Abstract
The extraction of brain tumor tissues in 3D Brain Magnetic Resonance Imaging (MRI) plays an important role in diagnosis before the gamma knife radiosurgery (GKRS). In this article, the post-contrast T1 whole-brain MRI images had been collected by Taipei Veterans General Hospital (TVGH) and stored in DICOM format (dated from 1999 to 2018). The proposed method starts with the active contour model to get the region of interest (ROI) automatically and enhance the image contrast. The segmentation models are trained by MRI images with tumors to avoid imbalanced data problem under model construction. In order to achieve this objective, a two-step ensemble approach is used to establish such diagnosis, first, classify whether there is any tumor in the image, and second, segment the intracranial metastatic tumors by ensemble neural networks based on 2D U-Net architecture. The ensemble for classification and segmentation simultaneously also improves segmentation accuracy. The result of classification achieves a F1-measure of \(75.64\%\), while the result of segmentation achieves an IoU of \(84.83\%\) and a DICE score of \(86.21\%\). Significantly reduce the time for manual labeling from 30 min to 18 s per patient.
Similar content being viewed by others
Introduction
Cancer is a disease that occurs when malignant cells grow in the body. These cells can form almost anywhere, including the brain, lungs, pancreas, and more. Cancerous cells cluster together to form a mass called a tumor and can spread throughout the body to other, more distant areas. In this article, we focus on the brain metastases which occur when cancer cells spread from their original site to the brain. Furthermore, an estimated 8–10% of adults with symptomatic brain metastases1, which is one of the most serious consequences, and it is essential to determine its stage. After diagnosis, it is necessary to take a treatment from the initial stage of cancer, such as gamma knife radiosurgery (GKRS) which is the major therapeutic strategy in this article.
GKRS is a type of radiation therapy used to treat tumors, vascular malformations, and other abnormalities in the brain. During GKRS, 201 cobalt 60 sources are arranged in a hemispheric shape in such a manner that all the gamma rays are focused at the center to create a cumulative radiation field2. The total energy of the rays is therefore transmitted precisely to the target with the preservation of normal brain tissue so that the lesion will degenerate2. To have this operation before, the radiologist should confirm the coordinate of the lesion on the MRI images. This treatment provides a new method for traditional neurosurgery and makes the surgical procedure be safer and faster. However, the time it takes for the radiologist to mark the location of the lesion on the MRI images depends on the shape and the volume of the tumors, and it often takes more than 30 min on average. It will even take several times longer to confirm the result in detail again.
This article developed an automated deep learning assistant diagnosis model for radiography. It can shorten the time to interpret the location of the lesion; moreover, it serves as a second suggestion for the doctors. Especially, for patients with terminal cancer, after the patient undergoes an MRI examination, the radiologist can quickly make a diagnosis report based on the preliminary data of the auxiliary diagnosis. Doctors can diagnose brain metastases, determine the stage of cancer, and develop treatment policies as soon as possible, which can greatly reduce the waiting time of patients. Relatively, doctors can also provide patients more care and design the best treatment strategies to make patients less anxious. In addition, it may speed up the hospital process, save hospital operating costs, and enhance competitiveness.
Our solution is to perform 2D U-Nets in ensemble learning for intracranial metastatic tumors classification and segmentation. The dataset contains the post-contrast \(T_{1}\)-weighted whole-brain MRI images which are an important clinical tool for cancer detection, diagnosis, prognosis, and treatment evaluation. In brief, the dataset is split as train, validation and test set first. We use train set to build a 2D U-Net model, and input validation set to this model to select an image classification threshold as the Youden index given by Receiver Operating Characteristic (ROC) curve. Furthermore, for segmentation, we calculate the detection rate of tumors in Precision-Recall (PR) curve, then set the mask threshold by the maximal F1-measure for labelling if a pixel has tumor. Finally, the test set is input to the model and we use the thresholds obtained above for classification and segmentation. In the last, the Intersection over Union (IoU) and DICE score are used as metrics for evaluation.
The contribution of this article is to design an automated 2D intracranial metastasis tumor segmentation system to overcome common problems in practice, such as hardware equipment, insufficient data and imbalanced data. Overview deep learning papers or related competitions on medical images of the brain, nothing more than using 3-dimensional models for construction or excellent hardware equipment to train a good model to refresh the ranking of the competition and publish a paper. However, in practice, there is no such abundant resource that can make AI produce the best possible results. Therefore, this article puts forward a new idea based on this position. Further design the ensemble strategy for classification and segmentation simultaneously, reducing the time for manual labeling from 30 min to 18 s per patient.
Materials and methods
Materials
The dataset of MRIs were originally acquired for guiding radiosurgical treatment using the Gamma Knife from 1999 to 2018. This dataset contained 556 patients suffered from intracranial metastases and referred for GKRS, including the precise lesion locations of total 1872 metastases lesions had been collected. This retrospective study was approved, and informed consent was waived by the Institutional Review Board of Taipei Veterans General Hospital (IRB-TPEVGH No.: 2017-10-017AC) and conformed to the tenets of the Declaration of Helsinki. All methods were carried out in accordance with relevant guidelines from IRB-TPEVGH. This article investigated tumors with a volume greater than 0.15 ml. In Fig. 1, the reason is that the size of the tumor accounts for almost half of the total number of tumors. A total of 492 patients with 904 tumors and 23354 \(T_{1}\)-weighted MRI images were found that an average of 48 images per patient, \(20\%\) of which had tumors, showed an imbalanced dataset. The image format is stored with the size of \(512\times 512\) pixels (approximately 5 mm per pixel) and the thickness of 3 mm. The original image of the whole brain in per patient will be presented in Fig. 2. In order to generate a corresponding mask for each image to use in the calculation of tumor volume, several DICOM tags need to be used.
The number and proportion of each tumor volume range.
Post-contrast T1-weighted MRI per patient.
Data pre-processing
The pre-processing step in this study involves three parts, selection for the region of interest (ROI), enhancement, and normalization. The pre-processing step firstly selects the ROI by the active contour model to overcome the imbalance pixel number for the skull and the background. Then, we enhance the contract to strengthen the image information. We also resize the image because the image size of ROI is varying. To reduce the noise in the MRI images, the snake model was utilized for delineating the skull outline, which makes use of the energy constraints and forces in the image for separation of the region of interest (ROI)3,4. The reason for this action is to overcome the imbalance pixel number for the skull and the background. Only focus on address the ROI part of image enhancement by z-score; besides, set the background to all black for decreasing the subsequent impact on convolutional neural network (CNN) training. Considering the structure of the models and the scale of the dataset, resize the image to \(256\times 256\) and normalize the data dimensions so that each image is approximately the same scale which the minimal and maximal along the dimension is \(-1\) and 15. At last, fill the background with intensity value zero again to ensure the image does not appear other values.
Methods
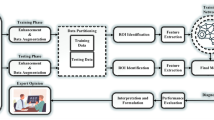
Figure 3 provides the framework of the experimental design. Input the processed images into four semantic segmentation models individually, 2D-UNet, 2D-UNet with backbone VGG-16, 2D-UNet with backbone ResNet-34, and 2D-UNet with backbone EfficientNet-B0, then get four different outputs, namely the metastasis probability maps6,7,8,9. For the evaluation, deploy the trained model to the validation set for determining the threshold of classification and segmentation. Input the test set through the function \(I_{T_{c}}(score_{i})\) which do the classification is given by Eq (1)
where
In classification, select the image threshold \(T_{c}\) by ROC curves to determine whether the image had tumors or not. If the predicted image had tumors, do the semantic segmentation by average the images \(f_{j,i}\) from four different models. Collect all the result be the \(Y_{pred}\) in this part, namely stage-one.
Next, the mask threshold level is selected where all pixel values below the threshold are mapped to zero (black) and an upper threshold value is chosen so that all pixel values above this threshold are mapped to one (white). Then the binary masks \(Y_{new\_pred}\) for an predicted image would be created. In Eq. (2), if the g(x, y) were the input image, the thresholded image h(x, y) is given by
Finally, computed the performance of the new output prediction \(Y_{new\_pred}\).
The framework of the experimental design in this article.
Experimental environment
The network has been implemented using the open source Keras framework on top of TensorFlow10. For the training process, each 2D U-Net model requires approximately 2 hours to finish on an NVIDIA GeForce GTX 1080 Ti graphics processing unit (GPU) with 11G memory and Intel®Core(TM) i7-6700 CPU @ 3.40GHz with 64GB random access memory (RAM).
Results
Evaluating the framework on the validation set with 80 patients and the test set with 100 patients both using the whole brain images. The performance of classification was evaluated by ROC curve, AUC, precision, and Recall11,12. The segmentation part was measured by PR curve, F1-measure, IoU and DICE13.
Classification
Use validation set to determine decision points and plot the ROC curve of the result in classification. As shown in Fig. 4, the AUC were 0.8514 in validation set, which around 0.8 to 0.9 is considered excellent4. According to the Youden Index, the best threshold \(T_{c}\) was 0.50. The confusion matrix of the classification in Fig. 4 indicated that the precision and Recall in Table 1.
In test set, given the \(T_{c}=0.5\), if the \(score_{i}\) calculated in stage-one was less than 0.50, then the image will be present in no tumors. Figure 4 presented the confusion matrix of the classification in test set. The Youden Index ensured the number of false negative be the minimal in this set, so the images were not missed. In addition, the precision and recall were shown in Table 1.
The result in classification. The left is the ROC curve of the classification. At \(T_{c}=0.5\), middle shows the confusion matrix from validation set, the right is from the test set.
Segmentation
Undertake the images classified as having tumors, then averaged the image segmentation results in validation set for determined the \(Mask_c\). Through different mask thresholds, the PR curve is drawn to obtain the AUC of the tumor volume bigger than 0.15 ml detected rate of the validation set was 89.90%, in which the value around 0.8–0.9 is extremely good; that is, a certain level of the tumor can be detected through this experimental. Figure 5 and Table 2 shown the performances in \(Mask_{c}=0.52\), which the maximal of F1-measure on the PR curve. Next, calculated the IoU and DICE score of each image which the performances shown in Table 2.
In test set, given the best mask threshold \(Mask_{c}=0.52\) from the validation set, which means that all intensity values below the threshold are mapped to zero (black) and an upper threshold value is chosen so that all intensity values above this threshold are mapped to one (white). So the binary masks were be the final prediction results of ensemble strategy. Then, calculate the IoU and DICE score of each image in the test set, and the measurement to get the results in Fig. 5 and Table 2.
The result in segmentation. The left is the PR curve of the detection rate of tumors. At the \(Mask_{c}=0.52\), middle shows the confusion matrix from validation set, the right is from the test set.
Visualization
Figure 6 visualize the segmentation results. The left-hand side was the original MRI image and the right-hand side shown the ground truth where the tumors was in the slice. The middle is the segmentation, the blue part was the true positive, the green region was the false negative and the area in red was false positive. More segmentation result shown in Figure 7.
Selected the tumor segmentation results compared in slice-by-slice.
Visualization the segmentation by the ensemble strategy.
Conclusion and comparison
The main contribution of this study is to design an automated 2D intracranial metastasis tumor segmentation system in practice. Significantly shorten the time for radiologists to mark lesions. Through a fully automated process, the MRI of a patient can be completed about every 18 s. In this way, doctors can also spend more time taking care of the psychological state of patients and their families; therefore, improve the comprehensive medical quality.
The technical part in this study, image pre-processing and two-stage ensemble strategy consider to overcome the imbalanced classification and the pixel-wise class imbalanced in binary image segmentation. Make good use of the characteristics of the semantic segmentation model and use different types of datasets, design a ensemble strategy for classification and segmentation simultaneously. Moreover, ensemble modelling improves segmentation accuracy. The performance of the model also exceeds the 3D U-Net.
This study proposes the integrated and automatic approach to classify and segment the intracranial metastatic tumors from MRI images. The proposed approach firstly classifies whether there is any tumor in the image. If this is positive, then it segments the intracranial metastatic tumors by ensemble learning based on the 2D U-Net architecture. For the first part of classification, recent studies14,15,16 have similar results. In14, they use the 3D architecture in CNN to classify seven classes for health and various tumor types. They also conduct the comparison for quantitative results by 2D ResNet 50. In the comparison, the sensitivity, which is equal to Recall in our paper, are 72, 72, 92, 94, 97, 83 and 57 in percentage, whereas our result is 82.33 in percentage. Note that, their sample sizes range from 45 to 359, whereas our sample size is 23,354. In15,16, they use the method of mRMR as the feature selection method and the similar architecture to classify the images. In15, they focus on the breast and use ultrasonography as input images. In16, they proposed the hybrid models with KNN to achieve great performance in F1-score and sensitivity. However, their total samples are 12,800 for four classes and they do not develop the segmentation part. Hence, the integrated and automatic approach developed in this study is more suitable for clinical practice because it provides comprehensive tools for medical experts.
References
Eichler, A. et al. The biology of brain metastasestranslation to new therapies. Nat. Rev. Clin. Oncol. 8(6), 344 (2011).
Gamma Knife Radiosurgery, Chapter: Intracranial Cavernomas and Gamma Knife Radiosurgery (SM Group eBooks DE-USA) pp. 1–18
Hemalatha, R. J. et al. Active contour based segmentation techniques for medical image analysis. Medical and biological image analysis17, (2018).
Kass, M., Witkin, A. & Terzopoulos, D. Snakes: active contour models. Int. J. Comput. Vis. 1(4), 321–331 (1988).
Ioffe, S., & Szegedy, C. ñBatch normalization: accelerating deep network training by reducing internal covariate shift. In îICML (2015).
Long, J., Shelhamer, E., & Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 3431–3440 (2015).
Ronneberger, O., Fischer, P., & Brox, T. U-net: convolutional networks for biomedical image segmentation. In International Conference on Medical Image Computing and Computer-Assisted Intervention (Springer, Cham, 2015), pp. 234–241.
Rokach, L. Ensemble-based classifiers. Artif. Intell. Rev. 33(1–2), 1–39 (2010).
Krizhevsky, A., Sutskever, I., & Hinton, G. E. Imagenet classification with deep convolutional neural networks. In: Advances in Neural Information Processing Systems (pp. 1097–1105). (2012).
Chollet, F., et al. (2015). Keras. https://github.com/fchollet/keras.
Lee, C.-T. Performance Analysis of 3-Class Classifiers Based on the Volume Under a 3D ROC Surface. https://hdl.handle.net/11296/6c2a3t
Mandrekar, J. N. Receiver operating characteristic curve in diagnostic test assessment. J. Thorac. Oncol. 5(9), 1315–1316 (2010).
Branco, P., Torgo, L. & Ribeiro, R. P. A survey of predictive modeling on imbalanced domains. ACM Comput. Surv. (CSUR) 49(2), 1–50 (2016).
Chakrabarty, S., Sotiras, A., Milchenko, M., LaMontagne, P., Hileman, M., & Marcus, D. MRI-Based Identification and Classification of Major Intracranial Tumor Types Using a 3D Convolutional Neural Network: A Retrospective Multi-institutional Analysis (RSNA) (2021).
Eroglu, Y., Yildirim, M. & Cinar, A. Convolutional neural networks based classification of breast ultrasonography images by hybrid method with respect to Benign, Malignant, and normal using mRMR. Comput. Biol. Med. 133(10016), 104407 (2021).
Eroglu, Y., Yildirim, M. & Cinar, A. mRMR-based Hybrid Convolutional Neural Network Model for Classification of Alzheimer’s Disease on Brain Magnetic Resonance Images (Wiley Online Library, 2021).
Acknowledgements
The brain tumor images used in this study were provided by TVGH. This work is supported by grants from the Ministry of Science and Technology, Taiwan (MOST 106-2634-F-075-001, MOST 107-2634-F-075-001, MOST 108-3011-F-075-001, MOST 110-2118-M-A49-002-MY3). We are grateful to the National Center for High-performance Computing for computer time and facilities.
Author information
Authors and Affiliations
Contributions
All authors contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, CC., Wu, MY., Sun, YC. et al. Ensemble classification and segmentation for intracranial metastatic tumors on MRI images based on 2D U-nets. Sci Rep 11, 20634 (2021). https://doi.org/10.1038/s41598-021-99984-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-021-99984-5