Abstract
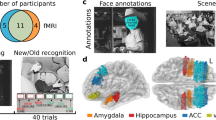
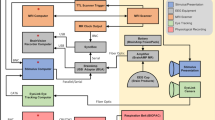
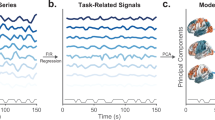
Dense, individual-level datasets are an important resource for social neuroscience that enable models of brain responses to a wide range of naturalistic features, often investigated with movie fMRI. This data release provides high spatiotemporal resolution 3 Tesla BOLD fMRI data in three healthy participants while viewing four feature-length movies with sound (Grand Budapest Hotel, Forrest Gump, Planet Earth, Jiro Dreams of Sushi; ca. 520 minutes) and a movie trailer composite, together with clip and movie repeats. All movie functional data were acquired with partial brain coverage from an approximately axial slab covering ventral prefrontal and temporal lobes, including the amygdala, with a repetition time of 556 ms and complex-valued image reconstruction. The dataset is released in both unprocessed and minimally preprocessed forms with individual, high quality anatomic templates. Preprocessed fMRI data are provided in both individual template and FreeSurfer average surface spaces. Reference raw data are provided for a conventional face-object functional localizer task. Auxiliary physiological data includes 4-lead ECG waveforms, image-estimated respiratory waveforms and pupillometry. Quality metrics include temporal SNFR maps, high-resolution B0 maps, EPI dropout and head motion parameters for all runs. Physiological noise estimation and cleaning used spatial independent component analysis and custom, per-subject component classification. Annotations for the movies provide automated low-level audiovisual and facial features, as well as emotion ratings from each participant. The presence of presumed BOLD neurovascular responses not associated with physiological or instrumentation noise was confirmed using temporally concatenated spatial independent component analysis (tcsICA).
Similar content being viewed by others
Data availability
This dataset is made available under a Creative Commons Zero (CC0) license through the OpenNeuro data sharing platform as ds00694755.
Code availability
Video presentation scripts implemented in PsychoPy 2021.2.3: https://github.com/adolphslab/DenseAmygdalaRelease.
slabpreproc slab-selective EPI preprocessing pipeline implemented in nipype: https://github.com/adolphslab/slabpreproc.
PhaseRespWave class for respiratory waveform estimation from temporally unwrapped EPI phase channel images: https://github.com/adolphslab/mriphysio.
voxface irreversible deidentification of the face region in structural MRI data: https://github.com/jmtyszka/voxface.
References
Finn, E. S. & Bandettini, P. A. Movie-watching outperforms rest for functional connectivity-based prediction of behavior. Neuroimage 235, 117963 (2021).
Finn, E. S. Is it time to put rest to rest? Trends Cogn. Sci. https://doi.org/10.1016/j.tics.2021.09.005 (2021).
Gordon, E. M. et al. Precision Functional Mapping of Individual Human Brains. Neuron https://doi.org/10.1016/j.neuron.2017.07.011 (2017).
Fedorenko, E. The early origins and the growing popularity of the individual-subject analytic approach in human neuroscience. Current Opinion in Behavioral Sciences 40, 105–112 (2021).
Naselaris, T., Allen, E. & Kay, K. Extensive sampling for complete models of individual brains. Current Opinion in Behavioral Sciences 40, 45–51 (2021).
Poldrack, R. A. et al. Long-term neural and physiological phenotyping of a single human. Nat. Commun. 6, 8885 (2015).
Adolphs, R. What does the amygdala contribute to social cognition? - Adolphs - 2010 - Annals of the New York Academy of Sciences - Wiley Online Library. Ann. N. Y. Acad. Sci. https://doi.org/10.1111/j.1749-6632.2010.05445.x/pdf (2010).
Varkevisser, T., Geuze, E. & van Honk, J. Amygdala fMRI-A critical appraisal of the extant literature. Neurosci. Insights 19, 26331055241270591 (2024).
West, H. V., Burgess, G. C., Dust, J., Kandala, S. & Barch, D. M. Amygdala activation in cognitive task fMRI varies with individual differences in cognitive traits. Cogn. Affect. Behav. Neurosci. 21, 254–264 (2021).
Canli, T., Sivers, H., Whitfield, S. L., Gotlib, I. H. & Gabrieli, J. D. E. Amygdala response to happy faces as a function of extraversion. Science 296, 2191 (2002).
Byrge, L. et al. Video-evoked fMRI BOLD responses are highly consistent across different data acquisition sites. Hum. Brain Mapp. https://doi.org/10.1002/hbm.25830 (2022).
Meer, J. N., van der, Breakspear, M., Chang, L. J., Sonkusare, S. & Cocchi, L. Movie viewing elicits rich and reliable brain state dynamics. Nat. Commun. 11, 5004 (2020).
Sonkusare, S., Breakspear, M. & Guo, C. Naturalistic stimuli in neuroscience: Critically acclaimed. Trends Cogn. Sci. 23, 699–714 (2019).
Vanderwal, T., Eilbott, J. & Castellanos, F. X. Movies in the magnet: Naturalistic paradigms in developmental functional neuroimaging. Dev. Cogn. Neurosci. 36, 100600 (2019).
Saarimäki, H. Naturalistic Stimuli in Affective Neuroimaging: A Review. Front. Hum. Neurosci. 15, 675068 (2021).
Hanke, M. et al. A high-resolution 7-Tesla fMRI dataset from complex natural stimulation with an audio movie. Sci Data 1, 140003 (2014).
Visconti di Oleggio Castello, M., Chauhan, V., Jiahui, G. & Gobbini, M. I. An fMRI dataset in response to “The Grand Budapest Hotel”, a socially-rich, naturalistic movie. Sci Data 7, 383 (2020).
Morgenroth, E. et al. Emo-FilM: A multimodal dataset for affective neuroscience using naturalistic stimuli. Sci. Data 12, 684 (2025).
Aliko, S., Huang, J., Gheorghiu, F., Meliss, S. & Skipper, J. I. A naturalistic neuroimaging database for understanding the brain using ecological stimuli. Sci Data 7, 347 (2020).
Poldrack, R. A. Precision neuroscience: Dense sampling of individual brains. Neuron 95, 727–729 (2017).
Hanke, M. et al. A studyforrest extension, simultaneous fMRI and eye gaze recordings during prolonged natural stimulation. Sci Data 3, 160092 (2016).
Agtzidis, I., Meyhöfer, I., Dorr, M. & Lencer, R. Following Forrest Gump: Smooth pursuit related brain activation during free movie viewing. Neuroimage 216, 116491 (2020).
Jiahui, G. et al. Predicting individual face-selective topography using naturalistic stimuli. Neuroimage 216, 116458 (2020).
Jiahui, G. et al. Modeling naturalistic face processing in humans with deep convolutional neural networks. Proc. Natl. Acad. Sci. USA. 120, e2304085120 (2023).
Bernal-Rusiel, J. L. et al. Statistical analysis of longitudinal neuroimage data with Linear Mixed Effects models. Neuroimage 66, 249–260 (2013).
Esteban, O. et al. fMRIPrep: a robust preprocessing pipeline for functional MRI. Nat. Methods 16, 111–116 (2019).
Esteban, O., Markiewicz, C. J. & Goncalves, M. SDCflows: Susceptibility Distortion Correction workFLOWS. https://doi.org/10.5281/ZENODO.12654387 (Zenodo, 2024).
Drew, P. J. Vascular and neural basis of the BOLD signal. Curr. Opin. Neurobiol. 58, 61–69 (2019).
Hillman, E. M. C. Coupling mechanism and significance of the BOLD signal: a status report. Annu. Rev. Neurosci. 37, 161–181 (2014).
Epp, S. M. et al. BOLD signal changes can oppose oxygen metabolism across the human cortex. Nat. Neurosci. 1–12 https://doi.org/10.1038/s41593-025-02132-9 (2025).
Visconti di Oleggio Castello, M., Haxby, J. V. & Gobbini, M. I. Shared neural codes for visual and semantic information about familiar faces in a common representational space. Proc. Natl. Acad. Sci. USA. 118, e2110474118 (2021).
Chen, J., Papies, E. K. & Barsalou, L. W. A core eating network and its modulations underlie diverse eating phenomena. Brain Cogn. 110, 20–42 (2016).
Sato, W., Kochiyama, T., Minemoto, K., Sawada, R. & Fushiki, T. Amygdala activation during unconscious visual processing of food. Sci. Rep. 9, 7277 (2019).
Douglass, A. M. et al. Central amygdala circuits modulate food consumption through a positive-valence mechanism. Nat. Neurosci. 20, 1384–1394 (2017).
Mormann, F. et al. A category-specific response to animals in the right human amygdala. Nat. Neurosci. 14, 1247–1249 (2011).
Psychtoolbox-3. http://psychtoolbox.org/.
StudyForrest Project. studyforrest.org. https://studyforrest.org/.
A. Everything Everywhere All At Once | Official Trailer HD | A24. https://www.youtube.com/watch?v=wxN1T1uxQ2g (2021).
Films, S. G. Big Gold Brick - Official Trailer. https://www.youtube.com/watch?v=uDoQ2qzO8Rw (2021).
Movies, L. Warning (2021 Movie) Official Trailer - Thomas Jane, Alex Pettyfer, Alice Eve. https://www.youtube.com/watch?v=H9Cpdh0eKuc (2021).
The FFmpeg Developer Team. FFmpeg - A complete, cross-platform solution to record, convert and stream audio and video. https://ffmpeg.org/ (2023).
Beck, A. T., Steer, R. A. & Brown, G. K. BDI-II, Beck Depression Inventory: Manual. (Psychological Corporation, 1996).
Spielberger, C. D., Gorsuch, R. L., Lushene, R., Vagg, P. R. & Jacobs, G. A. STAI: Manual for the State-Trait Anxiety Inventory. (1983).
Watson, D., Clark, L. A. & Tellegen, A. Development and validation of brief measures of positive and negative affect: the PANAS scales. J. Pers. Soc. Psychol. 54, 1063–1070 (1988).
Cohen, S., Kamarck, T. & Mermelstein, R. A Global Measure of Perceived Stress. J. Health Soc. Behav. 24, 385–396 (1983).
Cohen, S. & Janicki-Deverts, D. Who’s stressed? Distributions of psychological stress in the United States in probability samples from 1983, 2006, and 20091. J. Appl. Soc. Psychol. 42, 1320–1334 (2012).
Insightface: State-of-the-Art 2D and 3D Face Analysis Project. (Github).
Avants, B. B., Duda, J. T., Zhang, H. & Gee, J. C. Multivariate normalization with symmetric diffeomorphisms for multivariate studies. Med. Image Comput. Comput. Assist. Interv. 10, 359–366 (2007).
Avants, B. B., Tustison, N. J. & Johnson, H. J. Advanced Normalization Tools (ANTs). https://github.com/ANTsX/ANTs (2014).
Fischl, B. FreeSurfer. Neuroimage 62, 774–781 (2012).
Ségonne, F., Pacheco, J. & Fischl, B. Geometrically accurate topology-correction of cortical surfaces using nonseparating loops. IEEE Trans. Med. Imaging 26, 518–529 (2007).
Gorgolewski, K. et al. Nipype: a flexible, lightweight and extensible neuroimaging data processing framework in python. Front. Neuroinform. 5, 13 (2011).
Glasser, M. F. et al. The minimal preprocessing pipelines for the Human Connectome Project. Neuroimage 80, 105–124 (2013).
Friedman, L., Glover, G. H. & Consortium, F. Reducing interscanner variability of activation in a multicenter fMRI study: controlling for signal-to-fluctuation-noise-ratio (SFNR) differences. Neuroimage 33, 471–481 (2006).
Tyszka, J. M. et al. OpenNeuro ds006947. Dense complex-valued partial brain BOLD of passive audiovisual movie presentations https://doi.org/10.18112/openneuro.ds006947.v1.0.1 (2026).
Beckmann, C. F. & Smith, S. M. Probabilistic independent component analysis for functional magnetic resonance imaging. IEEE Trans. Med. Imaging 23, 137–152 (2004).
Kliemann, D. et al. Caltech Conte Center, a multimodal data resource for exploring social cognition and decision-making. Scientific Data 9, 1–15 (2022).
Power, J. D., Barnes, K. A., Snyder, A. Z., Schlaggar, B. L. & Petersen, S. E. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142–2154 (2012).
Esteban, O. et al. MRIQC: Advancing the automatic prediction of image quality in MRI from unseen sites. PLoS One 12, e0184661 (2017).
Power, J. D. et al. Distinctions among real and apparent respiratory motions in human fMRI data. Neuroimage 201, 116041 (2019).
Gratton, C. et al. Removal of high frequency contamination from motion estimates in single-band fMRI saves data without biasing functional connectivity. Neuroimage 116866, https://doi.org/10.1016/j.neuroimage.2020.116866 (2020).
Cox, R. W., Chen, G., Glen, D. R., Reynolds, R. C. & Taylor, P. A. FMRI Clustering in AFNI: False-Positive Rates Redux. Brain Connect. 7, 152–171 (2017).
Gong, Z.-Q. & Zuo, X.-N. Dark brain energy: Toward an integrative model of spontaneous slow oscillations. Phys. Life Rev. 52, 278–297 (2025).
Syväoja, S. S. et al. Ultrafast complex-valued 4D fMRI reveals sleep-induced brain respiratory pulsation changes in both magnitude and phase signals. Neuroimage 317, 121290 (2025).
Yu, M.-C. et al. ICA of full complex-valued fMRI data using phase information of spatial maps. J. Neurosci. Methods 249, 75–91 (2015).
Qiu, Y. et al. Spatial source phase: A new feature for identifying spatial differences based on complex-valued resting-state fMRI data. Hum. Brain Mapp. 40, 2662–2676 (2019).
Chen, Z., Fu, Z. & Calhoun, V. Phase fMRI reveals more sparseness and balance of rest brain functional connectivity than magnitude fMRI. Front. Neurosci. 13, 204 (2019).
Robinson, S. D. et al. Improved dynamic distortion correction for fMRI using single-echo EPI and a readout-reversed first image (REFILL). Hum. Brain Mapp. 44, 5095–5112 (2023).
Lancione, M. et al. Complementing canonical fMRI with functional Quantitative Susceptibility Mapping (fQSM) in modern neuroimaging research. Neuroimage 244, 118574 (2021).
Chen, Z. & Calhoun, V. T2* phase imaging and processing for brain functional magnetic susceptibility (χ) mapping. Biomed. Phys. Eng. Express 2, 025015 (2016).
Wiggins, G. C. et al. 32-channel 3 Tesla receive-only phased-array head coil with soccer-ball element geometry. Magn. Reson. Med. 56, 216–223 (2006).
Boubela, R. N. et al. fMRI measurements of amygdala activation are confounded by stimulus correlated signal fluctuation in nearby veins draining distant brain regions. Sci. Rep. 5, 10499 (2015).
Tyszka, J. M. & Pauli, W. M. An in vivo atlas of the human amygdala. DOI%252010.17605/OSF.IO/HKSA6 (2017).
Zhou, Q. et al. Charting the human amygdala development across childhood and adolescence: Manual and automatic segmentation. Dev. Cogn. Neurosci. 52, 101028 (2021).
Acknowledgements
We are greatly indebted to Dr. Lynn K. Paul for all her help administering and reviewing the session questionnaires in Qualtrics. This work was funded by a 2022 Research Grant Award from the T & C Chen Center for Social and Decision Neuroscience at Caltech (JMT and RA).
Author information
Authors and Affiliations
Contributions
R.A. and J.M.T. conceived the study. J.M.T., Y.X., and R.A. developed the methodology. J.M.T., U.K., and Z.D. developed the software and performed validation. Formal analysis was conducted by J.M.T., U.K., Z.D., N.K., W.Z., Q.W., and D.A.K. J.M.T. carried out the investigation and provided resources. J.M.T. and Z.D. curated the data. J.M.T., R.A., Z.D., and U.K. wrote the original draft and contributed to review and editing. J.M.T., U.K., and Z.D. produced the visualizations. R.A. and J.M.T. supervised the project, managed project administration, and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tyszka, J.M., Diamandis, Z., Keles, U. et al. “Dense Amygdala”: Extensive Complex-valued Functional MRI of the Ventral and Medial Temporal Lobe during Passive Movie Watching in Three Individuals. Sci Data (2026). https://doi.org/10.1038/s41597-026-07065-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07065-x