doi.org/10.1038/s41588-020-0671-9 (2020).
Hufford, M. B. et al. De novo assembly, annotation, and comparative analysis of 26 diverse maize genomes. Science 373, 655–662, https://doi.org/10.1126/science.abg5289 (2021).
Mayjonade, B. et al. Extraction of high-molecular-weight genomic dna for long-read sequencing of single molecules. BioTechniques 61, 203–205, https://doi.org/10.2144/000114460 (2016).
Workman, R. et al. High molecular weight dna extraction from recalcitrant plant species for third generation sequencing v1. https://doi.org/10.1038/protex.2018.059 (2018).
Cabanettes, F. & Klopp, C. D-GENIES: dot plot large genomes in an interactive, efficient and simple way. journal = PeerJ 6, https://doi.org/10.7717/peerj.4958 (2018).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nature Methods 18, 170–175, https://doi.org/10.1038/s41592-020-01056-5 (2021).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution hi-c experiments. Cell systems 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95, https://doi.org/10.1126/science.aal3327 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for hi-c contact maps with unlimited zoom. Cell Systems 3, 99–101, https://doi.org/10.1016/j.cels.2015.07.012 (2016).
Alonge, M. et al. Automated assembly scaffolding using ragtag elevates a new tomato system for high-throughput genome editing. Genome Biology 23, https://doi.org/10.1186/s13059-022-02823-7 (2022).
Bradnam, K. R. et al. Assemblathon 2: evaluating de novo methods of genome assembly in three vertebrate species. Gigascience 2, https://doi.org/10.1186/2047-217x-2-10 (2013).
Seppey, M., Manni, M. & Zdobnov, E. M. BUSCO: Assessing Genome Assembly and Annotation Completeness, 227–245 (Springer New York, 2019).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biology 21, https://doi.org/10.1186/s13059-020-02134-9 (2020).
Smolka, M. et al. Detection of mosaic and population-level structural variants with Sniffles2. Nature Biotechnology 42, https://doi.org/10.1038/s41587-023-02024-y (2024).
Kirsche, M. et al. Jasmine and Iris: population-scale structural variant comparison and analysis. Nature Methods 20, https://doi.org/10.1038/s41592-022-01753-3 (2023).
European nucleotide archive. https://www.ebi.ac.uk/ena/browser/view/PRJEB67812 (2025).
European variant archive. https://www.ebi.ac.uk/eva/?eva-study=PRJEB106599 (2026).
The 29 maize lines SNP and SV variant set. https://doi.org/10.57745/7AUTOL (2025).
Germplasm Resources Information Network (GRIN) — doi.org. https://doi.org/10.15482/USDA.ADC/1212393.
European nucleotide archive. https://www.ebi.ac.uk/ena/browser/view/PRJEB60075 (2023).
Byrne, P. F. et al. Sustaining the future of plant breeding: The critical role of the usda-ars national plant germplasm system. Crop Science 58, 451–468, https://doi.org/10.2135/cropsci2017.05.0303 (2018).
Camus-Kulandaivelu, L. et al. Maize adaptation to temperate climate: Relationship between population structure and polymorphism in the dwarf8 gene. Genetics 172, 2449–2463, https://doi.org/10.1534/genetics.105.048603 (2006).
Bouchet, S. et al. Adaptation of maize to temperate climates: Mid-density genome-wide association genetics and diversity patterns reveal key genomic regions, with a major contribution of the vgt2 (zcn8) locus. PLoS ONE 8, e71377, https://doi.org/10.1371/journal.pone.0071377 (2013).
Acknowledgements
We thank “La Région Occitanie” and European Union for funding the project as part of the Occitanie Region’s “Regional Research and Innovation Platforms” call for projects under the FEDER-FSE MIDI-PYRENEES ET GARONNE 2014-2020 Operational Program. We thank KWS, Maisadour, Euralis, Caussade semences, Syngenta, RAGT and Limagrain for their financial support and their inputs for choosing the genetic material analyzed. We thank Valérie Combes for sample preparation, Delphine Madur and Nathalie Rivière for genotype validation, Gaëtan Givry for EVA data submission and Jorge Duarte and Johann Joets for insightful discussions on maize genome scaffolding. We are grateful to Cyril Bauland for expertise in maize germplasm accession nomenclature. We thank Carine Palaffre and French maize inbred lines seed bank (CRB, INRAE Saint Martin de Hinx), the U.S. National Plant Germplasm System (NPGS)35 and the USDA Agricultural Research Service Germplasm Resources Information Network (GRIN)36 for providing seeds with traced seedlots, as well as Adrienne Ressayre and Christine Dillmann (GQE-Le Moulon) for providing seeds of F252 and MBS847, and Silvio Salvi (University of Bologna) for early access to seeds from the GF111 inbred line, Carlotta Balconi (CREA-Research Centre for Cereal and Industrial Crops) for providing access to Lo3, and CSIC (Consejo Superior de Investigaciones Científicas) for authorizing the use of EM1197. GeT core facility https://doi.org/10.15454/1.5572370921303193E12 is supported by France Génomique National infrastructure, funded as part of “Investissement d’avenir” program managed by the French Agence Nationale pour la Recherche (contract ANR-10-INBS-09). We are grateful to the genotoul bioinformatics platform Toulouse Occitanie (Bioinfo Genotoul, https://doi.org/10.15454/1.5572369328961167E12) for providing computing and storage resources.
Author information
Authors and Affiliations
Contributions
C.D., D.M., and Ch.G. conceived and supervised the whole “SeqOccIn” project. Cl.V. and A.C. conceived the maize-related sub-project of the “SeqOccIn” project. C.D., D.M., Ch.G., Cl.V. and A.C. secured funding. C.I. coordinated data generation and quality control. C.I., C.M., C.E., E.D. produced sequence data. Ch.K., T.F. and Cl.K. supervised bioinformatic analyses. C.B., A.D.F., T.F., J.D., S.N., Cl.V. and Ch.K. analysed the results. S.P. and A.C. coordinated the selection of the inbred lines with private partners. Cl.K. and Ca.V. secured data and submitted them to public databases. C.I., Cl.V., Ch.K., S.N. and T.F. wrote the original draft of the manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Marcuzzo, C., Birbes, C., Eché, C. et al. High-quality chromosome-scale genome assemblies of 29 maize inbred lines of European breeding relevance. Sci Data 13, 715 (2026). https://doi.org/10.1038/s41597-026-07055-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41597-026-07055-z