Abstract
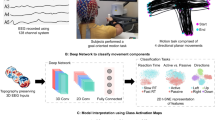
Physiological artifacts pose persistent challenges in electroencephalogram (EEG) data acquisition, often compromising interpretation and post-analysis of EEG signals across research and clinical applications. To address such limitations, including various artifact types, insufficient annotations, and low spatial resolutions, we present PhysioMotion Artifact, a large-scale, task-driven EEG dataset with point-wise artifact annotations. EEG data was acquired from 30 healthy participants performing 16 systematically designed single-type and multi-type movement tasks, inducing 14 distinct types of physiological artifacts. To demonstrate the utility of the dataset, we implemented a Convolutional Neural Networks-Transformer hybrid model for artifact detection and classification, achieving 95.4% accuracy in binary classification and 79.7% in 14-class classification tasks.
Similar content being viewed by others
Data availability
The dataset is publicly available on OpenNeuro platform (https://doi.org/10.18112/openneuro.ds006386.v1.0.1)30.
Code availability
All code utilized in this study has been made publicly available on GitHub (https://github.com/JiangweiYu221/PhysioMotion_Artifact), encompassing modules for data preprocessing, the annotation interface, the annotation verification tool, and model training procedures. Comprehensive instructions for setup and usage are provided in the accompanying README file included in the repository.
References
Praline, J. et al. Emergent EEG in clinical practice. Clin. Neurophysiol. 118, 2149–2155 (2007).
Islam, M. K., Rastegarnia, A. & Yang, Z. Methods for artifact detection and removal from scalp EEG: A review. Neurophysiol. Clin./Clin. Neurophysiol. 46, 287–305 (2016).
Rashmi, C. R. & Shantala, C. P. EEG artifacts detection and removal techniques for brain computer interface applications: a systematic review. Int. J. Adv. Technol. Eng. Explor. 9, 354 (2022).
Li, R. & Principe, J. C. Blinking artifact removal in cognitive EEG data using ICA. In 2006 International Conference of the IEEE Engineering in Medicine and Biology Society 5273–5276 (IEEE, 2006).
Bigirimana, A. D., Siddique, N. & Coyle, D. A hybrid ICA-wavelet transform for automated artefact removal in EEG-based emotion recognition. In 2016 IEEE International Conference on Systems, Man, and Cybernetics (SMC) 004429–004434 (IEEE, 2016).
Hartmann, M. M. et al. PureEEG: automatic EEG artifact removal for epilepsy monitoring. Neurophysiol. Clin./Clin. Neurophysiol. 44, 479–490 (2014).
Cassani, R. et al. The effects of automated artifact removal algorithms on electroencephalography-based Alzheimer’s disease diagnosis. Front. Aging Neurosci. 6, 55 (2014).
Kaur, C., Singh, P. & Sahni, S. EEG artifact removal system for depression using a hybrid denoising approach. Basic Clin. Neurosci. 12, 465 (2021).
Guarnieri, R. et al. Online EEG artifact removal for BCI applications by adaptive spatial filtering. J. Neural Eng. 15, 056009 (2018).
Benda, M. & Volosyak, I. Peak detection with online electroencephalography (EEG) artifact removal for brain–computer interface (BCI) purposes. Brain Sci. 9, 347 (2019).
Jafarifarmand, A. & Badamchizadeh, M. A. EEG artifacts handling in a real practical brain–computer interface controlled vehicle. IEEE Trans. Neural Syst. Rehabil. Eng. 27, 1200–1208 (2019).
Delorme, A., Sejnowski, T. & Makeig, S. Enhanced detection of artifacts in EEG data using higher-order statistics and independent component analysis. Neuroimage 34, 1443–1449 (2007).
Fitzgibbon, S. P. et al. Removal of EEG noise and artifact using blind source separation. J. Clin. Neurophysiol. 24, 232–243 (2007).
Winkler, I. et al. Robust artifactual independent component classification for BCI practitioners. J. Neural Eng. 11, 035013 (2014).
Inuso, G. et al. Wavelet-ICA methodology for efficient artifact removal from Electroencephalographic recordings. In 2007 International Joint Conference on Neural Networks 1524–1529 (IEEE, 2007).
Kumar, P. S. et al. An adaptive method to remove ocular artifacts from EEG signals using wavelet transform. J. Appl. Sci. Res. 5, 711–745 (2009).
Phadikar, S., Sinha, N. & Ghosh, R. Automatic eyeblink artifact removal from EEG signal using wavelet transform with heuristically optimized threshold. IEEE J. Biomed. Health Inform. 25, 475–484 (2020).
Mijović, B. et al. Source separation from single-channel recordings by combining empirical-mode decomposition and independent component analysis. IEEE Trans. Biomed. Eng. 57, 2188–2196 (2010).
Sweeney, K. T., McLoone, S. F. & Ward, T. E. The use of ensemble empirical mode decomposition with canonical correlation analysis as a novel artifact removal technique. IEEE Trans. Biomed. Eng. 60, 97–105 (2012).
Guarascio, M. & Puthusserypady, S. Automatic minimization of ocular artifacts from electroencephalogram: A novel approach by combining Complete EEMD with Adaptive Noise and Renyi’s Entropy. Biomed. Signal Process. Control 36, 63–75 (2017).
Pion-Tonachini, L., Kreutz-Delgado, K. & Makeig, S. ICLabel: An automated electroencephalographic independent component classifier, dataset, and website. NeuroImage 198, 181–197 (2019).
Khatwani, M. et al. Energy efficient convolutional neural networks for eeg artifact detection. In 2018 IEEE Biomedical Circuits and Systems Conference (BioCAS) 1–4 (IEEE, 2018).
Zhang, H. et al. A novel convolutional neural network model to remove muscle artifacts from EEG. In ICASSP 2021-2021 IEEE International Conference on Acoustics, Speech and Signal Processing (ICASSP) 1265–1269 (IEEE, 2021).
Qendro, L. et al. High frequency eeg artifact detection with uncertainty via early exit paradigm. Preprint at https://arxiv.org/abs/2107.10746 (2021).
Chuang, C.-H. et al. IC-U-Net: a U-Net-based denoising autoencoder using mixtures of independent components for automatic EEG artifact removal. NeuroImage 263, 119586 (2022).
Peh, W. Y., Yao, Y. & Dauwels, J. Transformer convolutional neural networks for automated artifact detection in scalp EEG. In 2022 44th Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC) 3599–3602 (IEEE, 2022).
Hamid, A. et al. The temple university artifact corpus: An annotated corpus of eeg artifacts. In 2020 IEEE Signal Processing in Medicine and Biology Symposium (SPMB) 1–4 (IEEE, 2020).
Obeid, I. & Picone, J. The temple university hospital EEG data corpus. Front. Neurosci. 10, 196 (2016).
Acharya, J. N. et al. American clinical neurophysiology society guideline 3: a proposal for standard montages to be used in clinical EEG. Neurodiagn. J. 56, 253–260 (2016).
Yu, J. & He, A. PhysioMotion_Artifact. https://doi.org/10.18112/openneuro.ds006386.v1.0.1 (2025).
Tatum, W. O., Dworetzky, B. A. & Schomer, D. L. Artifact and recording concepts in EEG. J. Clin. Neurophysiol. 28, 252–263 (2011).
Gorgolewski, K. J. et al. The brain imaging data structure, a format for organizing and describing outputs of neuroimaging experiments. Sci. Data 3, 1–9 (2016).
Gramfort, A. et al. MEG and EEG Data Analysis with MNE-Python. Front. Neurosci. 7, 1–13 (2013).
Bergstra, J. et al. Hyperopt: A Python library for model selection and hyperparameter optimization. Comput. Sci. Discov. 8, 014008 (2015).
Acknowledgements
This research was funded by the National Key Research and Development Program of China (Grant No. 2022YFC2405600), the National Natural Science Foundation of China (Grants No. T2225025, 31400842). We also gratefully acknowledge the support provided by the Big Data Computing Center of Southeast University and Visiting Scholar programme at the UGR (Spain).
Author information
Authors and Affiliations
Contributions
C.Y., G.Z., M.C., Y.Z., J.G. and Y.C. developed the conceptual framework for the study. C.Y., Y.Z. and W.X. provided the necessary resources. J.Y. conceived and conducted the experiments with the help of X.W., A.H. constructed the model and analysed the results. The whole process was supervised by C.Y., G.Z., Y.Z., J.G. and M.C. J.Y., A.H. and M.C. contributed to the writing of this manuscript. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, C., Yu, J., He, A. et al. PhysioMotion Artifact: A task-driven EEG dataset with point-wise motion artifact annotations. Sci Data (2026). https://doi.org/10.1038/s41597-026-07120-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-026-07120-7