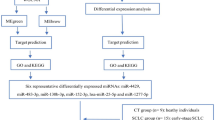
Abstract
MicroRNAs (miRNAs) are important regulators of gene expression and are involved in bacterial pathogenesis and host–pathogen interactions. In this study, we investigated the function of miRNAs in the regulation of host responses to Pasteurella multocida infection. Using next-generation sequencing, we analyzed miRNA expression pattern and identified differentially expressed miRNAs in Pasteurella multocida-infected goat lungs. In addition, we investigated the function of differentially expressed miRNAs andtheir targeted signaling pathways in bacterial infection processes. The results showed that Pasteurella multocida infection led to 69 significantly differentially expressed miRNAs, including 28 known annotated miRNAs with miR-497-3p showing the most significant difference. Gene target prediction and functional enrichment analyses showed that the target genes were mainly involved in cell proliferation, regulation of the cellular metabolic process, positive regulation of cellular process, cellular senescence, PI3K-Akt signaling pathway, FoxO signaling pathway and infection-related pathways. In conclusion, these data provide a new perspective on the roles of miRNAs in Pasteurella multocida infection.
Similar content being viewed by others
Introduction
Pasteurella multocida, is a Gram-negative anaerobic bacterium tha causes pneumonia and hemorrhagic septicemia, resulting in high morbidity and mortality as well as substantial economic losses1. It is a zoonotic pathogen that commomly found in the upperairways of animals, capable of transmitting to humans through animal bites, scratches, and contact with nasopharyngeal secretions2. Pasteurella multocida causes pasteurellosis, which is one of the most common diseases in goats3. Current vaccination strategies and antibiotics are inefficient4, highlighting the need for novel anti-bacterial strategies.
Pasteurella multocida produces multiple virulence factors that activate numerous cellular responses in the host5. Lipopolysaccharide (LPS), a crucial virulence factor, plays a significant role in the pathogenesis by interacting with the host immune system to promote infection6. In addition, protein toxins produced by Pasteurella multocida can regulate host immune system and mediate the production of pro-inflammatory cytokines7. Pasteurella multocida infection promotes inflammasome assembly and release, which activate caspase-1 and IL-1β secretion, triggering inflammation8. Pasteurella multocida mediates apoptotic and autophagic pathways causing liver injury9. Forthermore, Pasteurella multocida infection changes cell structure and regulates chromatin open regions, leading to transcriptome changes10. Undersranding exploring host factors involved in P.multocida infection can provide insights into the regulatory mechanisms of Pasteurella multocida-infected goat lungs.
MicroRNAs (miRNAs) are short (from 18 to 24 nucleotides), endogenous, highly conserved, non-coding RNA molecules11,12. MiRNAs are involved in the regulation of post-transcriptional gene expression. Theyplay crucial roles in multiple biological processes, including bacterial pathogenesis, immune response, and susceptibility13,14. For example, in caprine parainfluenza virus type 3 (CPIV3) infection,bta-miR-677 interacts with the 3'-untranslated region (3'-UTR) of mitochondrial antiviral signaling protein (MAVS), enhancing IFN pathway and suppressing viral replication in MDBK cells15. In addition, bta-miR-98 and bta-miR-222 suppress CPIV3 replication by targeting different genes16,17. Similarly, in Peste Des Petits Ruminants Virus (PPRV) infection, miR-1 is suppressed, promoting the expression of its target gene TWEAK18. Overexpression of miR-218 inhibits the expression of the SLAM gene, which is the primary receptor for PPRV and other morbilliviruses19. Research shows that miR-29-5p can repress Pasteurella multocida proliferation by directly targeting EMP2 and TBX420. High-throughput miRNA sequencing has been widely usedin identifying key miRNAs in goats diseases21,22. Differentially expressed miRNA were identified in goat peripheral blood mononuclear cells (PBMCs) during PPRV infection23,24. Similarly,miRNA sequnecing was used to detect the function of miRNAs against Brucella melitensis infection25. In this study, we used high-throughput miRNA sequencing to analyze the effects of P. multocida infection on the miRNA expression.
Materials and methods
Sample collection
7 Healthy 3-month-old males of South Sichuan black goats were kept in Zigong City, China. They were housed under the same conditions, including free access to water and food under natural lighting. They were all healthy and in good physical condition, with the same weight within the breeds, and then randomly separated into two groups: Control (n = 3) and Pasteurella multocida-treated group (n = 4). Both groups were inoculated with P.multocida strain serotype D or only equal volume of PBS at the same time. Once the infected goats showed clinical symptoms, both groups of goats were intraperitoneally euthanized by injection of sodium pentobarbital, and lung tissue samples measuring 1 cm3 were collected and stored in − 80 °C for miRNA-seq. All the experimental procedures were carried out under the authorization of the animal ethic committee of animal science academy of Sichuan province, China.
RNA extraction, and miRNA-Seq
Total RNA of goats lung from Pasteurella multocida-infected group and control group was extracted using TRIzol Reagent (Invitrogen). DNA digestion was carried out after RNA extraction by using DNaseI. RNA quality and integrity was determined by detecting A260/A280 with Nanodrop and agarose gel electrophoresis, respectively. Finally, Purified RNAs were finally quantified by Qubit3.0 with QubitTM RNA Broad Range Assay kit (Life Technologies) for miRNA library preparation. Proprietary adapters were ligated to the 5′ and 3′ terminals of the RNA, the eluted cDNA libraries were synthesized and then separated by 6% PAGE gel to sequence on Hiseq X-10 sequencer (Illumina) with PE150 model.
miRNA-Seq data analysis
After sequencing, the raw sequencing data was first filtered by discarding the low-quality reads and adapter sequences. Clean reads were mapped to the reference genome of Capra hircus (assembly ARS1) using bowtie program26 with default parameters and reads-related information was recorded. MiRDeep2 v2.0.1.3 (https://anaconda.org/bioconda/mirdeep2)27 was used to annotate known miRNAs of miRBase database, and unannotated miRNAs were recognized as novel miRNAs. In addition, the expression levels of miRNAs libraries were normalized with transcripts per million (TPM). The differential expressed miRNAs were identified using the edgeR v4.0.16 (https://bioconductor.org/packages/edgeR)28 following the screen cutoff: p-value < 0.05 and |log2Foldchange|> 1.
Functional enrichments analysis of predicted miRNA
miRwalk database (http://mirwalk.umm.uni-heidelberg.de/)29 that contains three different databases: miRDB (https://mirdb.org/), TargetScan (https://www.targetscan.org/), and miRTarBase (https://mirtarbase.cuhk.edu.cn/), was used to predict the target genes of differential expressed miRNAs. MiRNA–mRNA regulatory network was visualized with Cytoscape 3.10.2 (http//www.cytoscape.org/). The functional annotation of target genes was performed with Gene ontology (GO) and Kyoto encyclopedia of genes and genomes (KEGG30 enrichment analysis. with a corrected p-value cutoff of 0.05.
Ethics approval and consent to participate
The animal ethic committee of animal science academy of Sichuan province approved all experiments protocols for the animal trials used herein. The experiment’s procedures were performed in accordance with the guidelines approved by the institutional animal care and use committee of animal science academy of Sichuan province. All methods are reported in accordance with ARRIVE guidelines.
Results
Sequencing results of small RNA libraries
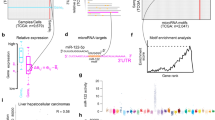
We generated approximately 29 million raw reads by performing functional enrichment analysis of target genes of differentially expressed miRNAs (Table 1). After removing low-quality and contaminated sequences, we obtained 19- to 23-nucleotide long miRNAs for further analysis (Fig. 1). The Q30 base percentage of all samples was above 95%, and the sequenceswere aligned with the goat reference genome, with a mapped rate of more than 98%.
miRNA length distribution.
Differentially expressed miRNAs after Pasteurella multocida infection
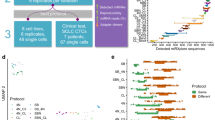
A total of 422 known miRNA were identified using miRBase database, and a total of 461 novel miRNAs were predicted using MIREAP_v0.2 software (Additional File 1). Using significant differences standard, 69 differentially expressed miRNAs were identified in P. multocida infected group compared with control group, including 42 upregulated and 27 downregulated miRNAs (Fig. 2A and B, Additional File 2).
Transcriptome analysis of miRNA in pasteurella multocida infected lung in southern sichuan black goats. (A) Volcano plots shows differentially expressed miRNAs. (B) Differentially expressed miRNAs clustering map.
Prediction and functional characterization of target genes of differentially expressed miRNAs
To understand the potential functions of these differentially expressed miRNAs, the target genes were predicted through using the miRwalk database. The analysis identified 28 miRNAs (let-7a-3p, miR-106b-3p, miR-130a-3p, miR-130a-5p, miR-144-3p, miR-144-5p, miR-146b-3p, miR-146b-5p, miR-155-5p, miR-15b-5p, miR-186-3p, miR-192-3p, miR-199b-5p, miR-223-5p, miR-27b-5p, miR-29c-3p, miR-330-5p, miR-335-5p, miR-363-3p, miR-374a-3p, miR-374b-3p, miR-409-5p, miR-423-3p, miR-493-3p, miR-493-5p, miR-497-3p, miR-497-5p and miR-92a-3p) and predicted 1229 target genes. Of these, 310 target genes had been validated (Fig. 3). A further functional enrichment analysis (GO and KEGG pathway enrichment) (Fig. 4) showed that the target genes were mainly enriched in epithelial cell proliferation, the Wnt signaling pathway and protein regulation. In terms of molecular function, the top categories were DNA-binding transcription activator activity, protein serine/threonine kinase activity and phosphatase binding. The analysis identified 77 items in the cellular component group, with the most significant being glutamatergic synapse and cytoplasmic ribonucleoprotein granule. The KEGG pathway analysis showed enrichments in the PI3K-Akt signaling pathway, focal adhesion, neurotrophic signaling pathway, and FoxO signaling pathway, suggesting that the dysregulated miRNAs may regulate genes involved in immune resposne, host–pathogen interaction, and inflammation, likely contributing to the development of pasteurellosis caused by Pasteurella multocida infection.
Construction and visualization of mRNA–miRNA networks.
Functional enrichment analysis of target genes of differentially expressed miRNAs. (A) The enriched BP distribution. (B) The enriched MF distribution. (C) Significantly enriched terms based on CC. (D) Enriched KEGG pathways of differentially expressed miRNA target genes.
Discussion
The Gram-negative Pasteurella multocida is one of the main pathogens in goats, causing pasteurellosis and hemorrhagic septicemia. The effectiveness of current antibiotics and vaccines is limited due to antimicrobial resistance and the specificity of vaccine targets, making it crucial to understand the molecular mechanisms of host–pathogen interactions. We used a high-throughput sequencing approach to identify differentially expressed miRNAs in response to P. multocida infection.
In this study, 884 miRNAs were detected, comprising 422 known and 462 newly predicted miRNAs, and 69 differentially expressed miRNAs were identified in Pasteurella multocida-infected group compared with the control group, including 42 upregulated and 27 downregulated miRNAs. We found that miR-497-3p was the most significantly upregulated miRNAs in Pasteurella multocida-infected group. Previous studies have shown that miR-497-3p was up-regulated in patients with bacterial pneumonia and SpT4-infected mice, and inhibition of miR-497-3p suppressed the level of inflammatory cytokines (IL-6 and TNF-α), attenuating inflammatory responses in bacterial pneumonia31. In addition, miR-497-3p causes DNA damage and apoptosis by inactivating the PI3K/AKT/mTOR signaling pathway32.
Target gene prediction analysis showed that miR-497-3p may regulate SNX2, FOXK1 and CAPRIN1 expression. Previous studies have shown that SNX2 knockdown increased viral transcripts and virus growth, and it could interact with influenza virus PA protein33. FOXK1 is a transcription factor with antiviral activity against RNA viruses34. CAPRIN1 is a stress granule-associated RNA-binding protein taht regulates IFN-γ-mediated control of murine norovirus (MNV) replication35. In CAPRIN1-depleted cells, IFN-inducible IFITM2, RIG-I/DDX58 ISG15 and STAT1were significantly reduced36. Based on these studies, we hypothesized that miR-497-3p is involved in the apoptosis and inflammation in Pasteurella multocida infection. In addition, we found that Pasteurella multocida infection induced miR-144-3p expression. miR-144-3p could enhance LPS-induced septic acute lung injury and promote Mycobacterium tuberculosis and Mycobacterium abscessus infection by targeting different genes and pathways37,38,39. The most downregulated known miRNAs in our study were miR-374a-3p and miR-130a-3p. miR-374a-3p regulates the expression of KLF14, RTN3, WNT3, S100A10, TLR4 and ROCK1 proteins involved in multiple biological processes, including immune response40,41,42,43,44. Up-regulating miR-374a-3p expression inhibited WNT5B and JNK/ERK/MAPK pathway to attenuate LPS-induced damage and inflammation45. The expression of miR-130a-3p was significantly decreased in our study. Schistosoma japonicum infection also significantly decreased the expression of miR-130a-3p, and miR-130a-3p can inhibit MAPK1, TGFBR1 and TGFBR2 expressions to attenuate the progression of liver fibrosis caused by Schistosoma japonicum infection46. Additionally, miR-130a-3p reduces inflammation and improves pulmonary lesions by targeting TNF-α and TGF-βRII and inhibiting the secretion of inflammatory cytokines47.
Using the functional enrichment analysis, we found that the target genes of differentially expressed miRNAs were mainly enriched in epithelial cell proliferation, the PI3K-Akt signaling pathway, the neurotrophin signaling pathway and the FoxO signaling pathway. Epithelial cells are the first barrier against pathogen invasion and play critical roles in host–pathogen interactions and immune response48. Research has shown that mycobacterium tuberculosis (Mtb) virulence factor Mce2E promotes epithelial cell proliferation for its survival49. Pasteurella multocida infection could induce epithelial cells polarization and increase epithelial permeability50,51, Moreover, Pasteurella multocida infection changed the structure of goat bronchial epithelial cells and increased chromatin open region52, suggesting that maintaining epithelial cell function is important against Pasteurella multocida infection. Additionally, neurotrophins expressed in the lung play important roles in lung development, inflammation, and lung fibrosis53,54. The PI3K-Akt and FoxO signaling pathways are widely involved in the pathogenesis and pathogen–host interactions in innate immune system. Helicobacter pylori virulence factor CagA interacts with PI3Kincreasing the risk of gastric cancer55, and Streptococcus pyogenes regulates PI3K-Akt for its adhesion and invasion56. FoxO is an important transcription factor that regulates the and innate immune system in respiratory epithelial cells57. Activated FoxO promots the expression of antimicrobial peptides and the phagocytosis of hemocytes against bacterial infection58. These proteins and signaling pathways may play important roles in Pasteurella multocida infection. Additional studies are required to investigate their functions.
Conclusions
Our result identified 69 differential expressed miRNAs, and miR-497-3p was the most significantly significantly upregulated miRNAs in Pasteurella multocida-infected group in this study. The targets genes of the differentially expressed miRNAs were principally enriched in signaling pathways related to regulation of cellular metabolic process, PI3K-Akt signaling pathway, FoxO signaling pathway and infection-related pathway. 28 keys miRNAs were screened by building miRNA–mRNA interaction networks, and these miRNA scould play critical roles in response to Pasteurella multocida infection and be valuable targets for antibacterial treatment. A combination of these roles may benefit the early diagnosis and treatment pasteurellosis caused by Pasteurella multocida.
Data availability
The analyzed contributions presented in the study are included in the article/Supplementary Materials, further inquiries can be directed to the corresponding author. The datasets generated and/or analysed during the current study are available in the figshare repository, https://figshare.com/s/e5938072fcd7d2eadfa2.
References
Peng, Z. et al. Pasteurella multocida: Genotypes and genomics. Microbiol. Mol. Biol. Rev. 83(4), e00014-19. https://doi.org/10.1128/MMBR.00014-19 (2019).
Marcin, P., Beata, B. L. & Jarosław, W. Pasteurella Multocida infection in humans. Pathogens 12(10), 1210. https://doi.org/10.3390/pathogens12101210 (2023).
Mostaan, S. et al. Pasteurella multocida vaccine candidates: A systematic review. Avicenna J. Med. Biotechnol. 12(3), 140–147 (2020).
Tabatabaei, M. & Abdolahi, F. Molecular evaluation of sheep and goats isolates of Pasteurella multocida and their antibiotic resistance. Vet. Res. Forum. 14(9), 481–487. https://doi.org/10.30466/vrf.2022.556438.3524 (2023).
Kubatzky, K. F. Pasteurella multocida toxin—Lessons learned from a mitogenic toxin. Front. Immunol. 13, 1058905. https://doi.org/10.3389/fimmu.2022.1058905 (2022).
Patiño, P., Gallego, C., Martínez, N., Rey, A. & Iregui, C. Intranasal instillation of Pasteurella multocida lipopolysaccharide in rabbits causes interstitial lung damage. Res. Vet. Sci. 152, 115–126. https://doi.org/10.1016/j.rvsc.2022.07.026 (2022).
Xiao, H. et al. IFN-γ promotes PANoptosis in Pasteurella multocida toxin-induced pneumonia in mice. Vet. Microbiol. 285, 109848. https://doi.org/10.1016/j.vetmic.2023.109848 (2023).
Fang, R. et al. High- and low-virulent bovine Pasteurella multocida induced differential NLRP3 inflammasome activation and subsequent IL-1β secretion. Vet. Microbiol. 243, 108646. https://doi.org/10.1016/j.vetmic.2020.108646 (2020).
Cai, Q. et al. Pasteurella multocida causes liver injury in ducks by mediating inflammatory, apoptotic and autophagic pathways. Microb. Pathog. 184, 106336 (2023).
Chen, Q. et al. Profiling chromatin accessibility responses in goat bronchial epithelial cells infected with Pasteurella multocida. Int. J. Mol. Sci. 24(2), 1312 (2023).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 11(9), 597–610. https://doi.org/10.1038/nrg2843 (2010).
Dysin, A. P., Barkova, O. Y. & Pozovnikova, M. V. The role of microRNAs in the mammary gland development, health, and function of cattle, goats, and sheep. Noncoding RNA 7(4), 78. https://doi.org/10.3390/ncrna7040078 (2021).
Riahi Rad, Z. et al. MicroRNAs in the interaction between host-bacterial pathogens: A new perspective. J. Cell. Physiol. 236(9), 6249–6270. https://doi.org/10.1002/jcp.30333 (2021).
Zhang, F., Zhou, Y. & Ding, J. The current landscape of microRNAs (miRNAs) in bacterial pneumonia: Opportunities and challenges. Cell. Mol. Biol. Lett. 27(1), 70. https://doi.org/10.1186/s11658-022-00368-y (2022).
Zhong, C. et al. Bta-miR-677 contribute to interferon pathway affecting the proliferation of caprine parainfluenza virus type 3. Microb. Pathog. 169, 105642. https://doi.org/10.1016/j.micpath.2022.105642 (2022).
Li, J. et al. Bta-miR-98 suppresses replication of caprine parainfluenza virus type 3 through inhibiting apoptosis by targeting caspase-3. Front. Immunol. 11, 1575. https://doi.org/10.3389/fimmu.2020.01575 (2020).
Li, J. et al. Cellular microRNA bta-miR-222 suppresses caprine parainfluenza virus type 3 replication via downregulation of interferon regulatory factor 2. Vet. Microbiol. 224, 58–65. https://doi.org/10.1016/j.vetmic.2018.08.028 (2018).
Qi, X. et al. MicroRNA-1 negatively regulates peripheral NK cell function via tumor necrosis factor-like weak inducer of apoptosis (TWEAK) signaling pathways during PPRV infection. Front. Immunol. 10, 3066. https://doi.org/10.3389/fimmu.2019.03066 (2020).
Qi, X. et al. MicroRNA-218 regulates signaling lymphocyte activation molecular (SLAM) mediated peste des petits ruminants virus infectivity in goat peripheral blood mononuclear cells. Front. Immunol. 10, 2201. https://doi.org/10.3389/fimmu.2019.02201 (2019).
Hu, J. et al. Characterization of microRNA Profiles in Pasteurella multocida-Infected Rabbits and Identification of miR-29-5p as a Regulator of Antibacterial Immune Response. Front. Vet. Sci. 8, 746638. https://doi.org/10.3389/fvets.2021.746638 (2021).
Wang, B. et al. Identification of novel and differentially expressed MicroRNAs in goat enzootic nasal adenocarcinoma. BMC Genom. 17(1), 896. https://doi.org/10.1186/s12864-016-3238-5 (2016).
Wang, A. et al. Differentially expressed MiRNAs of goat submandibular glands among three developmental stages are involved in immune functions. Front. Genet. 12, 678194. https://doi.org/10.3389/fgene.2021.678194 (2021).
Pandey, A. et al. Modulation of host miRNAs transcriptome in lung and spleen of peste des petits ruminants virus infected sheep and goats. Front. Microbiol. 8, 1146. https://doi.org/10.3389/fmicb.2017.01146 (2017).
Qi, X. et al. MicroRNA expression profiling of goat peripheral blood mononuclear cells in response to peste des petits ruminants virus infection. Vet. Res. 49(1), 62. https://doi.org/10.1186/s13567-018-0565-3 (2018).
Li, B. et al. Integrated mRNA-seq and miRNA-seq analysis of goat fibroblasts response to Brucella Melitensis strain M5–90. PeerJ 9, e11679. https://doi.org/10.7717/peerj.11679 (2021).
Tam, S., Tsao, M. S. & McPherson, J. D. Optimization of miRNA-seq data preprocessing. Brief Bioinform. 16(6), 950–963. https://doi.org/10.1093/bib/bbv019 (2015).
Friedländer, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40(1), 37–52. https://doi.org/10.1093/nar/gkr688 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1), 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Sticht, C., De La Torre, C., Parveen, A. & Gretz, N. miRWalk: An online resource for prediction of microRNA binding sites. PLoS One 13(10), e0206239. https://doi.org/10.1371/journal.pone.0206239 (2018).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28(1), 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Wang, W. et al. Inhibition of miR-497-3p downregulates the expression of procalcitonin and ameliorates bacterial pneumonia in mice. Inflammation 43(6), 2119–2127. https://doi.org/10.1007/s10753-020-01279-w (2020).
Zhou, Y. et al. MiR-497-3p induces Premature ovarian failure by targeting KLF4 to inactivate Klotho/PI3K/AKT/mTOR signaling pathway. Cytokine 170, 156294. https://doi.org/10.1016/j.cyto.2023.156294 (2023).
Koçmar, T., Çağlayan, E., Rayaman, E., Nagata, K. & Turan, K. Human sorting nexin 2 protein interacts with Influenza A virus PA protein and has a negative regulatory effect on the virus replication. Mol. Biol. Rep. 49(1), 497–510. https://doi.org/10.1007/s11033-021-06906-9 (2022).
Panda, D. et al. The transcription factor FoxK participates with Nup98 to regulate antiviral gene expression. mBio 6(2), e02509-e2514. https://doi.org/10.1128/mBio.02509-14 (2015).
Kurhade, C., Kang, S., Biering, S. B., Hwang, S. & Randall, G. CAPRIN1 is required for control of viral replication complexes by interferon gamma. mBio 14(3), e0017223. https://doi.org/10.1128/mbio.00172-23 (2023).
Bidet, K., Dadlani, D. & Garcia-Blanco, M. A. G3BP1, G3BP2 and CAPRIN1 are required for translation of interferon stimulated mRNAs and are targeted by a dengue virus non-coding RNA. PLoS Pathog. 10(7), e1004242. https://doi.org/10.1371/journal.ppat.1004242 (2014).
Xu, R., Shao, Z. & Cao, Q. MicroRNA-144–3p enhances LPS induced septic acute lung injury in mice through downregulating Caveolin-2. Immunol. Lett. 231, 18–25. https://doi.org/10.1016/j.imlet.2020.12.015 (2021).
Guo, L. et al. MicroRNA-144–3p inhibits autophagy activation and enhances Bacillus Calmette-Guérin infection by targeting ATG4a in RAW264.7 macrophage cells. PLoS One 12(6), e0179772. https://doi.org/10.1371/journal.pone.0179772 (2017).
Kim, H. J. et al. MiR-144-3p is associated with pathological inflammation in patients infected with Mycobacteroides abscessus. Exp. Mol. Med. 53(1), 136–149. https://doi.org/10.1038/s12276-020-00552-0 (2021).
Li, Z., Yao, H., Wang, S., Li, G. & Gu, X. CircTADA2A suppresses the progression of colorectal cancer via miR-374a-3p/KLF14 axis. J. Exp. Clin. Cancer Res. 39(1), 160. https://doi.org/10.1186/s13046-020-01642-7 (2020).
Fu, Q. et al. LINC02288 promotes chondrocyte apoptosis and inflammation through miR-374a-3p targeting RTN3. J. Gene Med. 23(5), e3314. https://doi.org/10.1002/jgm.3314 (2021).
Zhuang, Y. et al. Yiqi Jianpi Huayu Jiedu Decoction Inhibits Metastasis of Colon Adenocarcinoma by Reversing Hsa-miR-374a-3p/Wnt3/β-Catenin-Mediated Epithelial-Mesenchymal Transition and Cellular Plasticity. Front. Oncol. 12, 904911. https://doi.org/10.3389/fonc.2022.904911 (2022).
Zhu, J., Huang, Y., Zhang, Y., Huang, R. & Huang, C. KCNMB2-AS1 promotes bladder cancer progression through sponging miR-374a-3p to upregulate S100A10. Front. Genet. 12, 655569. https://doi.org/10.3389/fgene.2021.655569 (2021).
Wang, M., Wei, J., Shang, F., Zang, K. & Zhang, P. Down-regulation of lncRNA SNHG5 relieves sepsis-induced acute kidney injury by regulating the miR-374a-3p/TLR4/NF-κB pathway. J. Biochem. 169(5), 575–583. https://doi.org/10.1093/jb/mvab008 (2021).
Shi, F. L. & Ren, L. X. Up-regulated miR-374a-3p relieves lipopolysaccharides induced injury in CHON-001 cells via regulating Wingless-type MMTV integration site family member 5B. Mol. Cell. Probes 51, 101541. https://doi.org/10.1016/j.mcp.2020.101541 (2020).
Liu, L. et al. MiR-130a-3p alleviates liver fibrosis by suppressing HSCs activation and skewing macrophage to Ly6Clo phenotype. Front. Immunol. 12, 696069. https://doi.org/10.3389/fimmu.2021.696069 (2021).
Ding, Y. et al. MiR-130a-3p alleviates inflammatory and fibrotic phases of pulmonary fibrosis through proinflammatory factor TNF-α and profibrogenic receptor TGF-βRII. Front. Pharmacol. 13, 863646. https://doi.org/10.3389/fphar.2022.863646 (2022).
Hiemstra, P. S., McCray, P. B. Jr. & Bals, R. The innate immune function of airway epithelial cells in inflammatory lung disease. Eur. Respir. J. 45, 1150–1162. https://doi.org/10.1183/09031936.00141514 (2015).
Qiang, L. et al. Mycobacterium tuberculosis Mce2E suppresses the macrophage innate immune response and promotes epithelial cell proliferation. Cell. Mol. Immunol. 16(4), 380–391. https://doi.org/10.1038/s41423-018-0016-0 (2019).
Rabier, M. J. et al. Pasteurella multocida enters polarized epithelial cells by interacting with host F-actin. Vet. Microbiol. 54(3–4), 343–355. https://doi.org/10.1016/s0378-1135(96)01255-2 (1997).
Lin, L. et al. vascular endothelial growth factor A contributes to increased mammalian respiratory epithelial permeability induced by Pasteurella multocida infection. Microbiol. Spectr. 11(2), e0455422. https://doi.org/10.1128/spectrum.04554-22 (2023).
Chen, Q. et al. Profiling chromatin accessibility responses in goat bronchial epithelial cells infected with Pasteurella multocida. Int. J. Mol. Sci. 24(2), 1312. https://doi.org/10.3390/ijms24021312 (2023).
Braun, A. et al. Brain-derived neurotrophic factor (BDNF) contributes to neuronal dysfunction in a model of allergic airway inflammation. Br. J. Pharmacol. 141, 431–440. https://doi.org/10.1038/sj.bjp.0705638 (2004).
Prakash, Y. S., Thompson, M. A. & Pabelick, C. M. Brain-derived neurotrophic factor in TNF-α modulation of Ca2+ in human airway smooth muscle. Am. J. Respir. Cell Mol. Biol. 41, 603–611. https://doi.org/10.1165/rcmb.2008-0151OC (2009).
Yong, X. et al. Helicobacter pylori virulence factor CagA promotes tumorigenesis of gastric cancer via multiple signaling pathways. Cell Commun. Signal. 2015(13), 30. https://doi.org/10.1186/s12964-015-0111-0 (2015).
Kurosawa, M. et al. Streptococcus pyogenes CAMP factor promotes bacterial adhesion and invasion in pharyngeal epithelial cells without serum via PI3K/Akt signaling pathway. Microbes Infect. 20(1), 9–18. https://doi.org/10.1016/j.micinf.2017.09.007 (2018).
Seiler, F. et al. FOXO transcription factors regulate innate immune mechanisms in respiratory epithelial cells. J. Immunol. 190(4), 1603–1613. https://doi.org/10.4049/jimmunol.1200596 (2013).
Li, C., Hong, P. P., Yang, M. C., Zhao, X. F. & Wang, J. X. FOXO regulates the expression of antimicrobial peptides and promotes phagocytosis of hemocytes in shrimp antibacterial immunity. PLoS Pathog. 17(4), e1009479. https://doi.org/10.1371/journal.ppat.1009479 (2021).
Funding
This study was supported by funds from Basic scientific research business expenses of provincial scientific research institutes in Sichuan Province (SASA202206) and Wuhan polytechnic university (2022020801020397).
Author information
Authors and Affiliations
Contributions
Methodology: Hao Zheng; validation: Hao Zheng and Feng Xu; investigation: Xia Dong; data curation: Xia Dong and Ao Zhou; writing—original draft preparation: Ao Zhou and Quzhe Emu; writing—review and editing: Ao Zhou and Quzhe Emu; visualization: Ao Zhou; supervision: Quzhe Emu; funding acquisition: Quzhe Emu. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, F., Zheng, H., Dong, X. et al. miRNA expression signatures induced by pasteurella multocida infection in goats lung. Sci Rep 14, 19626 (2024). https://doi.org/10.1038/s41598-024-69654-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-69654-3