Abstract
In kidney transplant recipients, urine CXCL9 and CXCL10 (uCXCL9/10) chemokines have reached a sufficiently high level of evidence to be recommended by the European Society of Organ Transplantation for the monitoring of immune quiescence. To assess the risk of acute rejection (AR), the advantage of uCXCL9/10 is their cost-effectiveness and their high diagnostic performance. Here, we evaluated the feasibility of a next-generation immunoassay for quantifying uCXCL9/10 levels. It demonstrated high efficiency with minimal workflow and a 90-min time to result. Preanalytical studies indicated stability of uCXCL9/10 levels and analytical studies confirmed excellent linearity and precision. In a cohort of 1048 samples collected at biopsy, the results correlated significantly with ELISA quantification and were integrated into a previously validated 8-parameter urine chemokine model. The next generation immunoassay achieved an accuracy of 0.84 for AR diagnosis. This study validates this technology as a robust, locally available and unexpensive platform and marks a significant step towards the widespread implementation of uCXCL9/10, for immune quiescence monitoring. Therefore, we developed an open-access web application using uCXCL9/10 to calculate AR risk and improve clinical decision-making to perform biopsy, ushering in a new era in kidney transplantation, where personalized, data-driven care becomes the norm.
Similar content being viewed by others
Introduction
The standard of care for monitoring kidney transplant recipients (KTRs) has remained largely unchanged over the years, mostly relying on serum creatinine, proteinuria and anti-human leukocyte antigen (HLA) donor specific antibody (DSA) assessment. Any alarming modification of one or more of these parameters, prompts the decision to perform an allograft biopsy to assess the presence of an ongoing alloimmune injury. The histological evaluation, along with Banff classification, remains our most invaluable tool for routine surveillance, but comes with risk of bleeding, high cost, and requires trained pathologists. Therefore, addressing the shortcomings of current monitoring and developing diagnostic and prognostic biomarkers has been the aim of many studies1,2.
In this drive toward innovative biomarkers, two main lines emerges: research use only investigations and strategies to develop clinical applications. The former often relies on high-throughput techniques using various omics approaches or next-generation sequencing, and has provided novel insights and exciting perspective in disease pathways3,4,5,6,7,8. However, the complexity of these technologies, their cost and the fact that samples have to be sent to centralized, ultra-specialized and delocalized laboratories, makes it less fitted for routine work-up in the near future. The latter takes advantage of noninvasive body fluid collection, and focuses on clinical utility and practical implementation9. In this respect, urine can be regarded as an ideal fluid for kidney allograft assessment, at the same time body waste and condensed reflection of intragraft injuries10.
The use of urine chemoattractant cytokines (chemokines) levels as an integration of all ongoing inflammatory processes within the allograft has been proposed, and further validated in several studies from independent research groups11,12,13,14,15,16. Among them, CXCL9 and CXCL10 have been identified as robust biomarkers that can be measured at the messenger RNA14,16 or protein level11,12,13,15,17. Urinary CXCL9 (uCXCL9) and uCXCL10 essentially appear to have a very high negative predictive value (NPV) for the diagnosis of ongoing and short-term acute rejection (AR)18,19. In addition, we have refined the previously described confounding factors20 in using urinary chemokines as non-invasive markers of AR. As a first, we have built a multiparametric fully-validated model, including uCXCL9/10 and their confounders, which proved its clinical utility21. Besides, we have investigated uCXCL10 as predictive biomarker in the course of BK virus (BKV) reactivation, and we have shown that increased levels were associated with a significant decline in allograft function and an increased occurrence in antibody-mediated rejection (ABMR)22 in BKV viremic KTRs.
Recently, The European Society of Organ Transplantation (ESOT) convened a dedicated working group composed of experts in kidney transplant biomarkers to review literature pertaining to clinical and subclinical acute rejection biomarkers. A total of 298 published articles related to the key words “kidney transplantation, urine chemokine, and acute rejection” were reviewed, and 25 most robust articles were included in the grading process. After discussion and voting during the Consensus Conference (Prague, 2022), a 100% consensus was reached for uCXCL9 and uCXCL10, to recommend (recommendation strength: strong for) their measurement to inform the presence or absence of clinical acute rejection (TCMR or ABMR), and to suggest (recommendation strength: moderate for) their combined measurement to exclude subclinical acute rejection (TCMR or ABMR)23.
While urinary chemokines come to a high level of validation in experimental studies in research labs, clinical implementation requires the availability of robust, fast, reliable and affordable technics2. ELISA quantification appeared to be a good option with robust results, its availability in every hospital laboratory, and its affordable cost. However, in our preliminary study, the used CXCL9 ELISA kit (with a required overnight pre-coating step) raised concerns regarding its use in clinical practice, with a high intra-assay coefficient of variability (CV) and the need of repeating measures in almost 30% of the samples (unpublished data). Thus, we benchmarked the current CXCL9 ELISA landscape to find a pre-coated CXCL9 ELISA kit suitable for automatization and routine follow-up. Among the three candidates identified (none of which had been tested on urine samples), none was found suitable for routine use, most probably due to interference in antibodies fixation from the urine matrix (Supplementary Fig. 1 and Supplementary Results).
To overcome these issues, we investigated the next-generation immunoassay Ella® (ProteinSimple™), considering its alleged ease of use, reliability on urine samples and rapidity in result rendering. The Ella® method, relies on an immunoassay quantification, using cartridges of different formats, which is claimed to have a superior workflow and a higher throughput and quality than ELISA. Each well of the cartridge is sampled three times to give results in triplicate. Each well actually corresponds to a microfluidic channel, directly coated with the capture antibody. The antibodies as well as the fluorescent detector are pre-functionalized in the cartridge, with a built-in standard curve. Of note, some Ella® assays have been tested and validated for quantification from human urine samples, but not CXCL9 and CXCL10 cartridges.
In this study, we evaluated the Ella® platform as a feasible technique for routine implementation of urine chemokine monitoring after kidney transplantation, and validated each workflow step from sample collection to results reporting. More precisely, we investigated preanalytical sample processing from collection to storage and Ella® analytical performance from urine sample, as to provide suggestions of standard operating procedures, an essential factor in ensuring excellent sample quality and reliable results24. For clinical validation, we assessed whether the Ella® platform provided close and useful results in comparison to the reference ELISA technique. Finally, we reasoned that only a web application could advance the use of urine chemokine risk assessment for AR to daily care, and build a web app calculator as a handy tool for clinicians’ use as of now.
Results
Effects of preanalytical sample processing on urine chemokines assessment
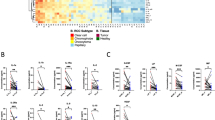
If urinary chemokines are to be used for routine surveillance of KTRs, urine sample collection and processing have to be optimized to fit hospital’s constraints (Fig. 1, Sample collection & storage). In research, protease inhibitors (PI) are usually added after urine supernatant collection to prevent protein degradation upon long-term storage. However, this additional step during sample preparation is time consuming, costly, and might prevent consistent practice between centers. Thus, valid information about necessity of PI addition to prevent CXCL9 and CXCL10 degradation is essential. Chemokine levels of 25 urine samples were compared according to the addition or not of PI during sample preparation. Median time from sample collection to quantification was 147 days [IQR: 127–184]. Figure 2a shows a nearly perfect correlation for CXCL9 (Spearman r = 0.98 [95% CI 0.96–0.99], P < 0.0001), with a mean with/without PI ratio = 1, as well as for CXCL10 (Spearman r = 0.96 [95% CI 0.92–0.98], P < 0.0001), with a mean with/without PI ratio = 1.1.
Overview on clinical workflow steps and research validation studies performed to assess Ella® quantification of urine chemokines: bringing research innovation to daily care. Left panel illustrates the 4-step workflow if urine chemokine were used in daily practice: sample collection and storage, assay preparation, chemokines quantification and acute rejection risk assessment. For each clinical step, the middle panel shows the corresponding validation studies which were conducted, independently from manufacturer’s certification. The right panel presents the web application (https://wise-master.admin.semeia.io/bioptim_computation) that was developed to prompt the use of urine chemokines in daily practice. A screen shot of the user interface is shown, illustrating how clinical and biological data can easily be filled in. Abbreviations: AUC, area under the curve; CV, coefficient of variation; LOQ, limit of quantification; min, minutes; prep, preparation; PI, protease inhibitors; RT, room temperature.
Effects of preanalytical sample processing on urine chemokine assessment. (a) Sample preparation study. Correlation plots illustrating relationships between uCXCL9 (left panel) and uCXCL10 (right panel) measurement from 25 pairs of urine samples, prepared and stored with or without protease inhibitors. P-value and r from a Spearman correlation test. (b, c). Sample storage study. Changes in urinary chemokine concentration over various processing delay (24H, 48H and 72H) and at 4 °C (Panel b) or room temperature (Panel c) storage conditions. Raw data from 5 individual samples are presented as solid lines for CXCL9 (navy blue dots) and for CXCL10 (green triangles). The percentage change was calculated at each timepoint in comparison to reference sample, and mean percentage change at each timepoint is presented by dashed lines (navy blue for CXCL9 and green for CXCL10). RM-ANOVA was performed to assess intra-patient variability. CV was calculated between all 4 urinary chemokine levels and mean CV is given as the mean of all intra-patient CVs. Baseline refers to standard sample processing (within 3H after collection) and storage (freezing at − 80 °C).
Effects of processing delay and storage conditions on urine chemokines assessment
If routinely implemented, urine chemokine assessment might not be available in each single hospital and shipment to a centralized reference center might be considered. Besides, freezing a urine specimen prior to centrifugation may cause cell lysis upon thawing, allowing cellular cytoplasmic protein to contaminate the urine specimen. In research, an early centrifugation step is thus usually performed to pellet cells, but it requires an available technician and a dedicated equipment. Thus, we investigated the influence of time and storage conditions on chemokine quantification. Fresh urine samples from 5 patients were kept at 4 °C or RT for respectively 24, 48 or 72 h. Centrifugation to pellet urine cells and collect urine supernatant was performed immediately before − 80 °C storage.
Within-person stability assessment showed similar results of CXCL9 and CXCL10 levels over time for samples kept at 4 °C (P = 0.26 and P = 0.79, Fig. 2b) or at RT (P = 0.13 and P = 0.51, (Fig. 2c). Up to 72 h at RT, mean intra-patient CV did not exceed 20% for CXCL9 (4 °C, CV = 19.28%; RT, CV = 17.51%) and 15% for CXCL10 (4 °C, CV = 13.12%; RT, CV = 10.41%). Percent change in chemokine level was consistently positive across conditions, indicating a minor increase in chemokine levels upon time. Main variation happened within the first 24 h, suggesting cell lysis from urine cell pellet with adds-on from intracellular chemokines.
Effects of repeated freeze–thaw cycles on urine chemokines assessment
Next, we investigated the influence of repeated freeze–thaw cycles on both chemokines quantified in 5 urine samples (Table 1). In comparison to samples thawed only once (T1), up to 4 additional cycles (T2–T5) did not significantly change within-patient levels of CXCL9 (P = 0.79) and CXCL10 (P = 0.26). The percentage change and CV in CXCL9 and CXCL10 concentration were calculated for each refrozen sample in comparison to the baseline T1 sample. With mean CV of 10.63% for CXCL9 and 18.90% for CXCL10, both assays were found to meet the FDA acceptance criteria for bioanalytical method validation (< 20%, https://www.fda.gov/files/drugs/published/Bioanalytical-Method-Validation-Guidance-for-Industry.pdf). Percent change remained low for both chemokines and was consistently negative for CXCL10, suggesting a more fragile protein, with possible degradation over repeated freeze-thaw cycles.
Linearity and range of measurement
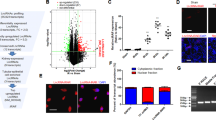
CXCL9 and CXCL10 Ella® cartridges have not been validated on human urine samples. Considering the wide range of pH and urine specific gravity, and that urine complex matrix may hinder immunologic testing, we run an in-house validation of all aspects of analytical performance of the assay (Fig. 1, Chemokines quantification). Ella® cartridges are provided with an internal calibration curve, i.e. a relationship between fluorescence and known concentrations of the analyte. But a calibration curve should be prepared in the same biological matrix as the sample. First, we investigated the ability of the assay to produce results that are directly proportional to the concentration of analyte in the urine sample. Linearity was assessed from 10 samples with a broad range of chemokine values from previous measurement, subjected to serial dilution (1:2, 1:4, 1:8 and 1:16). A high repeatability for each sample was assessed with mean intra-patient CV of 10.2% for CXCL9 and 9.3% for CXCL10 (Fig. 3a). From a Spearman correlation analysis between each dilution factor, all r values were ≥ 0.98 (Fig. 3b). For CXCL10, linearity was confirmed within the complete range given by the manufacturer (dilution-corrected range 1.2–1840 pg/mL). For CXCL9 (manufacturer’s range: 39.8–60,800 pg/mL), linearity was found reliable between 100 and 10,000 pg/mL, but was less clear for extreme values. Hence for sample 10 (Fig. 3a), CXCL9 deviated from 8732 pg/mL (1:8 dilution) to 17464 pg/mL (1:16 dilution, CV = 47.1%, %change = 100). To further define CXCL9 lower and upper limits of quantification (LLOQ-ULOQ) on urine, we used recombinant CXCL9 serially diluted (1:2) into Sample Diluent or into pooled urine from healthy kidney donors, all with undetectable CXCL9 levels (Supplementary Table 1). Recovery at 11 different spiked concentrations (N = 3) showed less reliable CXCL9 assessment below (expected value) 31.3 pg/mL and above 2000 pg/mL (Table 2). Overall, our linearity and recovery data support the following LLOQ and ULOQ on urine sample: 39.8–2000 pg/mL (CXCL9) and 0.6–920 pg/mL (CXCL10).
In-house validation of Ella® assay performance for chemokine quantification from a urine sample. (a, b). Linearity study. (a) Histograms showing uCXCL9 (upper) and uCXCL10 (lower) levels from 10 KTRs urine samples, serially diluted (1:2, 1:4, 1:8 and 1:16) before quantification. For each patient, CV was calculated between each diluted sample and the reference sample (1:2 dilution), and mean CV is given as the mean of all intra-patient CVs. (b) Correlogram illustrating correlation between uCXCL9 (upper) and uCXCL10 (lower) levels after serial dilution for the same 10 KTRs. R values from a Spearman correlation, all P-values < 0.0001. (c, d) ELISA to Ella® correlation study. (c) Correlations between uCXCL9 (N = 1048, navy blue dots) and uCXCL10 (N = 1048, green triangles) levels when measured by ELISA (x-axis) or by Ella® (y-axis) methods. R values from a Spearman correlation, all P-values < 0.0001. (d) Bland-Altman analysis. For each chemokine, Ella®/ELISA ratio (y-axis) is plotting against average chemokine levels (x-axis). Ella®/ELISA ratio of 1 would indicate perfectly superimposable values resulting from both techniques. Bias refers to deviation from this ideal ratio. For representation purpose, only average values < 500 pg/mL are shown. (e) Diagnostic accuracy (C-statistics) of Ella® results tested against reference ELISA results. ROC curves illustrating the diagnostic performance of the 8-parameter chemokine model for any acute rejection, when trained on Ella® or ELISA results (N = 976 samples with no missing value).
Within- and between-run precision
For both assays, precision was assessed on 5 urine samples quantified twice on the same cartridge (intra-assay precision), or quantified twice by different technicians on different days (inter-assay precision). The intra‐assay and inter-assay CVs were 4.7% and 15.3%, respectively for CXCL9, and 2.6% and 16.6%, respectively for CXCL10 (Table 3). For CXCL9, intermediate precision was further refined in a larger number of samples (N = 32) to assess technician-to-technician and day-to-day variations in high, mid and low CXCL9 concentration samples. Under the same set of conditions and within a short interval of time, repeatability ranged from 3.8% (mid CXCL9) to 11.6% (low CXCL9). When investigating the random error introduced by factors like specific technicians, between-run variation was also acceptable with CV ranging from 9.6% (low CXCL9) to 15.3% (high CXCL9). The inter-assay CV for all 37 tested samples averaged 10.3%.
Clinical accuracy study
To evaluate the clinical performances of uCXCL9 and uCXCL10 quantified by Ella®, over 1000 samples belonging to previously published cohorts21,25 were quantified again using the Ella® method. Clinical and biological characteristics of the patients are provided in Supplementary Table 2. We observed a high degree of correlation between uCXCL9 and uCXCL10 measurements by Ella® and by the reference ELISA method (P < 0.0001, Fig. 3c). More specifically, assessments from the two methods were compared using Bland-Altman test (Fig. 3d). For uCXCL10, both methods provided very superimposable values, with uCXCL10 Ella®/ELISA ratio mostly distributed around 1 (Bias = 1.08; 95%CI − 1.61:3.76). For uCXCL9, though highly correlated, numerical values were found higher when quantified by Ella® in comparison to ELISA (Bias = 2.58; 95%CI − 4.04:9.19). Combined with the previous recovery study using recombinant CXCL9, these results suggest that Ella® might provide a more accurate numerical quantification for CXCL9, than ELISA did. Finally, we previously established a model of AR risk using urine chemokines and their confounding factors. Using the same development cohort, we trained the model on Ella®-generated data, instead of ELISA data. Overall, discrimination performance of the 8-parameter model for the diagnostic of AR remained equivalent (AUCElla® = 0.8406, AUCELISA = 0.8481, DeLong’s P-value = 0.49, Fig. 3e). According to Youden’s method, the model cut-off for diagnosing acute rejection is 0.168. At this threshold, sensitivity is 88%, specificity is 68% and above all, negative predictive value is 95%.
The web application: assessment of acute rejection risk using urine chemokines
Our original model was derived in KTRs from Necker Hospital, and validated in an external single-center cohort and in a prospective multicenter unselected cohort21. All samples from these cohorts have since been quantified again by Ella® method, as well as all samples from our princeps study on urinary chemokines25, enabling to train and validate the model on a large set of Ella® data. The resulting model reached an AUC of 0.84 for any rejection diagnosis (Fig. 3e). For clinical assessment of a patient’s risk for AR, and either prompt the decision in performing a biopsy, either argue for avoiding a unnecessary biopsy, we have built an open access web application calculator (BiOptim, Sêmeia, https://wise-master.admin.semeia.io/bioptim_computation). Transplant specialists may easily enter the kidney transplant recipient date of birth and date of urine sample collection and select the recipient gender (Fig. 4a). Then users fill in blood biological data, including the eGFR, the previous identification of DSA and the presence or absence of BKV viremia. Additionally, but not mandatorily, users may provide details concerning the DSA Luminex supplier, the MFI of immuno-dominant DSA or the BK viral load. Finally, users may enter the urine biological data by selecting the presence or absence of a urinary tract infection (symptomatic or asymptomatic, low or upper tract infection), and by filling the urine creatinine, uCXCL9 and uCXCL10 values. Of note, an order of the expected values of uCXCL9 and 10 in these various conditions is shown in Supplementary Fig. 2 as a guide for clinicians. For ease of use, lab test results may be entered in various units with build-in conversion calculation. A screen shot of the results interface is shown in Fig. 4b, computing an individualized risk of rejection according to the 8-parameter model including Ella®-measured urine chemokines. Using the clinical and blood biological data of the simulated patient in Fig. 4a, three distinct urine chemokine patterns were run in the application and their respective acute rejection risk score is depicted, illustrating the additive value of uCXCL9 and uCXCL10 in a DSA positive KTR. The user guide for clinicians then suggests how to interpret the calculated risk score, with the decision to perform a biopsy nevertheless left to the transplant specialist's expertise. Health Care Professionals may register their unit preference for future use as well as a create a patient’s profile, allowing time intervals between score to be graphically displayed and listed. Finally, for flexibility missing data can be imputed by last recorded data (e.g. no recent DSA assessment but patient was always DSA negative), or by mean imputation (e.g. missing BKV viral load imputed by mean viral load).
Building of a web application for clinicians. An open access web application was developed to prompt the use of urine chemokines in daily practice (https://wise-master.admin.semeia.io/bioptim_computation). (a) The web application offers a simple interactive interface. Users may now easily enter their patients’ clinical data (age, gender), serum lab tests (creatinine, DSA and BKV viral load) and urine lab tests (creatinine, uCXCL9 and uCXCL10 levels), and rapidly run the risk score. Example for a 40-year old woman, with a urine chemokine assessment performed in March 2024. She presents with a 55 mL/min eGFR, has a history of DSA but no current blood BK virus replication or UTI. (b) A screen shot of the results interface is shown, computing an individualized risk of rejection according to the 8-parameter model including Ella®-measured urine chemokines. Using the same clinical and blood biological data as in Panel a, 3 distinct urine chemokine patterns were run in the application and their respective acute rejection risk score is depicted, illustrating the additive value of uCXCL9 and uCXCL10 in a DSA positive KTR.
Discussion
In the present study, we aimed to concretize the use of urine chemokines CXCL9 and CXCL10, the monitoring of which is now recommended for KTRs by the European Society of Organ Transplantation, with 2 key objectives: the validation of an automated assay for routine monitoring and the development of a web application for AR risk calculation.
We tested the Ella® platform for CXCL9 and CXCL10 measurement, which has been validated for quantification from plasma, serum and cell culture supernatant but not from urine samples. Our assessment of the Ella® platform from KTR urine sample shows adequate performances with regard to reproducibility, validity, stability after several freeze-thaw cycles, and upon various storage conditions. Results were for the most part reproducible, with laboratory intra- and inter-assay CVs between 3 and 17%, thus meeting the FDA acceptance criteria for bioanalytical method validation26. More importantly, chemokines quantified by Ella® showed similar performance to ELISA in terms of diagnostic accuracy for acute renal allograft rejection.
During sample collection and storage, concerns about degradation are significant. We investigated the impact of various storage conditions and observed that urine chemokine levels remained stable after multiple freeze-thaw cycles and storage at − 80 °C. Interestingly, immediate centrifugation or protease inhibitor addition may not be imperative for sample processing, simplifying the pre-analytical steps.
To investigate the feasibility of clinical implementation of Ella® quantification for urine proteins, we compared ELISA and Ella® workflow, from sample thaw to render of the results. Upon assay preparation, Ella® appeared superior to conventional ELISA with no plate coating and no reagent preparation. Sample preparation only included thawing and an additional centrifugation step as to pellet all debris which might cause microfluidics and glass nanoreactors obstruction (Fig. 1, Assay preparation). Ella® procedure further included a dilution step as for ELISA and a simple one-step sample deposition within the cartridge prior to running the assay (20 min). Once launched, time to result is approximately 70 min. Altogether, the estimated Ella® assay procedure is 1 h 30 as compared to 7 h for a conventional ELISA (let alone antibody coating the day before for uncoated CXCL9 ELISA kit). In the case of acute renal failure, the clinician can retrieve the chemokine assay in less than 2 h to make a decision.
After assessing that Ella® assay provided close results, in comparison to the reference ELISA technique, we assessed whether it provided useful results, i.e. similar diagnostic performance to assess the risk of AR. Although not identical, the high correlation between values from ELISA and Ella® ensured that individuals were ranked comparably by both methods, which is critical risk assessment models. Hence in our cohort of 1048 samples, training the 8-parameter model on Ella®-data lead to similar accuracy, enabling the building of a web application calculator for clinicians to use. This technology recently provided very similar results in a large cohort of 1559 urine samples, with a 5-parameter model yielding an AUC of 0.81 for the AR diagnosis27.
Moreover, the widespread adoption of Ella™ technology in hospitals worldwide, particularly in response to the challenges posed by the Covid-19 pandemic, advocates the local feasibility of implementing our novel approach on a global scale. Endorsement of chemokine measurement by esteemed organizations such as the European Society for Organ Transplantation (ESOT) further underscores the transformative potential of our research28.
It is imperative to emphasize that our article represents more than just a scientific contribution—it heralds a new era in renal transplantation, one where personalized, data-driven care becomes the standard. This shift in clinical practice is a rarity in the medical field and deserves recognition for its potential to significantly improve patient outcomes and redefine the boundaries of nephrological care.
Methods
Study samples and cohorts
Samples for the clinical validation study belong to previously published cohorts29,30 and comprised N = 1082 samples collected at the time of a for cause (71%) or a surveillance (29%) biopsy. The study adheres to the Declaration of Helsinki, was approved by the Ethics Committee of Ile-de-France XI (#13016), and all participating patients provided written informed consent. The clinical and research activities being reported are consistent with the Principles of the Declaration of Istanbul as outlined in the 'Declaration of Istanbul on Organ Trafficking and Transplant Tourism.
ELISA methods for uCXCL9/CXCL10
We previously described the extensive protocols for urinary chemokines quantification by enzyme-linked immunosorbent assay (ELISA). Briefly, uCXCL9 was measured using Human CXCL9/MIG DuoSet ELISA kit (Bio-Techne, Minneapolis, USA), with a protocol optimized for quantification from a urine sample. Human CXCL10/IP10 Quantikine ELISA Kit (Bio-Techne,) was used according to the manufacturer’s instructions. Optical densities were measured using a Multiskan FC plate reader (Thermo Fisher, Illkirch, France). All measurements were performed in duplicate.
Samples for Ella® technical validation
Samples for Ella® technical validation (preanalytical and analytical performance studies) were all taken from the local Transplantation Biobank (DC-2009-955, Necker Hospital, Paris, France). Urine samples were routinely collected from kidney transplant recipients as part of transplant care, with exception for the “storage study” where freshly emitted urine samples (N = 12) were prospectively collected. Besides, urine samples (N = 10) were collected from kidney living-donors (KD) at the time of their pre-donation evaluation.
Ella® immunoassay methods
Urine CXCL9 and CXL10 levels were measured using the Ella® microfluidic Single Plex cartridges (ProteinSimple™, Bio-Techne, Noyal Châtillon sur Seiche, France), following the manufacturers’ instructions. Briefly, urine samples were stored frozen at − 80 °C, thawed on ice, then centrifugated at 1500 relative centrifugal force (g) for 2 min, as to pellet all debris which might cause microfluidic channel obstruction. For Single Plex cartridge loading, 50 µL of each diluted urine supernatant sample (1:1 in Sample Diluent) or quality control was added to the wells, as well as 1 mL of Wash Buffer in the dedicated inlet. The automated Ella® immunoassay protocol was then initiated, including automated three times sampling of each well to give results in triplicate. Measurement of creatinine was performed in the same urine samples using the Creatinine Parameter Assay Kit (Bio-Techne).
Sample preparation study
As part of the local routine biobanking (reference method here), urine samples are collected and processed as follows: samples are kept at room temperature (RT) until centrifuged at 3300 rpm for 20 min at 4 °C within 3 h of collection. The supernatant is collected and stored with or without protease inhibitors (cOmplete™, Roche Diagnostics, Meylan, France) at − 80 °C. In this Sample Preparation Study, both aliquots of 25 urine samples were thawed on ice and urinary chemokines were quantified by Ella® technique in a single batch. Chemokine levels in each sample were compared according to the addition or not of protease inhibitors during sample preparation.
Clinical accuracy study
To evaluate the diagnostic performances of urinary CXCL9 and CXCL10 quantified by Ella® as compared to the reference ELISA method, we used all available urine samples from four previously published cohorts29,30. Details of quantification is provided in Supplementary Material. Among the 1123 urine samples included in the final analyses of the original publications, 1084 corresponded to unique urine samples, and enough material was available for Ella® quantification for 1048 of those. ELISA and Ella® results were compared using Spearman correlation tests and Bland-Altman analysis. These results were further used to investigate how the modification in quantification method might influence the performance of our 8-parameter chemokine model. All samples with no missing data for all 8 variables (N = 976) comprising N = 225 acute rejection (23%) were used to build several risk prediction models using logistic regression and random forest analyses. Accuracy was assessed by mean of an Area Under the recipient operating Curve (AUC).
Statistical analyses
Patient and donor characteristics are described by numbers, percentages and frequencies for categorical variables. We report continuous variables using means and standard deviations (SD) or medians and interquartile ranges (IQR) for variables with a skewed distribution. Changes in concentrations of chemokines over time, temperature or freeze-thaw cycles were analyzed using one-way repeated measures analysis of variance (RM-ANOVA) followed by Sidak’s multiple comparisons tests. AUC’s were compared with the Delong’s test. Statistical analyses were performed using Graphpad Prism version 9.0.1 (GraphPad Software, San Diego, USA) and with R software (R Development Core Team, R studio version 2022.12.0+353).
Data availability
All data produced in the present study are available upon reasonable request to the authors. An open access web application was developed to prompt the use of urine chemokines in daily practice (https://wise-master.admin.semeia.io/bioptim_computation).
References
Naesens, M. & Anglicheau, D. Precision transplant medicine: Biomarkers to the rescue. J. Am. Soc. Nephrol. 29, 24–34 (2018).
Anglicheau, D., Naesens, M., Essig, M., Gwinner, W. & Marquet, P. Establishing biomarkers in transplant medicine: A critical review of current approaches. Transplantation 100, 2024–2038 (2016).
Halloran, P. F. et al. Review: The transcripts associated with organ allograft rejection. Am. J. Transplant. 18, 785–795. https://doi.org/10.1111/ajt.14600 (2018).
Mas, V. R., Dumur, C. I., Scian, M. J., Gehrau, R. C. & Maluf, D. G. MicroRNAs as biomarkers in solid organ transplantation. Am. J. Transplant 13, 11–19 (2013).
Bontha, S. V., Maluf, D. G., Mueller, T. F. & Mas, V. R. Systems biology in kidney transplantation: The application of multi-omics to a complex model. Am. J. Transplant. 17, 11–21 (2017).
Tinel, Claire et al. Modulation of monocyte response by microrna-15b/106a/374a during antibody-mediated rejection in kidney transplantation. Transplantation 107(5), 1089–1101. https://doi.org/10.1097/TP.0000000000004393 (2023).
Tinel, C. et al. Integrative omics analysis unravels microvascular inflammation-related pathways in kidney allograft biopsies. Front. Immunol. 12, 738795 (2021).
Van Loon, E. et al. Biological pathways and comparison with biopsy signals and cellular origin of peripheral blood transcriptomic profiles during kidney allograft pathology. Kidney Int. https://doi.org/10.1016/j.kint.2022.03.026 (2022).
Naesens, M., Friedewald, J., Mas, V., Kaplan, B. & Abecassis, M. M. A practical guide to the clinical implementation of biomarkers for subclinical rejection following kidney transplantation. Transplantation 104, 700–707. https://doi.org/10.1097/TP.0000000000003064 (2020).
Mannon, R. B. Immune monitoring and biomarkers to predict chronic allograft dysfunction. Kidney Int. Suppl. 78, S59-65. https://doi.org/10.1038/ki.2010.425 (2010).
Hu, H. et al. Elevation of CXCR3-binding chemokines in urine indicates acute renal-allograft dysfunction. Am. J. Transplant. https://doi.org/10.1111/j.1600-6143.2004.00354.x (2004).
Jackson, J. A. et al. Urinary chemokines CXCL9 and CXCL10 are noninvasive markers of renal allograft rejection and BK viral infection. Am. J. Transplant. 11, 2228–2234 (2011).
Hirt-Minkowski, P. et al. Detection of clinical and subclinical tubulo-interstitial inflammation by the urinary CXCL10 chemokine in a real-life setting. Am. J. Transplant. 12, 1811–1823 (2012).
Suthanthiran, M. et al. Urinary-cell mRNA profile and acute cellular rejection in kidney allografts. N. Engl. J. Med. 369, 20–31 (2013).
Rabant, M. et al. Urinary C-X-C motif chemokine 10 independently improves the noninvasive diagnosis of antibody-mediated kidney allograft rejection. J. Am. Soc. Nephrol. 26, 2840–51 (2015).
Sigdel, T. K. et al. A urinary common rejection module (uCRM) score for non-invasive kidney transplant monitoring. PLoS One 14, e0220052 (2019).
Blydt-Hansen, T. D. et al. Validity and utility of urinary CXCL10/Cr immune monitoring in pediatric kidney transplant recipients. Am. J. Transplant. 21, 1545–1555 (2021).
Rabant, M. et al. Early low urinary CXCL9 and CXCL10 might predict immunological quiescence in clinically and histologically stable kidney recipients. Am. J. Transplant. 16, 1868–81 (2016).
Hricik, D. E. et al. Multicenter validation of urinary CXCL9 as a risk-stratifying biomarker for kidney transplant injury. Am. J. Transplant. 13, 2634–2644 (2013).
Ho, J. et al. Urinary CXCL10 chemokine is associated with alloimmune and virus compartment-specific renal allograft inflammation. Transplantation https://doi.org/10.1097/TP.0000000000001931 (2018).
Tinel, C. et al. Development and validation of an optimized integrative model using urinary chemokines for noninvasive diagnosis of acute allograft rejection. Am. J. Transplant. https://doi.org/10.1111/ajt.15959 (2020).
Tinel, C. et al. Deciphering the prognostic and predictive value of urinary CXCL10 in kidney recipients with BK virus reactivation. Front. Immunol. 11, 604353 (2020).
Park, S. et al. European society of organ transplantation consensus statement on testing for non-invasive diagnosis of kidney allograft rejection. Transpl. Int. 36, 12115 (2023).
Sturgeon, C., Hill, R., Hortin, G. L. & Thompson, D. Taking a new biomarker into routine use–a perspective from the routine clinical biochemistry laboratory. Proteomics Clin. Appl. 4, 892–903 (2010).
Rabant, M. et al. Early low urinary CXCL9 and CXCL10 might predict immunological quiescence in clinically and histologically stable kidney recipients. Am. J. Transplant. 16, 1868–1881 (2016).
U.S. Food and Drug Administration. Bioanalytical Method Validation. Guidance for Industry. https://www.fda.gov/files/drugs/published/Bioanalytical-Method-Validation-Guidance-for-Industry.pdf (2018).
Van Loon, E. et al. Automated urinary chemokine assays for noninvasive detection of kidney transplant rejection: A prospective cohort study. Am. J. Kidney Dis. 83, 467–476 (2024).
Park, S. et al. European society of organ transplantation consensus statement on testing for non-invasive diagnosis of kidney allograft rejection. Transplant. Int. https://doi.org/10.3389/ti.2023.12115 (2024).
Rabant, M. et al. Urinary C-X-C motif chemokine 10 independently improves the noninvasive diagnosis of antibody-mediated kidney allograft rejection. J. Am. Soc. Nephrol. 26, 2840–2851 (2015).
Tinel, C. et al. Development and validation of an optimized integrative model using urinary chemokines for noninvasive diagnosis of acute allograft rejection. Am. J. Transplant. 20, 3462–3476 (2020).
Acknowledgments
We are indebted to all Necker Hospital teams and patients who participated in this research. We thank Joffrey Alves-Gasnier, Xavier Lebreton, Charlotte Leclaire, Lise Morin, Corinne Normand, Juliette Olivré and Laïla Sahli for their technical assistance. CT is supported by the Société Francophone de Transplantation. The Emmanuel Boussard Foundation supported DA, LA, LM, CT and VS. DA received funds from the Day Solvay Foundation.
Funding
CT is supported by the Société Francophone de Transplantation. The Emmanuel Boussard Foundation supported DA, LA, LM, CT and VS. DA received funds from the Day Solvay Foundation.
Author information
Authors and Affiliations
Contributions
C.T. and D.A. conceived and designed the study. C.L. provided the urine samples. C.T., L.A. and M.R. collected the clinical data. V.S. and C.T. carried out the experiments. M.R. performed the histology reading of biopsies at Necker Hospital. B.L. participated in the creation of the figures. C.T. performed the statistical analyses and interpreted the data. C.L. and F.T. participated in the performance of the research. C.T. and D.A. wrote the draft of the report. All the authors revised the report.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Tinel, C., Sauvaget, V., Aouni, L. et al. Transforming kidney transplant monitoring with urine CXCL9 and CXCL10: practical clinical implementation. Sci Rep 14, 20357 (2024). https://doi.org/10.1038/s41598-024-70390-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-70390-x