Abstract
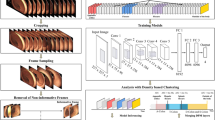
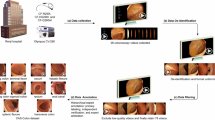
The presence of uninformative frames in colonoscopy videos is a major factor that reduces the accuracy and efficiency of various video analysis applications. To address this issue, research on informative frame classification has been conducted, but the lack of a publicly available dataset has made reproducibility difficult. In this study, we propose a novel dataset, InfoColon, which integrates video data collected from multiple medical institutions with major public colonoscopy datasets. All colonoscopy frames were labeled as either an informative frame or one of six types of uninformative frames. We also propose an active learning method to efficiently label large amounts of data with a small initial labeled dataset. Using the constructed InfoColon, we demonstrate the potential for its application in consecutive informative frame classification and 3D reconstruction. We expect that the proposed InfoColon will be valuable for various applications involving colonoscopy video analysis.
Similar content being viewed by others
Data availability
The colonoscopy videos, 7-class labels, calibration videos, and parameters for InfoColon have been uploaded and made publicly available on Synapse (https://www.synapse.org/InfoColon). Users must adhere to the data usage terms and conditions of the Synapse platform, and any research utilizing this dataset must cite the present paper.
Code availability
The code required for the data processing, model, and evaluation used in this study has been made publicly available at the following address: https://github.com/Choi-Tae-min/InfoColon.
References
Ali, S. et al. An objective comparison of detection and segmentation algorithms for artefacts in clinical endoscopy. Sci Rep 10, 2748, https://doi.org/10.1038/s41598-020-59413-5 (2020).
Koh, G. E. et al. Real-World Assessment of the Efficacy of Computer-Assisted Diagnosis in Colonoscopy: A Single Institution Cohort Study in Singapore. Mayo Clin Proc Digit Health 2, 647–655, https://doi.org/10.1016/j.mcpdig.2024.10.002 (2024).
Tang, C. P. et al. Polyp detection and false-positive rates by computer-aided analysis of withdrawal-phase videos of colonoscopy of the right-sided colon segment in a randomized controlled trial comparing water exchange and air insufflation. Gastrointest Endosc 95, 1198–1206 e1196, https://doi.org/10.1016/j.gie.2021.12.020 (2022).
Münzer, B., Schoeffmann, K. & Böszörmenyi, L. Content-based processing and analysis of endoscopic images and videos: A survey. Multimedia Tools and Applications 77, 1323–1362, https://doi.org/10.1007/s11042-016-4219-z (2018).
Tavanapong, W. et al. Artificial Intelligence for Colonoscopy: Past, Present, and Future. IEEE Journal of Biomedical and Health Informatics 26, 3950–3965, https://doi.org/10.1109/Jbhi.2022.3160098 (2022).
Ali, S. Where do we stand in AI for endoscopic image analysis? Deciphering gaps and future directions. npj Digital Medicine 5, 184 (2022).
De Carvalho, T. et al. Automated colonoscopy withdrawal phase duration estimation using cecum detection and surgical tasks classification. Biomed Opt Express 14, 2629–2644, https://doi.org/10.1364/boe.485069 (2023).
Kim, B. S. et al. Density clustering-based automatic anatomical section recognition in colonoscopy video using deep learning. Scientific Reports 14, 872, https://doi.org/10.1038/s41598-023-51056-6 (2024).
Batlle, V. M., Montiel, J. M. M. & Tardós, J. D. Photometric single-view dense 3D reconstruction in endoscopy. In Proc. IEEE/RSJ Int. Conf. Intell. Robots Syst. (IROS), 4904–4910 https://doi.org/10.1109/Iros47612.2022.9981742 (2022).
Li, Q., Yang, S., Shen, D. & Jin, Y. Free-dygs: Camera-pose-free scene reconstruction based on gaussian splatting for dynamic surgical videos. arXiv e-prints, arXiv: 2409.01003 (2024).
Brachmann, E. et al. Scene Coordinate Reconstruction: Posing of Image Collections via Incremental Learning of a Relocalizer. Computer Vision - Eccv 2024, Pt Lvi 15114, 421–440, https://doi.org/10.1007/978-3-031-72992-8_24 (2025).
Frank, N., Posner, E., Muhlethaler, E., Zholkover, A. & Bouhnik, M. ColNav: Real-Time Colon Navigation for Colonoscopy. Cancer Prevention through Early Detection, Caption 2023 14295, 119–131, https://doi.org/10.1007/978-3-031-45350-2_10 (2023).
Pore, A. et al. Colonoscopy Navigation using End-to-End Deep Visuomotor Control: A User Study. 2022 Ieee/Rsj International Conference on Intelligent Robots and Systems (Iros), 9582–9588 https://doi.org/10.1109/Iros47612.2022.9981480 (2022).
Wang, K. L. et al. EndoGSLAM: Real-Time Dense Reconstruction and Tracking in Endoscopic Surgeries Using Gaussian Splatting. Medical Image Computing and Computer Assisted Intervention - Miccai 2024, Pt Vi 15006, 219–229, https://doi.org/10.1007/978-3-031-72089-5_21 (2024).
Yao, H. M., Stidham, R. W., Soroushmehr, R., Gryak, J. & Najarian, K. Automated Detection of Non-Informative Frames for Colonoscopy Through a Combination of Deep Learning and Feature Extraction. 2019 41st Annual International Conference of the Ieee Engineering in Medicine and Biology Society (Embc), 2402–2406 https://doi.org/10.1109/embc.2019.8856625 (2019).
Bashar, M. K., Kitasaka, T., Suenaga, Y., Mekada, Y. & Mori, K. Automatic detection of informative frames from wireless capsule endoscopy images. Medical Image Analysis 14, 449–470, https://doi.org/10.1016/j.media.2009.12.001 (2010).
Akbari, M. et al. Classification of Informative Frames in Colonoscopy Videos Using Convolutional Neural Networks with Binarized Weights. 2018 40th Annual International Conference of the Ieee Engineering in Medicine and Biology Society (Embc), 65–68 (2018).
Oh, J. et al. Informative frame classification for endoscopy video. Medical Image Analysis 11, 110–127, https://doi.org/10.1016/j.media.2006.10.003 (2007).
Rungseekajee, N., Lohvithee, M. & Nilkhamhang, I. Informative frame classification method for real-time analysis of colonoscopy video. Ecti-Con: 2009 6th International Conference on Electrical Engineering/Electronics, Computer, Telecommunications and Information Technology, Vols 1 and 2, 1042–1045 (2009).
van der Putten, J. et al. Informative Frame Classification of Endoscopic Videos Using Convolutional Neural Networks and Hidden Markov Models. 2019 Ieee International Conference on Image Processing (Icip), 380–384 https://doi.org/10.1109/icip.2019.8802947 (2019).
An, Y. H. et al. Informative-frame filtering in endoscopy videos. Medical Imaging 2005: Image Processing, Pt 1-3 5747, 291–302, https://doi.org/10.1117/12.595622 (2005).
Azagra, P. et al. Endomapper dataset of complete calibrated endoscopy procedures. Scientific Data 10, 671, https://doi.org/10.1038/s41597-023-02564-7 (2023).
Borgli, H. et al. a comprehensive multi-class image and video dataset for gastrointestinal endoscopy. Scientific Data 7, 283, https://doi.org/10.1038/s41597-020-00622-y (2020).
Pogorelov, K. et al. NERTHUS: A Bowel Preparation Quality Video Dataset. Proceedings of the 8th Acm Multimedia Systems Conference (Mmsys'17), 170–174. https://doi.org/10.1145/3083187.3083216 (2017).
Ma, Y. T., Chen, X. J., Cheng, K., Li, Y. & Sun, B. LDPolypVideo Benchmark: A Large-Scale Colonoscopy Video Dataset of Diverse Polyps. Medical Image Computing and Computer Assisted Intervention - Miccai 2021, Pt V 12905, 387–396, https://doi.org/10.1007/978-3-030-87240-3_37 (2021).
Jang, S. et al. SeamXSim: Seamless-textured virtual colonoscopy simulator via unpaired long-term video translation. Computers in Biology and Medicine 198, 111217 (2025).
Zhang, Z. Y. A flexible new technique for camera calibration. Ieee Transactions on Pattern Analysis and Machine Intelligence 22, 1330–1334, https://doi.org/10.1109/34.888718 (2000).
Lai, E. J., Calderwood, A. H., Doros, G., Fix, O. K. & Jacobson, B. C. The Boston bowel preparation scale: a valid and reliable instrument for colonoscopy-oriented research. Gastrointestinal Endoscopy 69, 620–625, https://doi.org/10.1016/j.gie.2008.05.057 (2009).
Lee, D.-H. Pseudo-label: The simple and efficient semi-supervised learning method for deep neural networks. In Proc. ICML Workshop on Challenges in Representation Learning, 896 (2013).
Gal, Y., Islam, R. & Ghahramani, Z. Deep Bayesian active learning with image data. In Proc. Int. Conf. Mach. Learn. (ICML), PMLR 70, 1183–1192 (2017).
Choi, T. et al. InfoColon: A Real-World Dataset for Informative Frame Classification in Colonoscopy. Synapse. https://doi.org/10.7303/syn55251782 (2025).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929 (2020).
Golhar, M. V., Bobrow, T. L., Ngamruengphong, S. & Durr, N. J. GAN Inversion for Data Augmentation to Improve Colonoscopy Lesion Classification. Ieee Journal of Biomedical and Health Informatics 29, 3864–3873, https://doi.org/10.1109/Jbhi.2024.3397611 (2025).
Nie, C., Xu, C., Li, Z. P., Chu, L. L. & Hu, Y. Specular Reflections Detection and Removal for Endoscopic Images Based on Brightness Classification. Sensors 23, 974, https://doi.org/10.3390/s23020974 (2023).
Sharma, V., Bhuyan, M. K. & Das, P. K. Can adversarial networks make uninformative colonoscopy video frames clinically informative? (student abstract). In Proc. AAAI Conf. Artif. Intell. 37, 16322–16323 (2023).
Rau, A., Bhattarai, B., Agapito, L. & Stoyanov, D. Task-Guided Domain Gap Reduction for Monocular Depth Prediction in Endoscopy. Data Engineering in Medical Imaging, Demi 2023 14314, 111–122, https://doi.org/10.1007/978-3-031-44992-5_11 (2023).
Struski, L. et al. MeVGAN: GAN-based plugin model for video generation with applications in colonoscopy. PLoS One 20, e0312038, https://doi.org/10.1371/journal.pone.0312038 (2025).
Sengupta, A. & Bartoli, A. Colonoscopic 3D reconstruction by tubular non-rigid structure-from-motion. International Journal of Computer Assisted Radiology and Surgery 16, 1237–1241, https://doi.org/10.1007/s11548-021-02409-x (2021).
Bonilla, S. et al. Gaussian Pancakes: Geometrically-Regularized 3D Gaussian Splatting for Realistic Endoscopic Reconstruction. Medical Image Computing and Computer Assisted Intervention - Miccai 2024, Pt Vi 15006, 274–283, https://doi.org/10.1007/978-3-031-72089-5_26 (2024).
Shi, Y., Lu, B., Liu, J.-W., Li, M. & Shou, M. Z. Colonnerf: Neural radiance fields for highfidelity long-sequence colonoscopy reconstruction. arXiv preprint arXiv:2312.02015 (2023).
Chavarrias-Solano, P. E., Bhattarai, B. & Ali, S. Improving NeRF representation with no pose prior for novel view synthesis in colonoscopy. In Proc. MICCAI Workshop on Data Engineering in Medical Imaging, 144–154 (2023).
Acknowledgements
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2021R1I1A3047535), and was supported by Institute of Information & communications Technology Planning & Evaluation (IITP) grant funded by the Korea government(MSIT) [NO.RS-2021-II211343, Artificial Intelligence Graduate School Program (Seoul National University)].
Author information
Authors and Affiliations
Contributions
Taemin Choi was responsible for dataset construction, experiment design, and analysis. Hee Seok Moon and Eun Hyo Jin conducted clinical video acquisition and data labeling validation. Seunghyun Jan and Chang Min Park assisted with data set construction. Dongheon Lee was responsible for research planning, overall project supervision, and guiding the methodology development. All authors were involved in the manuscript preparation and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Choi, T., Moon, H.S., Jang, S. et al. InfoColon: A dataset for consecutive informative frames in Colonoscopy. Sci Data (2026). https://doi.org/10.1038/s41597-026-07060-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07060-2