Abstract
In the last few decades, the field of ancient DNA has taken a new direction towards using sedimentary ancient DNA (sedaDNA) for studying human and mammalian population dynamics as well as past ecosystems. However, the screening of numerous sediment samples from archaeological sites remains a time-consuming and costly endeavor, particularly when targeting hominin DNA. Here, we present a novel high-throughput method that facilitates the fast and efficient analysis of sediment samples by applying a pooled testing approach. This method combines multiple extracts, enabling early parallelization of laboratory procedures and effective aDNA screening. Pooled samples with detectable aDNA signals undergo detailed analysis, while empty pools are discarded. We have successfully applied our method to multiple sediment samples from Middle and Upper Paleolithic sites in Europe, Asia, and Africa. Notably, our results reveal that an aDNA signal remains discernible even when pooled with four negative samples. We also demonstrate that the DNA yield of double-stranded libraries increases significantly when reducing the extract input, potentially mitigating the effects of inhibition. By embracing this innovative approach, researchers can analyze large numbers of sediment samples for aDNA preservation, achieving significant cost reductions of up to 70% and reducing hands-on laboratory time to one-fifth.
Similar content being viewed by others
Introduction
Genomic material can be obtained from various environmental sources, including soil1,2, water3,4, or air5,6. Hence, environmental DNA (eDNA) is a valuable tool for ecological surveys of species that are challenging to observe directly or in the absence of their remains. The initial applications of eDNA focused on retrieving microbial genetic material from environmental samples rather than culturing them prior to extraction7,8,9. Since then, numerous studies have been published, focusing on microbial taxa recovered from sediments and permafrost10,11,12,13,14. These studies have exemplified that the retrieval of DNA from archaeological sediments and permafrost could be used to study our past15,16. In its early stages, the analysis of ancient sediments focused on simple presence/absence studies through metabarcoding quantification17,18 involving the use of taxon-specific primers to amplify short gene regions, followed by high throughput sequencing (HTS)19. This approach has demonstrated great potential for the reconstruction of past ecosystems and the impacts of climate change20 from various sources, including sediments21,22, lake sediments20,23,24,25,26, and frozen soils27, even in the absence of macrofossils. Metabarcoding offers a cost-effective option. However, it does not allow further analyses such as phylogenies or population genetic tests. Alternatively, shotgun sequencing of sedaDNA provides a greater taxonomic resolution, as it analyzes all the genetic material contained within a sample, potentially allowing for the retrieval of full genomes28,29,30. As shotgun sequencing is usually ineffective in terms of endogenous DNA sequencing, studies have applied targeted hybridization capture, particularly focusing on mammalian and hominin mitochondrial DNA31,32,33 and nuclear DNA in archaeological sediments34. These capture approaches, however, add significantly to the expense and time needed, particularly with rare hominin sedaDNA from archaeological sediments.
Even with the addition of targeted hybridization capture, sediment samples may not have any aDNA preservation. The preservation of sedaDNA in sediment is variable and depends on climatic conditions and the chemical composition of the sediment31,33,35,36,37. In Denisova Cave, for example, the majority of samples exhibited the presence of faunal and hominin mitochondrial (mt)DNA33, while, in contrast, in Sefunim Cave, only four out of 33 tested samples showed traces of mammalian mtDNA38. In their analysis of Pleistocene sediments from sites across Europe, Asia, Africa, and North America, Massilani et al. were able to extract ancient mammalian mtDNA from approximately 50% of the analyzed, resin-impregnated sediment blocks39. Their study highlights the potential of resin-impregnated sediments to preserve ancient DNA, even from small particles like bone fragments and coprolites. This finding is crucial as it demonstrates that ancient DNA can remain stably localized within sediments over time, allowing for the recovery of genetic material in a microstratigraphic context. These studies provide valuable insights into the variable preservation and spatial distribution of ancient DNA within archaeological sites, emphasizing the need for efficient screening protocols.
The potentially low success rate of mammalian sedaDNA calls for a methodological screening optimization to allow for a faster and more cost-effective assessment of samples. Sample pooling is a common practice in the field of eDNA, primarily used to assess the biodiversity of ecological communities. The concept of pooling multiple samples for collective analysis was established in the 1940s when it was initially employed to detect positive cases of Syphilis during World War II40. Over the years, this approach has found applications in various fields, including the screening of infectious diseases such as hepatitis viruses B and C41,42 human immunodeficiency virus (HIV)43, and, more recently, during the SARS-CoV-2 pandemic44,45. In the context of sedimentary eDNA screening, several pooling strategies have been proposed to reduce costs, including pre-extraction46,47,48, post-extraction49, and post-PCR pooling20,49. A recent study demonstrated the successful pooling of multiple sample libraries into a single capture reaction while optimizing reagents costs50. However, there is currently no experimental framework examining how post-extraction sample pooling of ancient sediments affects the behavior of the deamination signal and the abundance of authentic ancient DNA fragments.
In this study, we hypothesize that deamination signals can be maintained and detected with increased post-extraction pooling of up to five sediment samples. We further hypothesize that the abundance of unique DNA reads, mapping to our reference panel, will not significantly decrease in the various extract pools compared to the individual stand-alone sample. To test these hypotheses, we applied multiple degrees of extract-pooling to Paleolithic sediment samples from European, Asian, and African sites and one burial site. Our findings demonstrate that up to five extracts can be pooled while maintaining a detectable signal of aDNA and achieving an average 1.36-fold increased mammalian target DNA yield across all pooling levels compared to the average unpooled state. This high-throughput pooling method will allow for large-scale sediment analyses for aDNA preservation with great reductions in costs and time.
Materials and methods
In the following wet lab experiments, we used sediment samples obtained from multiple Paleolithic cave sites, an open air site, and one burial site: El Mirón (Spain), Velika Pećina—Kličevica (Croatia), Hovk-1 (Armenia), Krems-Wachtberg (Austria), and Shinfa-Metema 1 (Ethiopia) (Supplementary Information S1).
In silico simulation of sample pooling
Following previous evidence for detectable sedaDNA after pooling post-PCR extracts20, we first conducted an in-silico simulation to examine the impact of pooling on the characteristics of ancient mammalian mtDNA sequences using mitochondrial reference genomes of Homo sapiens and domestic goat (Capra hircus). To simulate the characteristics of ancient DNA we employed an online sequence manipulation tool (https://www.bioinformatics.org/sms2/split_fasta.html), fragmenting the original sequences into segments of 30, 35, and 40 base pair length. These fragmented sequences were then combined into composite files for each species, containing fragments of varying lengths. Next, we introduced deamination on the composite sequences with deamination rates set at 10, 30, and 50 percent per species. To model differential deamination levels between species within a single sediment sample, we combined human sequences with a 10 percent deamination rate with Capra sequences exhibiting deamination rates of 30 and 50 percent, respectively.
We carried out sequential pooling of the deaminated composite sequence files for human, Capra, and the combination of human and Capra. These pooling steps were simulated by adding random reads extracted from a sediment sample that showed no detectable aDNA reads after the initial screening process. The pooling was obtained up to a dilution of 1:10 (Supplementary Fig. S1). We assessed deamination and read count, which are the main measures of aDNA, at each pooling step using a custom bioinformatics pipeline detailed in the methods.
DNA extraction and library preparation
To mitigate the risk of modern contamination, we implemented strict measures such as the use of gloves, disposable lab suits, and other sterile equipment. All of the samples were prepared in dedicated clean room facilities at the University of Vienna. We included negative controls at each step (extractions, libraries, and PCR) to monitor potential reagent contamination. During the screening for candidate samples, amounts of 50–60 mg of sediment were used for DNA extraction following the protocol from Dabney51 with adaptations from Korlević52, and eluted in 50 µL TET buffer (10 mM Tris–HCl, 1 mM EDTA, 0.05% Tween 20, pH 8.0). Double-stranded libraries were prepared of half of the extract as described in53, omitting the shearing of DNA into smaller fragments. Instead of SPRI beads, a MinElute PCR Purification kit from Qiagen was used for cleaning up the samples and eluted in 40 µL EBT buffer (1 mM EDTA, 0.05% Tween-20). We added a positive control using 24 µL of deionized water and 1 µL of a 1:250 dilution of CL104.
Mammalian mtDNA hybridization capture and sequencing
The number of PCR cycles for optimal amplification was determined individually for each library using a qPCR machine. We double-indexed54 and amplified 25% of each library with PfuTurbo Cx HotStart DNA Polymerase. Amplification products were cleaned up using NGS clean-up magnetic beads, introducing a size selection through a concentration of 1.2 × beads per sample and eluted in a total amount of 25 µL EBT buffer (1 mM EDTA, 0.05% Tween-20). In preparation for the subsequent capture, we further amplified 3 × 2 µL of each indexed library using KAPA HiFi HotStart DNA Polymerase and the primer pairs IS5/IS653 at a rate of 20 cycles and cleaned up with magnetic beads. The concentration of the PCR product was measured on a Qubit.
We enriched for 51 mammalian mitogenomes, including human mitochondrial DNA, using custom-designed capture probes55 following the TWIST capture protocol56. After a hybridization time of 16 h at 65 °C and four rounds of washing the enriched DNA was cooked off the probes in a PCR cycler at 95 °C for 5 min (including a heated lid at 110 °C). A qPCR was performed to determine the correct number of PCR cycles for each library. Half of the captured library was re-amplified using KAPA HiFi HotStart DNA Polymerase and the primer pairs IS5/IS6 and cleaned up with magnetic beads. Library concentration (ng/µL) and molarity (nmol/L) were determined through Qubit and Tapestation. For sequencing on an Illumina NovaSeq6000XP SP lane with a total number of 300 million (3,000,000,000) reads, libraries were pooled by including 20 ng of DNA for each (which roughly equals five million reads per sample) and the amount for blanks was calculated to approximate 200,000 target reads per blank (for numbers of sequenced reads per sample refer to Supplementary Table S1). The sequencing was performed at the Vienna BioCenter Core Facilities in Vienna.
Pooling experiment
To generate sufficient extracts for the pooling experiment, we employed a modified extraction protocol that involved processing 1 g of sediment for the negative samples (i.e., samples previously sequenced but without detectable aDNA, defined as < 1000 unique reads mapping to the mammalian reference panel and < 10% of deamination) and 250 mg of test samples (i.e., samples previously sequenced with detectable aDNA, defined as either > 1000 unique reads mapping to the mammalian reference panel or > 10% of deamination, or both). Following the protocol described in Dabney et al.51 and Korlević et al.52, we adjusted the amounts of buffers to the amount of used sediment and eluted the extracts in TET buffer, using 1 mL and 250 µL for negative and test samples, respectively. To ensure reproducibility of the initial sequencing results across all samples (Supplementary Table S1), we conducted downstream lab analyses using the same methods as previously described, and again carried out shallow sequencing on a NovaSeq SP lane, alongside other samples. Any sample for which the screening results were reproducible was included in the experimental set-up. In cases where the screening results could not be replicated, samples were substituted with new candidate samples. The extracts of the previously sequenced sediment samples which showed signs of aDNA (Table 1) were diluted with the extract of a negative sample (< 1000 unique reads mapping to the mammalian reference panel and < 10% of deamination) in increasing amounts (ranging from 1:1 to 1:4) up to a total volume of 25 µL. Each pool of extracts was prepared in three replicates and converted into double-stranded libraries according to53. The library preparation process for pooling levels incorporated different test extract input volumes, combined with increasing volumes of a negative sample up to a total volume of 50 μL; That equals to 25 μL of test extract for the unpooled state, 12.5 μL for a 1:1 pooling, 8.33 μL for a 1:2 pooling, 6.25 μL for a 1:3 pooling, and 5 μL for a 1:4 pooling state. We proceeded with the downstream analyses (PCR, mammalian capture, and quality controls) as described above.
Bioinformatic processing
First, we removed the adapter sequences and sequences shorter than 30 bp using Cutadapt 4.057. The clipped reads were mapped to a composite reference file encompassing the 51 mammalian mt genomes included in the hybridization capture55 using bwa aln version 0.7.17-r118858 with an edit distance of 0.01, a gap penalty opening of 2 and seeding disabled. Reads were filtered using samtools59 retaining the ones with average mapping qualities above 30. We removed duplicate sequences with samtools rmdup and assessed damage levels through mapdamage 2-2.2.160. The quantity of the accepted reads was examined using Qualimap v.2.361.
Statistical analyses
All statistical analyses and plots were performed in Rstudio Version 1.1.41462 using R version 3.6.363. Boxplots were produced using the default settings: The boxes represent the Interquartile Range (IQR) and span from the first quartile (25th percentile) to the third quartile (75th percentile) of the data. The midline of the box is the median. The whiskers extend up to 1.5 times the IQR from both ends of the box to the furthest datum within that distance. Data plotting beyond that distance are represented as individual points and considered outliers.
P-values were calculated in R studio using a paired Wilcoxon rank sum test with continuity correction and considered to be significant at a p-value below 0.05.
Results
In silico simulation
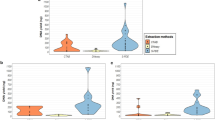
To evaluate the impact of sample pooling on ancient DNA detection, we conducted an in-silico simulation using fragmented mitochondrial sequences of Homo sapiens and Capra hircus, introducing varying levels of deamination and simulating pooling at different dilutions (Fig. 1). The simulation results (Supplementary Table S2) demonstrated stable read numbers and deamination levels across pooling steps with a non-significant decrease in read count (p > 0.05), indicating that an aDNA signal can be detected at a dilution up to 1:10-fold (Fig. 1).
In silico simulation of pooling. (a) Read numbers normalized by the unpooled state for each simulated mitochondrial dataset at different pooling steps, relative to the non-pooled ("NP") condition. (b) Simulated deamination values normalized by the unpooled state for each simulated mitochondrial dataset at different pooling steps.
Wet lab pooling of extracts does not decrease sensitivity
As part of the MINERVA project we screened archaeological sediments for ancient DNA employing a custom hybridization capture kit comprising 51 mammalian species (Supplementary Table S3)55. Due to the age of the sequences, aDNA displays distinct signs of degradation characterized by two main molecular features: strand breaks resulting in short DNA fragments and C to T substitutions caused by deamination64,65,66. We classified our samples using two parameters: (A) the quantity of DNA expressed as the number of unique reads mapping to the used mammalian reference panel and (B) the damage of these reads, measured by the occurrence of C to T transitions. We group our sediment samples into four states: (1) reads with a deamination level above 10%, and more than 1000 recovered unique reads, (2) reads with a deamination level above 10%, and fewer than 1000 recovered unique reads, (3) reads with a deamination level below 10% and more than 1000 recovered unique reads, and (4) reads with a deamination level below 10%, and fewer than 1000 recovered unique reads (Table 1). For convenience, all states are from here on referred to by their corresponding reference number (1–4) followed by the values for (a) deamination in percent and (b) the amounts of unique reads mapping to the mammalian reference panel. We further define state 1 as the threshold for a positive sediment sample (i.e., deamination level above 10% and > 1000 unique endogenous reads aligning to the mammalian reference genome panel used during the capture process). A failed sample is defined as a sample falling into states 2 (> 10%, < 1000), 3 (< 10%, > 1000), and 4 (< 10%, < 1000). In a different study on sedaDNA, a sample was considered to be positive for ancient hominin DNA if the number of DNA fragments assigned to hominins constituted at least 1% of the total identified fragments and if it had at least 10 fragments showing deamination which was significantly higher than 10%33. Another study also used a 10% deamination frequency at both ends as a cut-off (binomial CI ≥ 10%)34.
To evaluate our pooling simulation in a laboratory setting, we selected eight test samples to represent these four states (two samples per state according to earlier screening analyses). However, two of the test samples had to be excluded from the study. One test sample falling into state 2 (> 10%, < 1000) was contaminated with 1.67 × modern human DNA. Modern human contamination can be introduced while handling the sample on site or during the wet lab process and will be captured alongside ancient DNA fragments. We identified modern contamination using the software mapdamage 2-2.2.160, which assesses DNA damage patterns characteristic of ancient DNA, such as C-to-T substitutions at the ends of DNA fragments64,65,66. Additionally, the screening results for a state 3 (> 10%, < 1000) sample could not be reproduced; the sample displayed higher deamination after re-extraction. A negative sample, devoid of aDNA after screening (< 1000 unique reads mapping to the mammalian reference panel and < 10% of deamination) was chosen for the pooling. This sample had an average deamination of 4.7% at both ends and 651 reads aligning to our mammalian mtDNA panel. The extract of each test sample was pooled with the extract of our chosen negative sediment sample in increasing amounts up to a dilution of 1:4 (Fig. 2). This process was performed in two sets for each state, with two distinct candidate samples meeting states 1–4. To ensure accuracy, each dilution level was replicated three times (Supplementary Table S1).
Streamlining of initial screening of samples followed by pooling. Sediment samples were individually screened to distinguish samples with preserved ancient DNA (positive) from samples devoid of aDNA (negative). The extracts of positive samples were pooled with up to 4 extracts of a negative sample to recreate a pool testing method. The presence of an aDNA signal was evaluated by the deamination signal and the number of unique reads.
The experiment revealed that deamination remains stable with increased pooling in three states: state 1 (> 10%, > 1000, R2 = − 0.73), state 3 (< 10%, > 1000, R2 = 0.45), and state 4 (< 10%, < 1000, R2 = − 0.86), as shown in Fig. 3b. Notably, even in state 4, characterized by low deamination (< 10%) and low read levels (< 1000), the signal for the target aDNA could be maintained. In contrast, state 2 (> 10%, < 1000, R2 = − 0.94) exhibited a significant decrease in deamination level with increased pooling (p = 0.01789).
Average normalized read numbers percent of deamination across pooling levels. (a) Average read numbers of sediment samples normalized by the highest value across pooling levels. Each dataset is represented by a unique color in the legend. We calculated the differences between the unpooled state with each pooling step using a paired Wilcoxon rank sum test with continuity correction and considered it to be significant at a p-value below 0.05. Significance is indicated by a black asterisk. (b) Percent of deamination measured in sediment samples across increasing extract-pooling steps in comparison to the non-pooled (“NP”) sample. The colors indicate the four different states that sedaDNA samples commonly fall into, specified in Table 1. Different shapes within the same states indicate different datasets.
We observed an increase in normalized average read numbers when pooling was implemented (Fig. 3a). Despite considerable variation within each pooling level (SD = 0.12–0.23), there was a significant increase in average reads between the non-pooled state and the 1:3 dilution level (p = 0.018) as well as the 1:4 dilution level (p = 0.011). DNA yields were on average increased by 17, 47, 33, and 45 percent for pooling levels 1–4, respectively (Table 2). Overall, the DNA yield across all pooling levels was increased by 1.36-fold, compared to the average unpooled sample.
A cost-efficient method
The screening of sedaDNA samples is a laborious and expensive undertaking. We have modeled multiple situations in order to gain insights into the cost and labor reduction of this approach. We estimate that under conditions where 10% of the samples exhibit the characteristics previously defined as positive, the implementation of our approach would result in a 50% reduction in the number of libraries prepared. In the case of a 5% positivity rate, this would amount to a 75% decrease in the number of prepared libraries. Thus, we suggest that the pooling of five extracts is a cost-effective strategy in scenarios where the prevalence of positive samples is less than 20% among the total.
Discussion
With our extract-pooling approach, we set out to define a cost-efficient and high-throughput method for screening archeological sediments for aDNA. When pooling was implemented, we observed a cost reduction of approximately 50–75%, including costs for reagents and sequencing. Hands-on time in the lab was reduced to one-fifth. We demonstrate that the pre-PCR pooling of Paleolithic sediment extracts does not reduce the number of aDNA reads detected in double-stranded libraries53 across all pooling levels: extract-pooling, in fact, significantly (p = 0.0001) increases the efficiency of target aDNA recovery across all four states on average by 1.36-fold (Fig. 3a). While we observe an increase in unique DNA fragments mapping to the target panel with reduced extract input volume in double-stranded libraries, we acknowledge that understanding the underlying mechanisms requires experimental investigation beyond the scopes of this framework. One of the potential explanations for the observed increase could be the presence of substances in the sediment exerting a negative effect on PCR amplification. Inhibition is a prevalent obstacle in many PCR related applications such monitoring of foodborne diseases67,68,69,70, water and environmental surveillance71,72,73, plant pathogen detection74,75,76, soil, and sediment analyses77,78. For a detailed review on the occurrence, properties, and methods to remove inhibitors see79. Common PCR inhibitors present in the organic component of sediment include plant-derived substances such as tannic, fulvic, and humic acids77,78,80,81,82, as well as Phenols, Polysaccharides, Pectin, Polysaccharides, and Xylan83,84,85. Other substances known to impair PCR analysis include calcium ions, which compete with magnesium ions (a common component in commercial extraction kits) for binding to DNA polymerases, while humic acids interact directly with nucleic acids81. These inhibitors can be co-extracted with DNA from complex environmental samples like sediments and interfere with enzymatic reactions in qPCR and PCR amplification due to inhibition of polymerase activity86. This can lead to reduced PCR signals or an elevated number of false negative results87,88. Besides the diverse nature and mechanisms of PCR inhibitors, their concentration, which can vary within a sample, plays an important role in PCR results89. Many methodological approaches have been introduced to minimize the adverse effects of inhibitors at different steps of the wet lab pipeline. There are several treatments to reduce the co-purified amount of inhibitors during DNA extraction, such as adding polyvinylpolypyrrolidone (PVPP)90,91, hexadecyltrimethylammonium bromide (CTAB)90,92,93, and aluminum ammonium sulfate (AINH4(SO4)2)94. Further methods include the introduction of hydroxypatite columns95, column chromatography90,96, cesium chloride density centrifugations97,98,99,100, and DNA recovery from agarose after gel electrophoresis90,92,101. While most of these methods are laborious and and can effectively reduce the recovery of target DNA90,94, the dilution of extracts has been found to effectively mitigate inhibition; A recent study investigating PCR inhibitor removal methods found that the pre-dilution of extracts led to maximum amplification results71. The success of dilution and commercial PCR removal kits were, however, dependent on the initial concentration of inhibitors.
Our results show that a reduction of extract input into double-stranded libraries increases DNA yield significantly when comparing read numbers of the unpooled sample (25 µL) with those of pooling level 1:3 with 6.25 µL (p = 0.018) and pooling level 1:4 with 5 µL input (p = 0.011) (Fig. 3a). A study conducted on bones and teeth demonstrated that the preparation of single-stranded libraries with minimal extract input reduces the effects of inhibition102. A reduction of sample input in qPCR analyses provides additional support for our hypothesis that inhibition can be mitigated through this approach, as demonstrated by previous studies103,104. The extensive dilution of a DNA extract (often up to 400-fold), is a common practice across different scientific disciplines to circumvent inhibition87,103,105,106. As a consequence, the target DNA will simultaneously be diluted along with the inhibitory substances in PCR based methods107,108. In cases of underrepresented taxa or genes of interest, a dilution may even result in failed detection109,110. Another study, therefore, urges a moderate extract dilution of 40–60-fold to effectively reduce qPCR inhibition104.
Besides the reduction of PCR inhibiting agents, another potential explanation for the observed read increase is the improvement of PCR template availability in complex sediment samples: PCR is a highly sensitive process that can be affected by the availability and quality of endogenous DNA template and depends on the concentration and availability of PCR reagents, such as polymerase and dNTPs. In diverse environmental samples with high amounts of background DNA from non-target sources, endogenous DNA may be present in low quantities, thus reagents might be the limiting factor in a PCR reaction. Diluting the sample can reduce overall concentration, thereby improving the availability of PCR reagents for target template. This enhancement can lead to more efficient and consistent amplification, resulting in higher edogenous read numbers. We hypothesize that in our framework, the reduction of test sample input combined with increased amounts of a negative sample, containing very low amounts of target material mapping to the reference panel55, results in elevated read numbers.
To ensure consistency across our experimental set-up, the same DNA extraction protocol51,52 and library protocol53 were used for all samples included. This standardization rules out the possibility of any protocol-specific enhancements or adaptations contributing to the increase in reads within the experiment. Additionally, the same extract was used as a negative pooling extract consistently across the dilution series, ensuring that intrinsic properties of the sediment samples remained constant. Overall, this uniform approach across all samples eliminates the likelihood of variations in DNA extraction techniques, sample handling, or intrinsic sediment properties affecting the results.
The main objective of our experimental set-up was to mimic extract pooling of sedaDNA samples; This required the use of an authentic negative sediment sample for the dilution of the test samples. Therefore, the effect of inhibition was not specifically controlled for in this framework and we acknowledge that this aspect requires further testing. Moreover, due to the diverse nature and mechanisms of inhibitory substances, no one combination of PCR reagents can be used to work uniformly well against all of them, as tested by86. Hence, a test for the individual identification of inhibitory agents in different types of sediment in order to adapt extraction, purification, and PCR protocols has yet to be developed.
The minimal extract volume converted into double-stranded libraries in our study was 5 µL, which equals a 1:4 dilution of the regular input. In our approach, we intentionally pooled a positive sediment extract with increasing volumes of a negative extract. However, it is important to note that when applying pooling to several previously unscreened extracts, in which case the extent of inhibitors is unknown, we recommend pooling at small volumes. Another unknown factor in unscreened extracts is the extent of DNA preservation. Therefore, we cannot make a generalized conclusion that dilution consistently leads to increased DNA yields, as it relies on the assumption that the concentration of humic substances corresponds to DNA content, which would require further testing.
Additional experiments will be necessary to test the optimal volume of extract converted into libraries to examine the potential for even smaller volumes than those employed in this study for enhanced DNA recovery. We recommend conducting similar experiments to assess the optimal input concentrations of both extracts and pre-PCR libraries in the context of ancient DNA.
The extent of deamination remains stable as pooling increases, with one exception observed in state 2 (> 10%, < 1000) (Fig. 3b). For samples exhibiting the characteristics of state 2, it is plausible that the introduction of increased dilution with the negative sample containing a 2.4 × modern human contamination masks the impact of the limited aDNA portion within the positive sample. This aligns with the observations in Fig. 3, demonstrating a gradual reduction in C to T transitions with each increment in pooling. Notably, a parallel observation can be made for another sample categorized under state 2 which was ultimately excluded from the dataset due to a 1.67 × modern human contamination. This sample displayed a substantial reduction in its deamination signal as pooling intensified (Supplementary Fig. S2). This result supports the hypothesis that modern DNA can effectively mask the distinctive markers of aDNA: The various iterations of PCR amplification during the laboratory pipeline can introduce a technical bias towards the amplification of the more abundant and less fragmented modern DNA. Characteristic aDNA damage patterns including miscoding lesions and single-strand breaks111 can block polymerases, leading to chimeric sequences or the termination of amplification112. Consequently, this phenomenon leads to the prioritized sequencing of modern sequences over ancient ones during the screening process.
In this study, we employed equal-volume pooling of non-normalized extracts (according to concentration) to achieve increased pooling levels. Rather than merging different extracts, we utilized a single (test) extract and combined it with progressively larger volumes of another (negative) extract to attain higher pooling levels. This strategy was used to simulate the potential low success rate of retrieving ancient mammalian DNA from sedaDNA samples38,39. It is important to note that equal-volume pooling does not guarantee equal representation of each extract within the pool. When applying our extract-pooling method for the initial screening of different sediment samples, equal mass pooling might not circumvent heterogeneity in the extract pool. Quantifying the concentration of a sample’s extract prior to pooling is reasonable and necessary when dealing with modern DNA, as the target (endogenous) DNA would typically approximate 100 percent. However, for ancient DNA, especially ancient sedimentary DNA, it is more complicated as the total amount of DNA within a sample does not reflect endogenous target DNA content (which, in our case is mostly human and faunal DNA); The vast majority of the DNA stems from microorganisms, plants, and faunal components31,113,114. Thus, a bias could also, arguably, arise from pooling by qubit quantification: A sample with overall high DNA concentration may have a very low amount of human or faunal DNA and would thus be underrepresented if corrected for the DNA concentration.
It is a widely used approach in the field of eDNA to pool extracts in the order of hundreds to determine the presence or absence of organisms20. However, when the objective is to identify individual positive samples for downstream analyses, an elevated number is suboptimal as a large (financial) effort is required to produce individual libraries from sample pools that tested positive. Therefore, it is clear that an equilibrium must be found. To date, no studies have specifically addressed the positivity rates of sedaDNA from archaeological contexts and there is yet no compelling data on the success rate and its determinants. Based on extensive screening efforts published so far31,33,34, it appears that positivity—hitherto defined as the proportion of samples with sufficient DNA for classifying mammalian reads—may be around 10–15%. In our laboratory experiment, we established an upper threshold of five extracts per individual pool, assuming a more conservative 20% success rate for our samples. This decision was reached to ensure a balance between analytical efficiency and resource optimization, even if the success rate is high. One of the main objectives of sedaDNA analyses in the light of paleogenomics is to produce high-quality DNA data from hominins or fauna. When defining positivity as samples with enough reads to recover substantial information through population genetics, the success rate drops below 5%. The strength of our pooling method lies in its flexibility: the number of samples pooled can be adapted to specific research objectives. If the goal is to assess the overall preservation (presence or absence) of ancient DNA at a site, the number of samples within a pool can be increased. Conversely, if the objective is to identify hominins or specific mammals, the number of samples can be decreased (as pools with aDNA signals will have to be analyzed individually). Additionally, a subset of samples can first be pooled and pre-screened to determine the general state of aDNA preservation at a site, thereby concluding the optimal number of samples to pool.
Recently, there has been another notable methodological advancement aimed at enhancing the cost-effectiveness of large-scale sample screening, focusing on a step much closer to the end of the laboratory pipeline, referred to as multiplex capture50. In contrast to our method, multiplex capture involves individual library preparation, double-indexing, and initial amplification. The streamlining of lab processes is achieved at the step of hybridization capture, through the pooling of multiple double-indexed libraries into a single capture reaction. While reducing the costs for capture reactions considerably, the majority of lab processes have to be conducted individually. This method could additionally be tested in combination with our extract-pooling method.
In conclusion, we have successfully demonstrated two key findings that have important implications for the field of sedaDNA research: 1) Preservation of sensitivity: Our study shows that the pooling of up to five extracts did not compromise the sensitivity of ancient DNA detection. This finding implies that extract pooling can be employed as a cost-effective and efficient strategy for large-scale sediment sample screening without sacrificing the ability to detect ancient DNA. 2) Effective reduction of PCR drawbacks: Our results highlight that negative effects of undetermined nature in double-stranded libraries can be significantly reduced by using low extract input volumes. Although the nature of this observation requires further experimental testing, our approach allows researchers to optimize extract input volumes, leading to enhanced recovery of ancient DNA. Overall, this cost-effective and high-throughput approach has the potential to advance the field and facilitate the study of ancient ecosystems and populations.
Data availability
The datasets generated during the current study are available in the European Nucleotide Archive (ENA) through the accession number PRJEB67982.
References
Ariza, M. et al. Plant biodiversity assessment through soil eDNA reflects temporal and local diversity. Methods Ecol. Evol. 14, 415–430 (2023).
Allen, M. C. et al. Sampling environmental DNA from trees and soil to detect cryptic arboreal mammals. Sci. Rep. 13, 180 (2023).
Antich, A. et al. Marine biomonitoring with eDNA: Can metabarcoding of water samples cut it as a tool for surveying benthic communities?. Mol. Ecol. 30, 3175–3188 (2021).
Blattner, L., Ebner, J. N., Zopfi, J. & von Fumetti, S. Targeted non-invasive bioindicator species detection in eDNA water samples to assess and monitor the integrity of vulnerable alpine freshwater environments. Ecol. Indic. 129, 107916 (2021).
Lynggaard, C. et al. Airborne environmental DNA for terrestrial vertebrate community monitoring. Curr. Biol. 32, 701-707.e5 (2022).
Johnson, M. D., Fokar, M., Cox, R. D. & Barnes, M. A. Airborne environmental DNA metabarcoding detects more diversity, with less sampling effort, than a traditional plant community survey. BMC Ecol. Evol. 21, 218 (2021).
Ogram, A., Sayler, G. S. & Barkay, T. The extraction and purification of microbial DNA from sediments. J. Microbiol. Methods 7, 57–66 (1987).
Olsen, G. J., Lane, D. J., Giovannoni, S. J., Pace, N. R. & Stahl, D. A. Microbial ecology and evolution: A ribosomal RNA approach. Annu. Rev. Microbiol. 40, 337–365 (1986).
Pace, N. R., Stahl, D. A., Lane, D. J. & Olsen, G. J. The analysis of natural microbial populations by ribosomal RNA sequences. In Advances in Microbial Ecology (ed. Marshall, K. C.) 1–55 (Springer US, 1986).
Shi, T., Reeves, R. H., Gilichinsky, D. A. & Friedmann, E. I. Characterization of viable bacteria from Siberian permafrost by 16S rDNA sequencing. Microb. Ecol. 33, 169–179 (1997).
Coolen, M. J. & Overmann, J. Analysis of subfossil molecular remains of purple sulfur bacteria in a lake sediment. Appl. Environ. Microbiol. 64, 4513–4521 (1998).
Willerslev, E. et al. Long-term persistence of bacterial DNA. Curr. Biol. 14, R9-10 (2004).
Fierer, N. & Jackson, R. B. The diversity and biogeography of soil bacterial communities. Proc. Natl. Acad. Sci. U. S. A. 103, 626–631 (2006).
Johnson, S. S. et al. Ancient bacteria show evidence of DNA repair. Proc. Natl. Acad. Sci. U. S. A. 104, 14401–14405 (2007).
Hofreiter, M., Mead, J. I., Martin, P. & Poinar, H. N. Molecular caving. Curr. Biol. 13, R693–R695 (2003).
Willerslev, E. et al. Diverse plant and animal genetic records from Holocene and Pleistocene sediments. Science 300, 791–795 (2003).
Valentini, A., Pompanon, F. & Taberlet, P. DNA barcoding for ecologists. Trends Ecol. Evol. 24, 110–117 (2009).
Taberlet, P., Coissac, E., Pompanon, F., Brochmann, C. & Willerslev, E. Towards next-generation biodiversity assessment using DNA metabarcoding. Mol. Ecol. 21, 2045–2050 (2012).
Glenn, T. C. Field guide to next-generation DNA sequencers. Mol. Ecol. Resour. 11, 759–769 (2011).
Rijal, D. P. et al. Sedimentary ancient DNA shows terrestrial plant richness continuously increased over the Holocene in northern Fennoscandia. Sci. Adv. 7, eabf9557 (2021).
Jørgensen, T. et al. Islands in the ice: detecting past vegetation on Greenlandic nunataks using historical records and sedimentary ancient DNA meta-barcoding. Mol. Ecol. 21, 1980–1988 (2012).
ter Schure, A. T. M. et al. Sedimentary ancient DNA metabarcoding as a tool for assessing prehistoric plant use at the Upper Paleolithic cave site Aghitu-3, Armenia. J. Hum. Evol. 172, 103258 (2022).
Han, W. et al. Anthropogenic activities altering the ecosystem in Lake Yamzhog Yumco, southern Qinghai-Tibetan Plateau. Sci. Total Environ. 904, 166715 (2023).
Seeber, P. A. et al. Evaluation of lake sedimentary ancient DNA metabarcoding to assess fungal biodiversity in Arctic paleoecosystems. Environ. DNA 4, 1150–1163 (2022).
Jia, W. et al. Sedimentary ancient DNA reveals past ecosystem and biodiversity changes on the Tibetan Plateau: Overview and prospects. Quat. Sci. Rev. 293, 107703 (2022).
Giguet-Covex, C. et al. Long livestock farming history and human landscape shaping revealed by lake sediment DNA. Nat. Commun. 5, 1–7 (2014).
Bellemain, E. et al. Fungal palaeodiversity revealed using high-throughput metabarcoding of ancient DNA from arctic permafrost. Environ. Microbiol. 15, 1176–1189 (2013).
Parducci, L. et al. Shotgun environmental DNA, pollen, and macrofossil analysis of lateglacial lake sediments From Southern Sweden. Front. Ecol. Evol. 7, (2019).
Stahlschmidt, M. C. et al. Ancient mammalian and plant DNA from late quaternary stalagmite layers at Solkota Cave, Georgia. Sci. Rep. 9, 6628 (2019).
Gelabert, P. et al. Genome-scale sequencing and analysis of human, wolf, and bison DNA from 25,000-year-old sediment. Curr. Biol. 31, 3564-3574.e9 (2021).
Slon, V., Hopfe, C., Weiß, C. L. & Mafessoni, F. Neandertal and denisovan DNA from pleistocene sediments. Science 356, 605–608 (2017).
Zhang, D. et al. Denisovan DNA in late pleistocene sediments from Baishiya Karst Cave on the Tibetan Plateau. Science 370, 584–587 (2020).
Zavala, E. I. et al. Pleistocene sediment DNA reveals hominin and faunal turnovers at Denisova Cave. Nature 595, 399–403 (2021).
Vernot, B. et al. Unearthing Neanderthal population history using nuclear and mitochondrial DNA from cave sediments. Science 372, (2021).
Pietramellara, G. et al. Extracellular DNA in soil and sediment: Fate and ecological relevance. Biol. Fertil. Soils 45, 219–235 (2009).
Parducci, L. et al. Ancient plant DNA in lake sediments. New Phytol. 214, 924–942 (2017).
Nagler, M., Insam, H., Pietramellara, G. & Ascher-Jenull, J. Extracellular DNA in natural environments: Features, relevance and applications. Appl. Microbiol. Biotechnol. 102, 6343–6356 (2018).
Slon, V. et al. Extended longevity of DNA preservation in Levantine Paleolithic sediments, Sefunim Cave, Israel. Sci. Rep. 12, 14528 (2022).
Massilani, D. et al. Microstratigraphic preservation of ancient faunal and hominin DNA in Pleistocene cave sediments. Proc. Natl. Acad. Sci. U. S. A. 119, (2022).
Dorfman, R. The detection of defective members of large populations. Ann. Math. Stat. 14, 436–440 (1943).
García, Z. et al. Evaluation of a pooling method for routine anti-HCV screening of blood donors to lower the cost burden on blood banks in countries under development. J. Med. Virol. 49, 218–222 (1996).
Roth, W. K., Weber, M. & Seifried, E. Feasibility and efficacy of routine PCR screening of blood donations for hepatitis C virus, hepatitis B virus, and HIV-1 in a blood-bank setting. Lancet 353, 359–363 (1999).
Quinn, T. C. et al. Feasibility of pooling sera for HIV-1 viral RNA to diagnose acute primary HIV-1 infection and estimate HIV incidence. AIDS 14, 2751–2757 (2000).
Mutesa, L. et al. A pooled testing strategy for identifying SARS-CoV-2 at low prevalence. Nature 589, 276–280 (2021).
Sawicki, R., Korona-Glowniak, I., Boguszewska, A., Stec, A. & Polz-Dacewicz, M. Sample pooling as a strategy for community monitoring for SARS-CoV-2. Sci. Rep. 11, 3122 (2021).
Tveit, A., Schwacke, R., Svenning, M. M. & Urich, T. Organic carbon transformations in high-Arctic peat soils: Key functions and microorganisms. ISME J. 7, 299–311 (2013).
Keshri, J., Yousuf, B., Mishra, A. & Jha, B. The abundance of functional genes, cbbL, nifH, amoA and apsA, and bacterial community structure of intertidal soil from Arabian Sea. Microbiol. Res. 175, 57–66 (2015).
Song, Z. et al. Effort versus reward: Preparing samples for fungal community characterization in high-throughput sequencing surveys of soils. PLoS One 10, e0127234 (2015).
Hestetun, J. T., Lanzén, A., Skaar, K. S. & Dahlgren, T. G. The impact of DNA extract homogenization and replication on marine sediment metabarcoding diversity and heterogeneity. Environ. DNA 3, 997–1006 (2021).
Zavala, E. I. et al. Quantifying and reducing cross-contamination in single- and multiplex hybridization capture of ancient DNA. Mol. Ecol. Resour. 22, 2196–2207 (2022).
Dabney, J. et al. Complete mitochondrial genome sequence of a Middle Pleistocene cave bear reconstructed from ultrashort DNA fragments. Proc. Natl. Acad. Sci. U. S. A. 110, 15758–15763 (2013).
Korlević, P. et al. Reducing microbial and human contamination in DNA extractions from ancient bones and teeth. Biotechniques 59, 87–93 (2015).
Meyer, M. & Kircher, M. Illumina sequencing library preparation for highly multiplexed target capture and sequencing. Cold Spring Harb. Protoc. 2010, db.prot5448 (2010).
Kircher, M., Sawyer, S. & Meyer, M. Double indexing overcomes inaccuracies in multiplex sequencing on the Illumina platform. Nucleic Acids Res. 40, e3 (2012).
Tejero, J.-M. et al. Cervidae antlers exploited to manufacture prehistoric tools and hunting implements as a reliable source of ancient DNA. Heliyon 10, e31858 (2024).
Rohland, N. et al. Three assays for in-solution enrichment of ancient human DNA at more than a million SNPs. Genome Res. 32, 2068–2078 (2022).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Jónsson, H., Ginolhac, A., Schubert, M., Johnson, P. L. F. & Orlando, L. mapDamage2.0: Fast approximate Bayesian estimates of ancient DNA damage parameters. Bioinformatics. 29, 1682–1684 (2013).
Okonechnikov, K., Conesa, A. & García-Alcalde, F. Qualimap 2: Advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics 32, 292–294 (2016).
Team & R. S,. RStudio: Integrated Development for R (RStudio. Inc., 2015).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2013). http://www.R-project.org/.
Briggs, A. W. et al. Patterns of damage in genomic DNA sequences from a Neandertal. Proc. Natl. Acad. Sci. U. S. A. 104, 14616–14621 (2007).
Brotherton, P. et al. Novel high-resolution characterization of ancient DNA reveals C > U-type base modification events as the sole cause of post mortem miscoding lesions. Nucleic Acids Res. 35, 5717–5728 (2007).
Sawyer, S., Krause, J., Guschanski, K., Savolainen, V. & Pääbo, S. Temporal patterns of nucleotide misincorporations and DNA fragmentation in ancient DNA. PLoS One 7, e34131 (2012).
Suther, C. & Moore, M. D. Quantification and discovery of PCR inhibitors found in food matrices commonly associated with foodborne viruses. Food Sci. Hum. Wellness 8, 351–355 (2019).
Andersen, M. R. & Omiecinski, C. J. Direct extraction of bacterial plasmids from food for polymerase chain reaction amplification. Appl. Environ. Microbiol. 58, 4080–4082 (1992).
Jaykus, L. A., De Leon, R. & Sobsey, M. D. A virion concentration method for detection of human enteric viruses in oysters by PCR and oligoprobe hybridization. Appl. Environ. Microbiol. 62, 2074–2080 (1996).
Richards, G. P. Limitations of molecular biological techniques for assessing the virological safety of foods. J. Food Prot. 62, 691–697 (1999).
Hamza, I. A. & Leifels, M. Assessment of PCR inhibitor removal methods to monitor viruses in environmental water samples: DAX-8 outperforms competitors. Water Air Soil Pollut. 235, 20 (2024).
Haramoto, E. et al. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 135, 168–186 (2018).
Hamza, I. A. et al. Detection of human viruses in rivers of a densly-populated area in Germany using a virus adsorption elution method optimized for PCR analyses. Water Res. 43, 2657–2668 (2009).
de Chaves, M.Q.-G. et al. A new and accurate qPCR protocol to detect plant pathogenic bacteria of the genus ‘Candidatus Liberibacter’ in plants and insects. Sci. Rep. 13, 3338 (2023).
Kageyama, K., Komatsu, T. & Suga, H. Refined PCR protocol for detection of plant pathogens in soil. J. Gen. Plant Pathol. 69, 153–160 (2003).
Approaches for eliminating PCR inhibitors and designing PCR primers for the detection of phytopathogenic fungi. Crop Prot. 26, 145–161 (2007).
Albers, C. N., Jensen, A., Bælum, J. & Jacobsen, C. S. Inhibition of DNA polymerases used in Q-PCR by structurally different soil-derived humic substances. Geomicrobiol. J. 30, 675–681 (2013).
Tebbe, C. C. & Vahjen, W. Interference of humic acids and DNA extracted directly from soil in detection and transformation of recombinant DNA from bacteria and a yeast. Appl. Environ. Microbiol. 59, 2657–2665 (1993).
Schrader, C., Schielke, A., Ellerbroek, L. & Johne, R. PCR inhibitors—Occurrence, properties and removal. J. Appl. Microbiol. 113, 1014–1026 (2012).
Hering, J. G. & Morel, F. M. Humic acid complexation of calcium and copper. Environ. Sci. Technol. 22, 1234–1237 (1988).
Sutlovic, D., Gamulin, S., Definis-Gojanovic, M., Gugic, D. & Andjelinovic, S. Interaction of humic acids with human DNA: Proposed mechanisms and kinetics. Electrophoresis 29, 1467–1472 (2008).
Matheson, C. D., Gurney, C., Esau, N. & Lehto, R. Assessing PCR inhibition from humic substances. Open Enzym. Inhib. J. 3, 38–45 (2014).
Demeke, T. & Adams, R. P. The effects of plant polysaccharides and buffer additives on PCR. Biotechniques 12, 332–334 (1992).
Henson, C., Truchot, D. & Canevello, A. PTSD and PTG in French and American firefighters: A comparative study. Int. J. Environ. Res. Public Health 19, 11973 (2022).
Wei, T., Lu, G. & Clover, G. Novel approaches to mitigate primer interaction and eliminate inhibitors in multiplex PCR, demonstrated using an assay for detection of three strawberry viruses. J. Virol. Methods 151, 132–139 (2008).
Uchii, K. et al. Comparison of inhibition resistance among PCR reagents for detection and quantification of environmental DNA. Environ. DNA 1, 359–367 (2019).
McKee, A. M., Spear, S. F. & Pierson, T. W. The effect of dilution and the use of a post-extraction nucleic acid purification column on the accuracy, precision, and inhibition of environmental DNA samples. Biol. Conserv. 183, 70–76 (2015).
Opel, K. L., Chung, D. & McCord, B. R. A study of PCR inhibition mechanisms using real time PCR. J. Forensic Sci. 55, 25–33 (2010).
Rossen, L., Nørskov, P., Holmstrøm, K. & Rasmussen, O. F. Inhibition of PCR by components of food samples, microbial diagnostic assays and DNA-extraction solutions. Int. J. Food Microbiol. 17, 37–45 (1992).
Zhou, J., Bruns, M. A. & Tiedje, J. M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. https://doi.org/10.1128/aem.62.2.316-322.1996 (1996).
Frostegård, A. et al. Quantification of bias related to the extraction of DNA directly from soils. Appl. Environ. Microbiol. 65, 5409–5420 (1999).
Malik, M., Kain, J., Pettigrew, C. & Ogram, A. Purification and molecular analysis of microbian DNA from compost. J. Microbiol. Methods 20, 183–196 (1994).
Cho, J.-C., Lee, D.-H., Cho, Y., Cho, J.-C. & Kim, S.-J. Direct extraction of DNA from soil for amplification of 16S rRNA gene sequences by polymerase chain reaction. J. Microbiol. 34, 229–235 (1996).
Braid, M. D., Daniels, L. M. & Kitts, C. L. Removal of PCR inhibitors from soil DNA by chemical flocculation. J. Microbiol. Methods 52, 389–393 (2003).
Torsvik, V. L. Isolation of bacterial DNA from soil. Soil Biol. Biochem. 12, 15–21 (1980).
Cullen, D. W. & Hirsch, P. R. Simple and rapid method for direct extraction of microbial DNA from soil for PCR. Soil Biol. Biochem. 30, 983–993 (1998).
Holben, W. E., Jansson, J. K., Chelm, B. K. & Tiedje, J. M. DNA probe method for the detection of specific microorganisms in the soil bacterial community. Appl. Environ. Microbiol. 54, 703–711 (1988).
Leff, L. G., Dana, J. R., McArthur, J. V. & Shimkets, L. J. Comparison of methods of DNA extraction from stream sediments. Appl. Environ. Microbiol. 61, 1141–1143 (1995).
Lovell, C. R. & Piceno, Y. Purification of DNA from estuarine sediments. J. Microbiol. Methods 20, 161–174 (1994).
Walia, S., Khan, A. & Rosenthal, N. Construction and applications of DNA probes for detection of polychlorinated biphenyl-degrading genotypes in toxic organic-contaminated soil environments. Appl. Environ. Microbiol. 56, 254–259 (1990).
Moré, M. I., Herrick, J. B., Silva, M. C., Ghiorse, W. C. & Madsen, E. L. Quantitative cell lysis of indigenous microorganisms and rapid extraction of microbial DNA from sediment. Appl. Environ. Microbiol. 60, 1572–1580 (1994).
Glocke, I. & Meyer, M. Extending the spectrum of DNA sequences retrieved from ancient bones and teeth. Genome Res. 27, 1230–1237 (2017).
Schneider, S., Enkerli, J. & Widmer, F. A generally applicable assay for the quantification of inhibitory effects on PCR. J. Microbiol. Methods 78, 351–353 (2009).
Wang, H., Qi, J., Xiao, D., Wang, Z. & Tian, K. A re-evaluation of dilution for eliminating PCR inhibition in soil DNA samples. Soil Biol. Biochem. 106, 109–118 (2017).
Shutler, G. G. et al. Removal of a PCR Inhibitor and Resolution of DNA STR Types in Mixed Human-Canine Stains from a Five Year Old Case (ASTM International, 1999).
Imaizumi, K. et al. DNA typing of bone specimens—The potential use of the profiler test as a tool for bone identification. Leg. Med. 7, 31–41 (2005).
Bustin, S. A. et al. The MIQE guidelines: Minimum information for publication of quantitative real-time PCR experiments. Clin. Chem. 55, 611–622 (2009).
Lebuhn, M., Effenberger, M., Garcés, G., Gronauer, A. & Wilderer, P. A. Evaluating real-time PCR for the quantification of distinct pathogens and indicator organisms in environmental samples. Water Sci. Technol. 50, 263–270 (2004).
Lindberg, E., Albrechtsen, H.-J. & Jacobsen, C. S. Inhibition of real-time PCR in DNA extracts from aquifer sediment. Geomicrobiol. J. 24, 343–352 (2007).
Jane, S. F. et al. Distance, flow and PCR inhibition: eDNA dynamics in two headwater streams. Mol. Ecol. Resour. 15, 216–227 (2015).
Lindahl, T. lnstability and decay of the primary. Nature 362, 22 (1993).
Pääbo, S., Higuchi, R. G. & Wilson, A. C. Ancient DNA and the polymerase chain reaction. The emerging field of molecular archaeology. J. Biol. Chem. 264, 9709–9712 (1989).
Willerslev, E. & Cooper, A. Ancient DNA. Proc. Biol. Sci. 272, 3–16 (2005).
Pedersen, M. W. et al. Postglacial viability and colonization in North America’s ice-free corridor. Nature 537, 45–49 (2016).
Acknowledgements
This study was supported by a grant from the internal Research Platform MINERVA (https://minerva.univie.ac.at). P.G. was funded through COST iNEAL STSM Grant CA19141-8d068698. S.Sa. was funded by the Austrian Science Fund (FWF) M3108-G. The excavation and sampling of Velika Pećina in Kličevica was financially supported by the Croatian Science Foundation (NECEM, HRZZ‐IP‐2019‐04‐6649) and M.S. received funding from the European Research Council (ERC) MicroStratDNA project under Grant Agreement No. 101042570. The authors would like to express their gratitude to the following individuals and institutions for their invaluable contributions to this research: Paul Knabl for the artistic expertise demonstrated in creating the figures and Natalija Čondić and Archaeological Museum Zadar for various forms of assistance during the sampling of Velika Pećina in Kličevica.
Author information
Authors and Affiliations
Contributions
Study conceptualization: S.Sa., P.G., V.O., R.P. Methodological design: V.O., S.Sa., P.G., R.P., O.C., Experimental work: V.O., F.B., S.F., E.Z., S.Sz., F.TC., F.E., B.Z. Data analysis: V.O., P.G., and S.Sa. Archaeological conceptualization and sampling: I.K., M.B., L.G.S., M.R.G.M., B.G., M.B., J.K. Project administration and funding acquisition: R.P., T.R., S.M.K., M.S. Writing: V.O., S.Sa., P.G., R.P. with input from the rest of the authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Oberreiter, V., Gelabert, P., Brück, F. et al. Maximizing efficiency in sedimentary ancient DNA analysis: a novel extract pooling approach. Sci Rep 14, 19388 (2024). https://doi.org/10.1038/s41598-024-69741-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-69741-5
This article is cited by
-
Optimizing ancient DNA recovery from archaeological plant seeds
Scientific Reports (2025)