Abstract
The chloroplast genomes of wild loquat can help to determine their place in the history of evolution. Here, we sequenced and assembled two novel wild loquat’s chloroplast genomes, one is Eriobotrya elliptica, and the other is an unidentified wild loquat, which we named “YN-1”. Their sizes are 159,471 bp and 159,399 bp, respectively. We also assembled a cultivated loquat named ‘JFZ’, its chloroplast genome size is 159,156 bp. A comparative study was conducted with six distinct species of loquats, including five wild loquats and one cultivated loquat. The results showed that both E. elliptica and “YN-1” have 127 genes, one gene more than E. fragrans, which is psbK. Regions trnF-GAA-ndhJ, petG-trnP-UGG, and rpl32-trnL-UAG were found to exhibit high variability. It was discovered that there was a positive selection on rpl22 and rps12. RNA editing analysis found several chilling stress-specific RNA editing sites, especially in rpl2 gene. Phylogenetic analysis results showed that “YN-1” is closely related to E. elliptica, E. obovata and E. henryi.
Similar content being viewed by others
Introduction
The chloroplast genomes always contain two inverted repeats (IR), one large single copy (LSC), and one small single copy (SSC)1. The SSC can exhibit reverse complementary in different haplotypes of some chloroplast genomes2. Chloroplast genome sequences are useful markers for species identification3, whether using the entire chloroplast genome or specific fragments such as simple sequence repeats (SSR) or variable regions. Plant phylogenetic classification can also be studied using boundaries that vary among LSC, SSC, and IRs4. Most chloroplast genomes contain 110 to 130 genes, including protein-coding genes, rRNAs, and tRNA genes5. These genes are involved in fatty acid and amino acid synthesis pathways, as well as in the process of photosynthesis4,6. RNA editing of genes in the chloroplast genome is a post-transcriptional modification process, and these have been reported in some plants7,8,9. Cytidine to uridine conversion RNA editing events occur frequently, and occasionally an adenosine to inosine transition takes place8. The C-to-U editing rates were reported to be significantly reduced under heat or cold stress compared to normal conditions10.
Eriobotrya japonica (Thunb.) Lindl. (cultivated loquat, also known as common loquat) is a perennial fruit tree in the Rosaceae family, similar to apple and peach11,12. It is commonly grown in China and other countries12. The fruits of loquats are delicious, and their leaves are also used as a material in Chinese medicine containing triterpene13,14. Wild loquats are important materials for breeding programs aimed at cultivating loquats because they may be more resistant to stressful environments or have specific colors in their fruits15. Several wild loquats’ chloroplast genomes had been sequenced, like Eriobotrya malipoensis, Eriobotrya bengalensis et al.16,17,18, but no comparisons have been made among them.
Here, we sequenced and assembled the chloroplast genomes of two wild loquats, one is Eriobotrya elliptica, and the other is an unknown wild loquat (At present, we are calling it “YN-1”) that we found in Yunnan province, China. “YN-1” leaves are oval in shape; however, they are smaller than those of E. elliptica, measuring 10–15 cm long and 4.4–7 cm wide. Furthermore, the petiole base of YN-1 has hair, and its leaves are smooth and hairless. To date, the tallest "YN-1" tree we have seen is higher than ten meters in height. We also assembled the chloroplast genome of a cultivated loquat, Eriobotrya japonica ‘Jiefangzhong’. The chloroplast genomes of five wild loquats and one cultivated variety, "Jiefangzhong," were compared globally. Phylogenetic evolution analysis was conducted using all the published wild loquats, including the two newly assembled wild loquats and one cultivated loquat (Eriobotrya japonica ‘Jiefangzhong’).
Result
Chloroplast genomes global comparison
Here, we assembled two novel wild loquat chloroplast genomes, Eriobotrya elliptica (TY) and “YN-1” (an unknown wild loquat found in Yunnan Province), and one cultivated loquat chloroplast genome (‘Jiefangzhong’, JFZ). We also utilized three other publicly available wild loquats chloroplast genomes (Eriobotrya bengalensis, here named as NY; Eriobotrya malipoensis, here named as MLP; Eriobotrya fragrans, here named as XH) for a comprehensive comparison of the loquat chloroplast genomes.
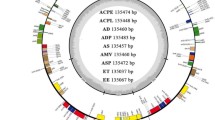
After assembly, the final chloroplast genome size of Eriobotrya elliptica (TY) was 159,471 bp and the chloroplast genome size of the newly discovered wild loquat “YN-1” was 159,399 bp (Fig. 1). The chloroplast genome size of “JFZ” was 159,156 bp (Fig. 1). Although the inverted repeated regions (IRs) of “YN-1” and “TY” only have 1 bp differentiation, the sizes of the large single copy (LSC) and small single copy (SSC) exhibit more differences. The LSC of “TY” was 87,426 bp and the LSC of “YN-1” was 87,359 bp, while the SSC of “TY” was 19,353 bp and 19,346 bp for “YN-1”. Both the chloroplast genomes of “TY” and “YN-1” contain 83 protein-coding genes, 8 rRNA, and 36 tRNA (Fig. 2), similar to the other three kinds of loquats. This is in contrast to the “XH” loquat, which is missing the psbK gene. After analyzing the boundaries of the junction sites of the chloroplast genomes, the results showed that there are three genes located in the junction sites: rps19, ndhF, and ycf1. Both ndhF in “MLP” and rps19 in “JFZ” are 3-4 bp different from their corresponding genes in the other four kinds of loquats. Gene ycf1 in “JFZ” and “NY” showed more variability compared to other loquats, with 30 and 36 bp differences, respectively (Fig. 3). Further alignment analysis revealed that both ycf1 genes in “JFZ” and “NY” contained two identical deletions compared to the ycf1 gene located in the other four kinds of loquats. Additionally, there was a unique 6 bp deletion “TATCAA”, which is only present in the ycf1 of “NY” (Fig. S1).
Gene map of the Eriobotrya elliptica, “YN-1”, and “JFZ”. Thick lines indicate the extent of the inverted repeat regions (IRa and IRb), which separate the genome into small (SSC) and large (LSC) single copy regions. Genes drawn inside the circle are transcribed clockwise, and those outside are transcribed counter clockwise. Different colors represent different functional groups of genes.
Comparison of gene number, gene type, and different region sizes of of different species of loquats.
Comparison of the borders of LSC, SSC, and IR regions of chloroplast genomes of different species of loquats.
Structural variation and DNA polymorphism
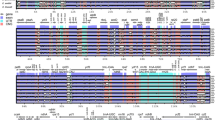
We compared the similarity of the six chloroplast genome sequences using mVISTA with the“YN-1” chloroplast genome as a reference. A high degree of synteny exists between the six chloroplast genomes, but the sequences of the CNS are highly variable (Fig. S2). In order to elucidate the sequence nucleotide divergence among six chloroplast genomes, we used DnaSP6.0 to analyze the sequence nucleotide variability within 600 bp window, 200 bp as a step. The highest average sequence divergence is located in trnF-GAA-ndhJ, followed by petG-trnP-UGG, and rpl32-trnL-UAG (Fig. 4). The average DNA sequence variation of different regions differs among various regions, LSC > SSC > IRs (Table S1). We also detected single nucleotide polymorphisms (SNPs) and indels between the two genomes, and we found 717 SNPs and 213 indels (Table S2).
Sliding window analysis (window length: 600 bp, step size: 200 bp) of the whole chloroplast genomes of six different kinds of loquat. X-axis, position of the mid point of a window; Y-axis, nucleotide diversity of each window.
SSR and repetitive sequences
In total, five types of SSRs were detected in the six chloroplast genomes; while only the “TY” and “XH” had tri nucleotide type SSRs. Besides, the penta nucleotide type SSRs were only missed in the “XH” loquat. The number of SSRs detected in different chloroplast genomes ranges from 91 to 99, the “XH” loquat contained the most amount of SSRs (Figure 5B, Table S3). All six chloroplast genomes had 50 repetitive sequences, and only the “MLP” loquat did not have complementary repeats. In terms of the number of repetitive sequences, the “MLP” loquat seems very specific; it contained the highest number of inverted repeats while having the fewest palindromic repeats, only 1. The “NY” loquat also had more inverted repeats compared to others, except for the “MLP” loquat. The most popular type of repetitive sequence detected was direct repeats (Fig. 5A). The repeat sizes mainly ranged from 20 to 50 bp, with “YN-1” containing two direct repeats with a size larger than 90 bp, the longest one is 110 bp. The “MLP” loquat chloroplast genome appeared to be very specific; all the repeats’ sizes are larger than 70, with an average size of 78.7 bp (Table S4).
Repeats (A) and SSRs (B) number comparison of six different species of loquats.
Codon usage bias
Protein-coding genes with more than 300 nucleotides were used to explore the codon usage pattern of the six loquat chloroplast genomes19. First, we determined each gene’s relative synonymous codon usage (RSCU) value. Across the six varieties of loquats, we observed no discernible differences (Fig S3, Table S5). Then we investigated the base composition of each position of each codon. For the entire gene, the GC content of the “JFZ” and “NY” protein-coding genes is lower than the other four loquats, the median GC content value of “JFZ” and “NY” is 0.385, while for the rest, this value is 0.39 (Fig. 6A). The GC content of the first codon (GC1) in different loquat chloroplast genomes varies. The GC1 median value of the “JFZ” loquat is the highest, which is 0.485; while the GC1 median value of “NY” is the lowest, which is 0.475. The median value of GC1 for the other four kinds of loquat is 0.48. There is no difference in the GC content median value of the second codon (GC2) and the third codon (GC3) among all six kinds of loquat. The correlation between GC12 and GC3 was significant (R2 < 0.01), indicating that mutations affect the CUB in all six types of loquat chloroplast genomes (Fig. 6B). The codon adaptation index (CAI) indicates directional synonymous codon usage bias20, only a minor difference between the CAI values of all the six kinds of loquat. The CAI value of “JFZ” is 0.216, while the CAI value for all the remaining varieties is 0.215 (Fig. 6C). The effective number of codons (ENC) value is used to quantify how far the codon usage of a gene departs from the equal usage of synonymous codons21. The ENCs of all the chloroplast genes in the six kinds of loquats range from 34 to 56, indicating diverse and relatively weak CUB in these genes. The relationship between the base composition and ENC was shown using an ENC plot. Most genes did not fall within the standard curve, indicating that the base composition did not affect the codon usage bias (Fig. 6D).
Chloroplast genome codon usage pattern related plot. (A) GC content of different codon sites. (B) Neutrality plot (GC12 against GC3). (C) The codon adaptation index (CAI) value of different function gene sets. (D) Relationship between GC3 and effective number of codons (ENC) (ENC-plot).
RNA editing
Some of the transcription products (RNAs) of the chloroplast genome will be edited during posttranscriptional modification. In seed plants, RNA editing in the chloroplast often converts cytidine into uridine (C to T RNA editing)22. The RNA editing profile under the two distinct stresses, freezing and chilling, was examined using cultivated loquat “JFZ”. Only if the same RNA editing sites were detected in at least 2 of the 3 sample repeats would the RNA editing sites count as a true editing site. In total, 53 RNA editing sites were detected in all the samples. Most RNA editing sites were located in gene bodies (including 30 genes), and 4 RNA editing sites were located in intergenic regions. The rpl2 contained the most RNA editing sites (6 RNA editing sites), followed by petB, atpA, rpoA, and rpl23, which contained 4, 3, 3, and 3 RNA editing sites, respectively (Table. S6). Among the 53 RNA editing sites, 29 were detected in the control group samples, 23 were in the samples under freezing stress group samples, and 50 in samples under chilling stress. The chilling samples contained 19 unique RNA editing sites, and 5 of them are located in the gene rpl2. We then analyzed different time points under chilling stress. We detected 6 time-specific RNA editing sites at 1 h time point, with 4 of them located in rpl2.
RNA editing profiles were also detected across different developmental stages (from young to old) of leaves in the “TY” wild loquat. There were a total of 35 RNA editing sites detected in “TY”. From young leaf to old leaf samples, the number of detected RNA editing sites was gradually increasing, they were 18, 24, and 27, respectively (Table. S7).
Phylogenetic analysis
Protein coding genes from all the six kinds of loquats were used to perform Ka/Ks analysis, Ka/Ks is an index that indicates evolutionary selection. The results showed that two genes, rpl22 and rps12 were under positive selection. Most of the other genes were under purifying selection, and three genes, ndhD (p-value < 0.01), atpA (p-value < 0.01), and rps14 (p-value < 0.05), were under significant purifying selection.
Chloroplast genomes of 13 different kinds of wild loquats, 1 cultivated loquat, Pyrus ussuriensis, Photinia lanuginosa, and Photinia prionophylla were used to construct a phylogenetic tree (Table S8). Plants of Eriobotrya were separated from the other two genera, while among the Eriobotrya, wild loquats and cultivated loquat ‘JFZ’ were divided into three groups. A group contained most of the wild loquats we used, B groups included two E. laoshanica isolates, E. cavaleriei, and cultivated loquat--Eriobotrya japonica ‘JFZ’, C group only contained E. bengalensis. As shown in Fig. 7, the unknown wild loquat “YN-1” is closely related to E. malipoensis (MLP) and E. henryi, followed by E. elliptica (TY) and E. obovata.
Phylogenetic tree, including 13 wild loquats, 1 cultivated loquat, Pyrus ussuriensis, Photinia lanuginosa. and Photinia prionophylla. The phylogenetic tree was constructed using whole chloroplast genome sequences of the 17 species of loquats (raxml-ng with parameters—model TVM+G—bs-trees 1000.).
Materials and methods
Sample sequencing, and chloroplast genome assembly
The leaves of the wild loquats (“YN-1” and “TY”) and cultivated loquat ‘Jiefangzhong’ (JFZ) were collected from trees planted on the Putian University campus located on Longqiao Street, Putian, China. The samples were sent to Biomark Company for DNA extraction, library construction, and second-generation sequencing (Illumina novaseq 6000 paired end sequencing, 150 bp). The chloroplast genomes of both “YN-1” and “TY” were assembled using GetOrganelle v1.7.6.123 with defined parameters and annotated using CPGAVAS224 (based on the ‘43-plastome’ dataset). The results were manually corrected. The final gbf files were uploaded to OGDRAW25 for chloroplast genome map drawing.
Collection of plant material comply with relevant institutional, national, and international guidelines and legislation.
Chloroplast genome comparative analysis
The chloroplast genomes of wild loquats Eriobotrya malipoensis (MLP), Eriobotrya bengalensis (NY), and Eriobotrya fragrans (XH) were downloaded from the Lauraceae Chloroplast Genome Database (https://lcgdb.wordpress.com/). All the chloroplast genomes’ sequences, including the three newly assembled ones, were aligned using the MAFFT26 software (Unweighted Pair Group Method with Arithmetic Mean), The result was then uploaded to DnaSP6.027 for DNA polymorphism analysis. GC content and codon usage bias analysis were performed using EMBOSS-6.6.028 and CodonW software(https://codonw.sourceforge.net/, defined parameter). Codon adaptation index (CAI) was performed on the GALAXY platform29. The mVISTA30 was used to do a comparative genomics analysis. MISA31(misa.ini parmeter: 1–10, 2–5, 3–4, 4–3, 5–3, 6–3) and REPuter32 (minimal repeat size, 18 bp) were used to analyze SSR and repeats. Single nucleotide polymorphisms and indels were detected using DnaSP6.0 and a Python script (https://www.biostars.org/p/119214/). Non-synonymous and synonymous rates of substitution analysis were conducted using ParaAT 2.033 and KaKs_Calculator 2.034. All related figures were drawn using R and OFFICE.
SNP calling for RNA editing site
After being treated to −3 °C (freezing) and −4 °C (chilling), leaf samples from “JFZ” were gathered and sent for RNA-seq in Metware Biotechnology Inc. (Illumina novaseq 6000 paired-end sequencing, 150 bp). SNP calling for RNA editing site detection was performed using RNA-seq data of wild loquat “TY” and cultivated loquat ‘JFZ’. The paired-end reads were mapped to the chloroplast genome using HISAT2 and sorted using samtools35,36. SNP calling was performed using samtools (mpileup and bcftools) and filleted with parameters: quality >= 20 and depth > 10. SNPs in at least two replications were selected as the final RNA editing candidate sites.
Phylogenetic analysis
All the chloroplast genomes of wild loquats, except the 3 members used in the comparison step were downloaded from NCBI database (https://www.ncbi.nlm.nih.gov/), along with chloroplast genomes of Pyrus ussuriensis, Photinia lanuginosa, and Photinia prionophylla. All the chloroplast sequences were aligned using MAFFT, and a phylogenetic tree was constructed using raxml-ng37 with parameters --model TVM+G --bs-trees 1000.
Discussion
Two novel chloroplast genomes, TY and YN-1, were assembled in this research. Their sizes are 159,471 bp and 159,399 bp, respectively. The findings of the gene contents analysis showed that there are 83 protein-coding genes, 8 rRNA, and 36 tRNA in these two wild loquats. This is not the same as the XH loquat, which is devoid of the psbK gene. Although the psbK gene is one of the genes related to PS II, reports suggest that psbK is not crucial for PS II activity in Synechocytis38.
Simple sequence repeats (SSRs), typically consisting of 1 to 6 bp repeat units, are important genetic markers that are widely used in evolutionary and population genetics studies39,40. Here, we conducted an SSR analysis on 6 kinds of loquats, the “XH” loquat contained the highest number of SSRs. Two types of SSRs, “AGC/CTG” and “AAAG/CTTT”, were exclusively identified in “XH”, while “AAT/ATT” was only detected in “TY”. These species-specific SSRs can be used in species differentiation analysis41. In the repetitive sequences analysis, most of the repetitive sequences range from 25 to 50 bp. The “MLP” loquat is an exception; most repetitive sequences contained were more than 70 bp, indicating that rearrangement events may be more intense in “MLP”. It has been reported that repetitive sequences are associated with rearrangement endpoints42.
RNA editing in chloroplast genes has been reported in several studies43,44. There is a report also showing that RNA editing profiles changed under stress situation10. Here, we did detect several RNA editing sites in some genes, using RNA_seq data of cultivating loquat ‘Jiefangzhong’, like rpl2, petB, atpA, rpoA, and rpl23 et al. rpl2 gene seems to play an important role when ‘Jiefangzhong’ response to chilling stress, there were 6 RNA editing sites detected in rpl2. rpl2 is related to plasmid development, and mutant-caused editing defective in rpl2 showed abnormal phenotype during cold stress45,46. RNA editing was also performed in “TY” loquat, results showed that the number of RNA editing sites was gradually increased in leaves from young to mature, and stage-specific RNA editing sites were also found, indicating that RNA editing dynamic in different development stages of leaves. In Coprinopsis cinerea, stage-specific patterns and dynamic RNA editing sites were also detected47.
In the phylogenetic analysis, “TY” (E. elliptica) was close to E. obovata, and then E. henryi. The “YN-1” is also very close to “TY”, indicating a close relationship among these species of wild loquats. Based on these findings, we can infer that “YN-1” is a new wild loquat. “NY” (E. bengalensis) loquat is the only wild loquat that was separated from other wild loquats, suggesting an origin from a different region. There is a report showing that sisterhood exists in E. bengalensis with nine other species of Eriobotrya in their analysis. In their phylogenetic tree, E. bengalensis was also separated from the other species of Eriobotrya they used, located in a solo branch18.
Data availability
The three new assembled chloroplast genomes can be download from link https://db.cngb.org/cnsa/ with accession code CNP0005553
References
Menezes, A. P. A. et al. Chloroplast genomes of Byrsonima species (Malpighiaceae): Comparative analysis and screening of high divergence sequences. Sci. Rep. 8, 1–12 (2018).
Lin, Z. et al. Comparative analysis of chloroplast genomes in Vasconcellea pubescens A.DC. and Carica papaya L. Sci. Rep. 10, 1–12 (2020).
Nock, C. J. et al. Chloroplast genome sequences from total DNA for plant identification. Plant Biotechnol. J. 9, 328–333 (2011).
Li, W., Zhang, C., Guo, X., Liu, Q. & Wang, K. Complete chloroplast genome of Camellia japonica genome structures, comparative and phylogenetic analysis. PLoS One 14, 1–18 (2019).
Amiryousefi, A., Hyvönen, J. & Poczai, P. The chloroplast genome sequence of bittersweet (Solanum dulcamara): Plastid genome structure evolution in Solanaceae. PLoS One 13, 1–23 (2018).
Song, Y., Zhao, W., Xu, J., Li, M. F. & Zhang, Y. Chloroplast Genome Evolution and Species Identification of Styrax (Styracaceae). Biomed Res. Int. 2022, 1–13 (2022).
Gommans, W. M. & Mullen, S. P. RNA editing: A driving force for adaptive evolution?. BioEssays 31, 1137–1145 (2010).
Sasaki, T., Yukawa, Y., Miyamoto, T., Obokata, J. & Sugiura, M. Identification of RNA editing sites in chloroplast transcripts from the maternal and paternal progenitors of tobacco (Nicotiana tabacum): Comparative analysis shows the involvement of distinct trans-factors for ndhB editing. Mol. Biol. Evol. 20, 1028–1035 (2003).
Wang, W., Zhang, W., Wu, Y., Maliga, P. & Messing, J. RNA editing in chloroplasts of Spirodela polyrhiza, an aquatic monocotelydonous species. PLoS One 10, 1–13 (2015).
Chu, D. & Wei, L. Reduced C-to-U RNA editing rates might play a regulatory role in stress response of Arabidopsis. J. Plant Physiol. 244, 153081 (2020).
Wang, Y. A draft genome, resequencing, and metabolomes reveal the genetic background and molecular basis of the nutritional and medicinal properties of loquat (Eriobotrya japonica (Thunb.) Lindl). Hortic. Res. 8, 1–18 (2021).
Peng, Z. et al. Integration of genomics, transcriptomics and metabolomics identifies candidate loci underlying fruit weight in loquat. Hortic. Res. https://doi.org/10.1093/hr/uhac037 (2022).
Wang, Y. & Paterson, A. H. Loquat (Eriobotrya japonica (Thunb.) Lindl) population genomics suggests a two-staged domestication and identifies genes showing convergence/parallel selective sweeps with apple or peach. Plant J. 106, 942–952 (2021).
Su, W. et al. Polyploidy underlies co-option and diversification of biosynthetic triterpene pathways in the apple tribe. Proc. Natl. Acad. Sci. U. S. A. 118, 1–11 (2021).
Su, W. et al. Comprehensive metabolome and transcriptome analyses demonstrate divergent anthocyanin and carotenoid accumulation in fruits of wild and cultivated loquats. Front. Plant Sci. 14, 1–16 (2023).
Qu, S. et al. Complete plastome of a subtropical tree Eriobotrya malipoensis (Rosaceae) in Yunnan. Mitochondrial DNA Part B 4, 3534–3535 (2019).
Chen, S. F. et al. A new species of Eriobotrya (Rosaceae) from Yunnan province, China. PhytoKeys 146, 61–69 (2020).
Xu, J. et al. Whole chloroplast genome sequence of a subtropical tree Eriobotrya bengalensis (Rosaceae). Mitochondrial DNA Part B Resour. 5, 697–698 (2020).
Yan, M. et al. The complete chloroplast genomes of Punica granatum and a comparison with other species in Lythraceae. Int. J. Mol. Sci. 20, 2886 (2019).
Sharp, P. M. & Li, W. H. The codon adaptation index-a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 15, 1281–1295 (1987).
Wright, F. The ‘effective number of codons’ used in a gene. Gene 87, 23–29 (1990).
Xiao, H. et al. A rice dual-localized pentatricopeptide repeat protein is involved in organellar RNA editing together with OsMORFs. J. Exp. Bot. 69, 2923–2936 (2018).
Jin, J. J. et al. GetOrganelle: A fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 1–31 (2020).
Shi, L. et al. CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res. 47, W65–W73 (2019).
Greiner, S., Lehwark, P. & Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: Expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 47, W59–W64 (2019).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302 (2017).
Rice, P., Longden, I. & Bleasby, A. EMBOSS: The European molecular biology open software suite. Trends Genet. 16, 276–277 (2000).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 32, 273–279 (2004).
Beier, S., Thiel, T., Münch, T., Scholz, U. & Mascher, M. MISA-web: a web server for microsatellite prediction. Bioinformatics 33, 2583–2585 (2017).
Kurtz, S. REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res. 29, 4633–4642 (2001).
Zhang, Z. et al. ParaAT: A parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 419, 779–781 (2012).
Zhang, Z. et al. KaKs_calculator: Calculating Ka and Ks through model selection and model averaging. Genomics, Proteomics Bioinforma. 4, 259–263 (2006).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Kozlov, A. M., Darriba, D., Flouri, T., Morel, B. & Stamatakis, A. RAxML-NG: A fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35, 4453–4455 (2019).
Zhang, Z. H., Mayes, S. R., Vass, I., Nagy, L. & Barber, J. Characterization of the psbK locus of Synechocystis sp. PCC 6803 in terms of photosystem II function. Photosynth. Res. 38, 369–377 (1993).
Ahmad, W. et al. Complete chloroplast genome sequencing and comparative analysis of threatened dragon trees Dracaena serrulata and Dracaena cinnabari. Sci. Rep. 12, 1–15 (2022).
Vieira, L. D. N. et al. The complete chloroplast genome sequence of Podocarpus lambertii: Genome structure, evolutionary aspects, gene content and SSR detection. PLoS One 9, 1–10 (2014).
Martin, G., Baurens, F. C., Cardi, C., D’Hont, A. & Aury, J. M. The complete chloroplast genome of banana (Musa acuminata, Zingiberales): insight into plastid monocotyledon evolution. PLoS One 8, e67350 (2013).
Xu, S. et al. Chloroplast genomes of four Carex species: Long repetitive sequences trigger dramatic changes in chloroplast genome structure. Front. Plant Sci. 14, 1–14 (2023).
Duan, Y., Cai, W. & Li, H. Chloroplast C-to-U RNA editing in vascular plants is adaptive due to its restorative effect: testing the restorative hypothesis. RNA 29, 141–152 (2023).
Cui, X. et al. The RNA editing factor DUA1 is crucial to chloroplast development at low temperature in rice. New Phytol. 221, 834–849 (2019).
Liu, X. et al. WSL5, a pentatricopeptide repeat protein, is essential for chloroplast biogenesis in rice under cold stress. J. Exp. Bot. 69, 3949–3961 (2018).
Lan, J. et al. Young leaf white stripe encodes a P-type PPR protein required for chloroplast development. J. Integr. Plant Biol. 65, 1687–1702 (2023).
Xie, Y., Chan, P. L., Kwan, H. S. & Chang, J. The genome-wide characterization of alternative splicing and RNA editing in the development of Coprinopsis cinerea. J. Fungi 9, 915 (2023).
Funding
The study was funded by Fujian Provincial Natural Science Foundation Youth Project, 2022J05254, Putian University scientific research launch project, 2022051, Fujian Provincial University industry-university Cooperation project, 2022N5006, Putian University Putian City Science and Technology Plan Project, 2021ZP09
Author information
Authors and Affiliations
Contributions
Z.C.L. wrote the main manuscript, Z.C.L. and Q.G. performed the chloroplast genome analysis. S.W.M. collected the samples treated with chiling and freezing stress. H.L., S.L., S.L. and J.W. reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, Z., Guo, Q., Ma, S. et al. Chloroplast genomes of Eriobotrya elliptica and an unknown wild loquat “YN-1”. Sci Rep 14, 18816 (2024). https://doi.org/10.1038/s41598-024-69882-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-69882-7