Abstract
Attention Deficit Hyperactivity Disorder (ADHD) is characterized by deficits in attention, hyperactivity, and/or impulsivity. Resting-state functional connectivity analysis has emerged as a promising approach for ADHD classification using resting-state functional magnetic resonance imaging (rs-fMRI), although with limited accuracy. Recent studies have highlighted dynamic changes in functional connectivity patterns among ADHD children. In this study, we introduce Skip-Vote-Net, a novel deep learning-based network designed for classifying ADHD from typically developing children (TDC) by leveraging dynamic connectivity analysis on rs-fMRI data collected from 222 participants included in the NYU dataset within the ADHD-200 database. Initially, for each subject, functional connectivity matrices were constructed from overlapping segments using Pearson’s correlation between mean time series of 116 regions of interest defined by the Automated Anatomical Labeling (AAL) 116 atlas. Skip-Vote-Net was then developed, employing a majority voting mechanism to classify ADHD/TDC children, as well as distinguishing between the two main subtypes: the inattentive subtype (ADHDI) and the predominantly combined subtype (ADHDC). The proposed method was evaluated across four classification scenarios: (1) two-class classification of ADHD from TD children using balanced data, (2) two-class classification between ADHD and TD children using unbalanced data, (3) two-class classification between ADHDI and ADHDC, and (4) three-class classification among ADHDI, ADHDC, and TD children. Using Skip-Vote-Net, we achieved mean classification accuracies of 97% ± 1.87 and 97.7% ± 2.2 for the balanced and unbalanced classification cases, respectively. Furthermore, the mean classification accuracy for discriminating between ADHDI and ADHDC reached 99.4% ± 1.21. Finally, the proposed method demonstrated an average accuracy of 98.86% ± 1.03 in classifying ADHDI, ADHDC, and TD children collectively. Our findings highlight the superior performance of Skip-Vote-Net over existing methods in the classification of ADHD, showcasing its potential as an effective diagnostic tool for identifying ADHD subtypes and distinguishing ADHD from typically developing children.
Similar content being viewed by others
Introduction
Attention Deficit/Hyperactivity Disorder (ADHD) is a prevalent mental disorder which affects children and primarily caused by genetics and environmental factors1. It is estimated that 8–11% of school-age children are affected by ADHD and 30–50% of them keep ADHD symptoms in their adulthood2,3,4,5,6. Children with ADHD struggle to maintain focus on tasks, which can lead to challenges in academic success, social interactions, and professional achievement4,5,6. This multifaceted disorder is defined by enduring deficits in attention, hyperactivity, and/or impulsivity. ADHD is categorized into three subtypes: a rarely identified hyperactive-impulsive subtype (ADHDH), an inattentive subtype (ADHDI), and a predominantly combined subtype (ADHDC) characterized by both inattention and hyperactivity, as delineated in the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV)5,7.
The diagnosis process based on symptoms relies on the experience of clinicians, could cause biases and may result in delays in treatment initiation8,9,10,11,12,13. Thus, integration of machine learning (ML) methods can significantly reduce the duration of the diagnostic process, thereby enhancing efficiency and enabling timely medical interventions.
Advanced neuroimaging modalities, such as structural and functional magnetic resonance imaging (sMRI/fMRI)14 serve as valuable sources of biomarkers for diagnosing ADHD. Functional connectivity networks (FCNs) derived from resting-state functional magnetic resonance imaging (rs-fMRI) data can be used to investigate alterations in interconnection between various brain regions. Recent investigations have underscored the efficacy of FC in elucidating disparities between individuals with ADHD and typically developing counterparts15,16.
Numerous ML methodologies have been proposed for ADHD classification based on brain functional connectivity (FC), broadly categorized into classical and deep learning (DL) approaches. Classical methods typically follow a sequential procedure comprising feature extraction, feature selection, and decision via classifiers. Notable among these are methodologies such as t-test analysis17, support vector machine with recursive feature elimination (SVM–RFE)18,19, and least absolute shrinkage and selection operator (LASSO)20,21. However, classical classification methods are burdened by inherent complexities, including the need for extensive knowledge about characteristic features of different ADHD subtypes, which significantly affects the accuracies of these methods, ranging between 65 and 85%22,23,24.
In recent years, representation learning has become a prominent area of focus in classification, leading to advancements in DL, especially convolutional neural networks (CNNs). This obviates the necessity for explicit data transformation, instead enabling the implicit representation of subtype-specific characteristics. Numerous studies have explored the efficacy of three- and four-dimensional CNN models based on the spatial and spatio-temporal characteristics of ADHD traits using sMRI and fMRI data10,25 for automatic ADHD classification9,26,27. Among the DL tools, auto-encoding and attention networks have been shown to better capture discriminative high-level features from rs-fMRI data, resulting in higher classification accuracy28,29.
Thus far, the DL methodologies utilized for ADHD classification have primarily relied on static functional connectivity (sFC), involving the derivation of FCNs spanning the entire scan durations. However, emerging evidence suggests that dynamic changes in brain FCNs offer a more comprehensive understanding of the underlying pathophysiology of ADHD4. Dynamic functional connectivity (dFC) allows us to capture the dynamic changes in FC over time, providing valuable insights into the complex interactions between brain regions. In essence, dFC focuses on capturing functional alterations within brain networks over brief intervals, thereby potentially enhancing the accuracy of classification algorithms. Additionally, while the majority of existing methodologies classify subjects into broad categories of normal or ADHD, discerning between different subtypes of ADHD, such as ADHDI and ADHDC, is pertinent.
In this investigation, we present a novel DL approach called Skip-Vote-Net for the automatic classification of ADHD subtypes and typically developing children (TDC) utilizing dFC. We hypothesize that dFC encompasses distinctive patterns facilitating a superior discrimination between ADHD subjects and TDC, along with ADHD subtypes. Within the Skip-Vote-Net framework, skip connections were integrated to increase convergence rates and improve the network’s efficacy. Additionally, we explored the potential enhancement in the network’s generalization capabilities by balancing the data distribution across classes.
Materials and methods
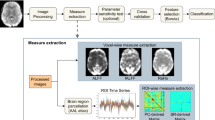
Figure 1 depicts the processing pipeline employed for ADHD classification, comprising three main steps: (i) preprocessing of rs-fMRI data, (ii) dynamic functional connectivity analyses, and (iii) ADHD classification using the proposed Skip-Vote-Net. The Skip-Vote-Net was utilized to categorize each subject into different states, involving: (i) classification of ADHD and TDC children using balanced training data, (ii) classification of ADHD and TDC children using unbalanced training data, (iii) classification of ADHD children into ADHDI and ADHDC, and (iv) classification of subjects into ADHDI, ADHDC, and TDC.
Database
For method development and evaluation, we utilized rs-fMRI data collected at the New York University (NYU) Child Study Center, which is part of the publicly available ADHD-200 repository30. This repository includes several hundred rs-fMRI scans gathered from eight distinct acquisition sites, each operating under its own recording protocols. The local Internal Review Board approved the recording protocols at each site. To maintain consistency and mitigate potential discrepancies related to differences in recording protocols, we included only the training set of the imaging data from the NYU center. The training set comprised rs-fMRI data from 222 participants, including 44 individuals with inattentive ADHD (aged 7–18), 77 individuals with combined ADHD (aged 7–18), 2 individuals with hyperactive ADHD, and 99 typically developing children (aged 7–18). Due to the insufficient number of subjects, the hyperactive ADHD class was excluded from the classification analysis. Additionally, in the NYU database, some participants had two resting-state fMRI scans. To enhance data robustness and improve the generalizability of our model, each scan was considered as a separate subject (see supplementary materials for further details).
The data have been collected using a 3-T Siemens Allegra scanner. Each participant’s data includes a high-resolution T1-weighted (T1w) scan, as well as at least one resting-state eyes closed T2*-weighted (BOLD-weighted) multi-echo scan. The T1w images have been obtained through a magnetization prepared, rapid-acquisition gradient-echo sequence with a voxel size of 1.3 mm×1 mm×1.3 mm. Each subject underwent approximately 6 min of rs-fMRI using a gradient-echo echo planar imaging (EPI) sequence with a repetition time (TR) of 2 s and a voxel size of 3 mm×3 mm×4 mm. We analyzed 176 functional volumes from each subject to match data quantity consistency across all participants30.
Data preprocessing
In the preprocessing step, the T1-weighted images of each subject were initially normalized to the standard structural space template, Montreal Neurological Institute (MNI152)31, employing both affine registration (FLIRT)32,33 and non-linear registration (FNIRT)34 tools. Then, skull stripping was performed using AFNIs 3dSkullStrip, followed by automated tissue-type segmentation applying FAST35 to obtain a high-resolution mask (1 mm isotropic) for brain tissues consisting of cerebrospinal fluid (CSF), white matter (WM), and gray matter (GM). To match the rs-fMRI data, low-resolution masks (4 mm isotropic) are generated by AFNI36.
For rs-fMRI preprocessing, the first four volumes were initially removed from each subject’s data. Subsequently, AFNI’s 3D DESPIKE was used to eliminate large transient fluctuations based on the median absolute deviation. Slice-timing correction was performed based on the middle slice, using AFNI’s 3dTshift, followed by rigid registration conducted with AFNI’s 3dvolreg to estimate subject movement parameters. The rs-fMRI images were then realigned and co-registered to structural data and normalized into the MNI atlas space31. The images were resliced to 4 mm×4 mm×4 mm voxels and smoothed with a Gaussian kernel with 6 mm full-width at half maximum. A temporal band pass filter with a frequency band of 0.009–0.1 Hz was used for noise attenuation36. Finally, linear detrending was applied to reduce the effects of noise and signal drift.
After preprocessing, 44 out of the 220 participants were excluded from further analysis due to excessive motion artifacts, defined as a displacement of more than 3.0 mm and a rotation index higher than 3.0°. Consequently, 176 individuals were included in the classification analysis, comprising 42 individuals with inattentive ADHD, 66 individuals with combined ADHD, and 68 typically developing children. Since some individuals had two scans, a total of 75 scans from individuals with inattentive ADHD, 91 scans from individuals with combined ADHD, and 99 scans from typically developing children were used for training and performance evaluation.
Processing pipeline used for ADHD classification using Skip-Vote-Net, including rs-fMRI preprocessing, dynamic connectivity analysis (P represents the number of parcels), and ADHD/TDC classification using Skip-Vote-Net.
Static and dynamic functional connectivity analysis
For the functional connectivity analysis, we utilized the Automated Anatomical Labeling (AAL) atlas37 to partition the brain into 116 regions-of-interest (ROIs). The mean time series of each region were extracted by averaging the preprocessed data of all voxels within the region. For the sFC analysis, the connectivity matrix was computed over the entire scan duration (herein 6 min) by computing the pair-wise Pearson correlation coefficients between the time courses of the brain regions. To construct the dFC matrix38 from the preprocessed rs-fMRI data, we employed a sliding window with a step size to partition the rs-fMRI data into K segments. K was chosen as [5, 7, or 9] by adjusting the window length with a 50% overlap. For instance, with seven segments, the window length was set to 44 TR (88s) with a step size of 22 TR (44s). Subsequently, K dFC matrices were generated for each subject and utilized to assess the performance of the Skip-Vote-Net.
Proposed Skip-Vote-Net
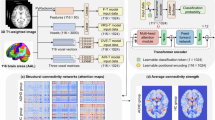
The proposed Skip-Vote-Net architecture was designed to extract the most relevant features. Figure 2 illustrates the Skip-Vote-Net, which is composed of three main components: convolutional, fully connected, and majority voting layers. Skip connections were used in the proposed model to increase the convergence rate by allowing gradients to flow more easily through the network. These connections also enabled the network to operate at its optimal level, resulting in improved efficiency and accuracy. The term “Vote” refers to the majority voting strategy used to enhance the accuracy of the proposed model at the subject level by combining the predictions from subjects’ segments.
The convolutional layers serve as the feature extraction step and were primarily comprised of four Skip-Blocks and a concatenation layer. The first three Skip-Blocks sequentially extract features, and the output features of each one were concatenated after passing through a max-pooling layer to reduce the size of the feature maps. The skip connections as illustrated in Fig. 3 were employed to increase the convergence rate and enhance the network’s overall performance39,40.
Within the convolutional network, the fully connected layer plays a crucial role in determining the final classification outcome for each segment. This layer takes the output features of the last pooling layer to make a prediction. Since there were K dFC matrices for each subject, the predicted classification labels were used to classify the subject into one class through the majority voting strategy. We utilized the hyperbolic tangent function, \(\:\sigma\:\left(x\right)=tanh\left(x\right)\), as the activation function in each layer to incorporate both positive and negative correlation values in the classification process.
Skip-Vote-Net architecture. P represents the number of parcels (i.e. 116, resulting in floor(\(\:\frac{P}{3}\)) = 38), and \(\:x\) was set to 4.
Skip-Block
Figure 3 illustrates the architecture of the Skip-Block, comprised of a detail path, an approximation path, and an element-wise sum component. The detail path augments the number of feature maps while preserving a combination that accentuates the key feature elements of the original input. Meanwhile, the approximation path enables the network to extract primary features, incorporating neighboring information. The incorporation of element-wise summation into these paths enhances the network’s ability to capture discriminative features.
Skip-Block Architecture. F and P represent the number of filters and the number of parcels (herein 116), respectively.
Max pooling, skip connection and concatenation
MaxPooling and skip connections (Fig. 4) can have a significant impact on the accuracy of the model. In the proposed Skip-Vote-Net, MaxPooling was used to down-sample feature maps and reduce the spatial dimensions of the input features, thereby decreasing the number of parameters and computation40,41. Skip-Blocks 1 to 3 were downsampled using MaxPooling with a kernel size of \(\:(3\times\:3)\), and the output feature maps of Skip-Block 4 were downsampled with a kernel size of (\(\:4\times\:4\)).
Skip connections provide alternative pathways for gradients to flow through the network during backpropagation, helping mitigate the vanishing gradient problem. This problem arises when gradients become increasingly small as they propagate through many layers, making it difficult for the network to effectively learn discriminative features42. Furthermore, skip connections enable the training of deeper networks. In the proposed architecture, both max pooling and skip connections were utilized in the first three Skip-Blocks.
Max pooling, skip connection and Concatenation layers. F and P represent the number of filters and the number of parcels (herein 116, resulting in floor(\(\:\frac{P}{3}\)) = 38), respectively.
Majority voting
The majority voting strategy was employed to classify each subject into either ADHD or TDC in the two-class mode, based on the majority of predicted labels assigned by the fully connected layer for the subject’s segments. In the 3-class mode, each subject was initially assigned ADHD or TDC based on the majority voting. If the assigned class was ADHD, then, using the same majority voting approach, the subtype class was determined between ADHDI and ADHDC.
Model validation
The proposed Skip-Vote-Net architecture was evaluated on four classification modes: (i) ADHD/TDC with unbalanced folds, (ii) ADHD/TDC with balanced folds, (iii) ADHDI/ADHDC, and (iv) ADHDI/ADHDC/TDC. For this purpose, the dataset was randomly partitioned into five subsets, both balanced and unbalanced, to conduct five-fold cross-validation (refer to Table 1). In the balanced folds, subjects were randomly selected to ensure an equal number of subjects for ADHD/TDC groups. This methodology enabled us to train our model on four subsets and evaluate its performance on the remaining subset. The average performance was then assessed for each mode (see Supplementary Materials for further details on the subjects selected for each fold and classification mode).
As described in Sect. 2.2, for data augmentation, both scans were included for individuals with two scans. In the “Augmented Data Mode”, there were instances where scans from the same individuals were distributed between the training and test sets. This could potentially introduce data leakage during performance evaluation due to the dependency between scans of the same subjects. To assess the impact of this leakage on the performance of the SVM and Skip-Vote-Net models, we conducted additional evaluations. Specifically, we removed the second scans from the test set in each fold, referred to as the “Baseline Data Mode”, to mitigate the potential leakage effect.
Network implementation
TensorFlow was used to implement the Skip-Vote-Net model, employing cross-entropy as the loss function for the two-class classification mode as:
where N was the number of subjects in training, \(\:{y}_{i}\) denotes the true label for the \(\:{i}_{th}\) training subject (0 for ADHD and 1 for TDC), and \(\:{\stackrel{\prime }{y}}_{i}\) represents the prediction made by Skip-Vote-Net. The cross-entropy loss function evaluates the class prediction of individuals by comparing the probability distribution of the actual and predicted labels. When the distributions are similar, the value of the cross-entropy is small; conversely, it is large when they differ significantly. The three-class classification was performed by utilizing a cross-entropy loss as follows:
where M was set to 3, \(\:{y}_{ij}\) represents the true label of the \(\:{i}_{th}\) training subject belonging to the \(\:{j}_{th}\) class, and \(\:{p}_{ij}\) is the probability of the \(\:{ij}_{th}\) subject. This allowed us to effectively handle the multi-class classification task and optimize the model’s performance across all classes. Additionally, we utilized the Adam optimizer, known for its efficiency in optimizing DL models, to further enhance the training process and improve convergence.
For each classification mode, the batch size, learning rate, and the maximum number of epochs were set to 8, 10− 5, and 100, respectively. Additionally, we employed the following strategies to mitigate overfitting: (i) early stopping with a patience parameter of 10, monitored based on validation accuracy; (ii) a dropout rate of 0.3; and (iii) L2 regularization with a penalty value of 10− 3.
Static FC-based classification
To evaluate the efficacy of the dynamic approach, we performed a comparative analysis against standard SVM-based classification applied to sFC. We employed the SVM classifier with an RBF kernel due to its superior efficiency relative to alternative variants. Each subject’s data was represented as a single vector derived from the upper-triangle elements of the sFC matrix. Subsequently, the SVM classifier was evaluated using the same cross-validation folds established in Sect. 2.5. Notably, the Skip-Vote-Net method was not viable for classification with sFC matrices due to its intricate architecture and limited input sample size, preventing network convergence.
Parameter selection
To optimize classification accuracy, we determined the optimal Skip-Vote-Net configuration by varying the number of filters across layers within predefined ranges for the unbalanced two-class (ADHD/TDC) classification. The numbers of filters for the first through fourth Skip-Blocks were selected from the following sets: [32, 64, 128, 256], [64, 128, 256, 512], or [128, 256, 512, 1024]. We employed a five-fold cross-validation strategy on segments, utilizing all dFC segments from subjects selected for model validation (Sect. 2.5) for both training and testing. The optimal network configuration identified through this process was then applied to subject-level classification evaluations, using the majority voting strategy.
Evaluation metrics
The classification performance of the proposed method was evaluated using accuracy (ACC), defined as:
where a true positive (TP) refers to a correct classification as ADHD, a false positive (FP) indicates a wrongful classification as ADHD, a true negative (TN) signifies a correct classification as TDC, and a false negative (FN) denotes a wrongful classification as TDC.
We further compared the performance of the proposed Skip-Vote-Net model with that of the SVM with an RBF kernel at the segment level using dFC segments and at the subject level, employing the majority voting strategy described in Sect. 2.4.3.
Results
Static FC-based classification performance
Table 2 presents the classification accuracy based on sFC using SVM for both the augmented and baseline data modes. As shown in the augmented data mode, the mean accuracy for the ADHD/TDC classification increased from 63% in the balanced mode to 68% in the unbalanced mode, highlighting the inefficacy of the balancing strategy in enhancing performance. Moreover, the classification accuracy for distinguishing the ADHD subtypes was 55.15%, indicating a performance close to chance level. Furthermore, in the three-class classification mode, the overall mean accuracy was 43.4%, underscoring the challenges in effectively distinguishing between the ADHD subtypes and TDCs.
In the baseline data mode, within the two-class classification scenarios, the mean accuracy in the balanced mode decreased by 4%. The observed reduction indicates that the performance in the augmented data mode was likely inflated due to leakage (ranging from 8.1 to 18.4%) between the training and test sets. In the remaining scenarios, while the decrease in mean accuracy was less pronounced, there was a significant increase in the cross-fold standard deviation in accuracy. In the three-class scenario, the mean accuracy for the sFC-based classification using SVM dropped by 5%, highlighting the performance bias potentially introduced by leakage arising from the dependency between scans of the same individuals.
The confusion matrices illustrating the classification results are presented in Table 3 for the augmented data mode. Notably, in the initial two-class classification scenarios, the classifiers exhibited a tendency to misclassify more TDC as having ADHD, indicating a directional bias. In the classification between ADHDI and ADHDC, a notable pattern emerged wherein a higher frequency of misclassifications occurred from ADHDC to ADHDI. This observation suggests a potential underlying connection between common features associated with inattention present in both classes.
Network configuration parameters
Table 4 illustrates the impact of varying the number of dFC segments on classification accuracy in the unbalanced two-class mode for the augmented data mode. The performance evaluation was conducted utilizing the five-fold cross-validation strategy at the segment level, wherein all dFC segments from both the training and test subjects selected for the unbalanced mode (Sect. 2.5) were included. At this step, the number of filters was configured as 64, 128, 256, and 512 for the first to fourth Skip-Blocks, respectively. As illustrated in the Table 4, the highest accuracy was achieved with 7 dFC segments. As shown, classification accuracy of dFC segments for ADHD vs. TDC decreased approximately 4% compared to the other two cases.
Table 5 illustrates the impact of varying the number of filters in each Skip-Block on classification accuracy in the unbalanced two-class mode. With the optimal number of segments set to 7 (44×TR), the Skip-Vote-Net model achieved its highest mean classification accuracy of 85.87% when utilizing 64, 128, 256, and 512 filters in successive layers within the Skip-Blocks. Alternative configurations yielded suboptimal results. Notably, excluding the detail paths and skip connections resulted in a mean accuracy reduction of 3% and 8%, respectively, emphasizing the significance of these components in the classification process.
Segment-level classification performance
Tables 6 and 7 present the mean accuracy and confusion matrices for the Skip-Vote-Net and SVM on dFC segments (7 segments per subject). In the augmented data mode, each segment was assigned a class label by both classifiers at the segment level. The results indicate that SVM achieved a maximum mean accuracy of 68.46% in the unbalanced two-class (ADHD-TDC) mode. In comparison, the Skip-Vote-Net significantly enhanced classification accuracy by 30% in the two-class mode. For the three-class mode, the Skip-Vote-Net achieved a classification accuracy of 85.55%, representing a 39% improvement over SVM. In this data mode, the maximum misclassification rates were observed for Skip-Vote-Net when distinguishing between ADHD and TDC, as well as between ADHDI and ADHDC. Overall, SVM exhibited significantly higher misclassification rates between the ADHD subtypes and TDC compared to Skip-Vote-Net at the segment level.
In the baseline data mode, the accuracy of the SVM classifier decreased across all classification modes at the segment level, with a maximum mean accuracy reduction of 4.8% observed in the three-class classification mode, where there were also greater variations in accuracy across folds. This decline was attributed to leakage between the training and test sets. In contrast, the Skip-Vote-Net demonstrated a maximum reduction in cross-fold mean accuracy of less than 1.6% in the two-class (ADHDI - ADHDC) mode, indicating its greater robustness to leakage.
Subject-level classification performance
Tables 8 and 9 present the mean accuracy values and confusion matrices for Skip-Vote-Net and SVM at the subject level. These results were obtained by setting the optimal number of dFC segments to 7 for the majority voting stage and configuring the number of filters in the first to fourth Skip-Blocks to 64, 128, 256, and 512, respectively, based on the best-performing setup.
As demonstrated for the augmented data mode, the performance of SVM improved by up to 10% in the two-class and three-class modes using the dFC approach compared to the sFC-based classification (Table 2). However, the mean classification accuracy for the three-class mode did not exceed 53.21% at the subject level. In contrast, Skip-Vote-Net outperformed SVM using both sFC and dFC analyses. The proposed model achieved an average accuracy of up to 99.39% in distinguishing between ADHD subtypes, with an overall accuracy of 98.86% in the three-class mode. Additionally, no significant differences were observed between balanced and unbalanced classification modes using the proposed model.
In the baseline data mode, similar to the segment-level results, the accuracy of the SVM decreased significantly by nearly 10% in the three-class mode for dFC-based classification. This reduction was accompanied by greater variability in accuracy across folds, attributed to leakage at the subject level. In contrast, Skip-Vote-Net exhibited only a minimal reduction in cross-fold mean accuracy (up to 1.7%) coupled with higher robustness (indicated by lower variation in performance across folds) across all classification modes.
As illustrated in Table 9, the misclassification rates for SVM were substantially higher between ADHD and TDC, as well as between ADHDI and ADHDC. For the Skip-Vote-Net model, the maximum misclassification rate from the ADHD group to TDC was limited to 4% in the balanced mode, whereas SVM exhibited a misclassification rate of approximately 33%. This indicates that Skip-Vote-Net reduced the misclassification rate by 29% compared to SVM.
Discussion
Alterations in brain functional connectivity patterns among children with ADHD, as compared to typically developing children, have been a subject of investigation in brain connectome research4. Prior studies have found evidence suggesting that deviations in resting-state brain FC observed in children with ADHD can be utilized to effectively differentiate them from their typically developing peers43. In this study, we introduced Skip-Vote-Net, a DL-based architecture used for the classification of ADHD/TDC children using dFC patterns derived from rs-fMRI data. Our results showed that the Skip-Blocks enhanced the network ability to extract discriminative features using two distinct paths, which were subsequently fused to accentuate prominent discriminative characteristics, thereby facilitating accurate feature learning within the network. Moreover, the inclusion of the skip connections empowered the network with varying resolutions of information, ensuring the preservation of critical information and gradients that might otherwise diminish with increasing the network depth. In the final layer of the Skip-Vote-Net, a voting mechanism was employed to help the network in classifying each subject based on the majority class of segments. This approach served to enhance the robustness of the proposed method, particularly in scenarios involving misclassification of segments.
Several studies (Table 10) have documented an average accuracy of 73% through various methodologies in the classification of ADHD/TDC utilizing sFC analysis on the NYU Dataset8,18,43,44,45,46. The prevalent method for evaluating classification performance has been the hold-out strategy (216 in the training set and 41 in the test set) employing the AAL atlas with 90 regions. In our research, employing SVM, we achieved a mean accuracy of 68% (± 4%) via five-fold cross-validation over 216 subjects utilizing the AAL116 atlas. Although DL-based methodologies have demonstrated superior accuracy compared to SVM when utilizing sFC, they have typically been evaluated over a single fold. Conversely, in our SVM-based approach, the accuracy reached 72%, highlighting the performance dependence on the dataset.
Several studies (Table 10) have demonstrated incremental advancements in classification accuracy, generally falling within the range of 73% to 76%, employing dFC analysis and DL methodologies43,44,45. In our study, we significantly advanced upon these results by employing connectivity strengths from FC matrices divided into seven segments per subject. As demonstrated, the SVM performance improved by up to 10% in the two-class and three-class modes using the dFC approach compared to the sFC-based classification. However, the mean classification accuracy achieved by SVM for the three-class mode did not exceed 53.21% at the subject level. By integrating Skip-Vote-Net and employing majority voting with a five-fold cross-validation strategy, we achieved a mean accuracy of 97.73% at the subject level for the unbalanced two-class (ADHD/TDC) case. However, when altering the number of segments per subject, specifically utilizing five or nine segments based on common window lengths used in dFC analysis (typically ranging from 30 to 240 s)47,48, our accuracy decreased by 4%. Furthermore, our findings indicated no substantial performance improvement when balancing the number of samples across classes, aligning with previous studies that utilized ensemble sampling techniques for data balancing [50]. The Skip-Vote-Net, grounded in dFC analysis, notably decreased misclassification rates compared to sFC-based SVM, primarily due to majority voting on dynamic segments. Additionally, we observed that increasing the network size, as indicated by the number of filters per Skip-Block, enhanced the classification accuracy by up to 5%. However, further increasing network complexity led to a 2% decline in performance. The ablation experiments highlighted the importance of detailed paths and skip connections, contributing to a 3.6% and 8.1% improvement, respectively, in dFC-segment classification accuracy. Importantly, our approach uniquely focused on not only accurately distinguishing between ADHD and TDC children but also effectively classifying ADHD subtypes, achieving an exceptional mean accuracy of 98.86%. This marks a significant advancement in the field and underscores the potential of our method for precise clinical characterization of ADHD.
In this study, several limitations have been identified that should be addressed in future research. Initially, the evaluation of the proposed method was restricted to classifying subjects exclusively from the NYU dataset. The performance of the SVM and Skip-Vote-Net models was assessed using a data augmentation strategy involving two rs-fMRI scans for some participants - one for training and one for testing. This approach introduced a risk of data leakage between the training and test sets. Our results indicated a significant decrease in accuracy for the SVM model, while the Skip-Vote-Net model exhibited greater resilience to this leakage. However, the exclusion of the second scans from the test sets to mitigate data leakage led to increased cross-fold variability in accuracy due to the reduced number of scans in the test sets. To achieve a more comprehensive evaluation of the Skip-Vote-Net model’s performance on unseen data, it is essential to extend the analysis to include datasets from multiple centers within the ADHD-200 database. This broader approach will further allow the investigation of how different acquisition settings and parameters affect classification accuracy, thereby providing a more robust assessment of the model’s generalizability and effectiveness. Furthermore, the application of the Skip-Vote-Net method for classification with sFC matrices presented challenges due to its complex architecture, which resulted in convergence issues likely attributed to the limited sample sizes. Therefore, additional optimization efforts are required to refine the model’s configuration for improved performance using sFC features. Additionally, we utilized a low-resolution atlas comprising only 116 regions, which may affect classification accuracy. Recent studies4,49 have underscored the significance of functional parcellation at different scales, highlighting the potential influence on classification accuracy. Moreover, it has been demonstrated that high-resolution functional parcellation holds promise for capturing changes in connection strengths, particularly among smaller-sized brain structures46.
Conclusion
In conclusion, our study introduced Skip-Vote-Net, a deep learning-based architecture tailored for the classification of ADHD/TDC children using dFC patterns derived from rs-fMRI data. Our findings demonstrated the efficacy of Skip-Blocks in enhancing the network’s capability to extract discriminative features by employing two distinct paths, fused to emphasize salient discriminative characteristics. Furthermore, the integration of skip connections empowered the network with multi-resolution information, preserving crucial gradients and information that might diminish with increasing network depth. In the final layer of Skip-Vote-Net, a voting mechanism was deployed to aid the network in subject classification. Further investigation into the performance and generalization of Skip-Vote-Net across diverse datasets and clinical populations is warranted to fully assess its potential as a robust diagnostic tool in neuroimaging research.
Data availability
The dataset analyzed in the current study is publicly available in the ADHD-200 repository.
References
Blum, K. et al. Attention-deficit-hyperactivity disorder and reward defi ciency syndrome. Neurosychiat Dis. Treat. 5, 893–918 (2008).
De Silva, S., Dayarathna, S., Ariyarathne, G., Meedeniya, D. & Jayarathna, S. A survey of attention deficit hyperactivity disorder identification using psychophysiological data. Int. Assoc. Online Eng. 15(13), 61–76. https://doi.org/10.3991/ijoe.v15i13.10744 (2019).
Luo, Y., Weibman, D., Halperin, J. M. & Li, X. A review of heterogeneity in attention deficit/hyperactivity disorder (ADHD). Front. Hum. Neurosci. 13(February), 1–12. https://doi.org/10.3389/fnhum.2019.00042 (2019).
Ahmadi, M., Kazemi, K., Kuc, K., Cybulska-Klosowicz, A., Helfroush, M. S. & Aarabi, A. Resting state dynamic functional connectivity in children with attention deficit/hyperactivity disorder. J. Neural Eng.. 18(4), 0460d1 (2021). https://doi.org/10.1088/1741-2552/ac16b3.
Ahmadi, M., Kazemi, K., Kuc, K., Cybulska-Klosowicz, A., Helfroush, M. S. & Aarabi, A. Disrupted functional rich-club organization of the brain networks in children with attention-deficit/hyperactivity disorder, a resting-state EEG study. Brain Sci. 11(7), 2021. https://doi.org/10.3390/brainsci11070938.
Ahmadi, M. et al. Cortical source analysis of resting state EEG data in children with attention deficit hyperactivity disorder. Clin. Neurophysiol. 131(9), 2115–2130. https://doi.org/10.1016/j.clinph.2020.05.028 (2020).
A. P. Association and others. Diagnostic and statistical manual of mental disorders. Text Revis. (2000).
Dey, S., Rao, R. & Shah, M. Attributed graph distance measure for automatic detection of attention deficit hyperactive disordered subjects. Front. Neural Circuits 8(JUNE), 1–11. https://doi.org/10.3389/fncir.2014.00064 (2014).
Riaz, A., Asad, M., Alonso, E. & Slabaugh, G. DeepFMRI: End-to-end deep learning for functional connectivity and classification of ADHD using fMRI. J. Neurosci. Methods 335, 108506. https://doi.org/10.1016/j.jneumeth.2019.108506 (2020).
Zou, L., Zheng, J., Miao, C., McKeown, M. J. & Wang, Z. J. 3D CNN based automatic diagnosis of attention deficit hyperactivity disorder using functional and structural MRI. IEEE Access 5, 23626–23636. https://doi.org/10.1109/ACCESS.2017.2762703 (2017).
Van Den Heuvel, M. P. & Pol, H. E. H. Exploring the brain network: A review on resting-state fMRI functional connectivity. Eur. Neuropsychopharmacol. 20(8), 519–534. https://doi.org/10.1016/j.euroneuro.2010.03.008 (2010).
Liu, S. et al. Deep spatio-temporal representation and ensemble classification for attention deficit/hyperactivity disorder. IEEE Trans. Neural Syst. Rehabil. Eng. 29, 1–10. https://doi.org/10.1109/TNSRE.2020.3019063 (2021).
Hawi, Z., Yates, H., Kent, L., Gill, M. & Bellgrove, M. A case-control genome wide association study of childhood attention deficit hyperactivity disorder (Adhd). Eur. Neuropsychopharmacol. 29, S956. https://doi.org/10.1016/j.euroneuro.2017.08.311 (2019).
Slater, J. et al. Can electroencephalography (EEG) identify ADHD subtypes? A systematic review. Neurosci. Biobehav. Rev. 139(July), 2022. https://doi.org/10.1016/j.neubiorev.2022.104752 (2021).
Tang, Y. et al.. ADHD classification using auto-encoding neural network and binary hypothesis testing. Artif. Intell. Med. 123(January 2021), 102209. https://doi.org/10.1016/j.artmed.2021.102209 (2022).
Hoekzema, E. et al. An independent components and functional connectivity analysis of resting state fMRI data points to neural network dysregulation in adult ADHD. Hum. Brain Mapp. 35(4), 1261–1272 (2014).
Sun, Y., Zhao, L., Lan, Z., Jia, X. Z. & Xue, S. W. Differentiating boys with ADHD from those with typical development based on whole-brain functional connections using a machine learning approach. Neuropsychiatr. Dis. Treat. 16, 691–702. https://doi.org/10.2147/NDT.S239013 (2020).
Colby, J. B., Rudie, J. D., Brown, J. A., Douglas, P. K., Cohen, M. S. & Shehzad, Z. Insights into multimodal imaging classification of ADHD. Front. Syst. Neurosci. 6, 1–18. https://doi.org/10.3389/fnsys.2012.00059 (2012).
Y. Chen, Y. Tang, C. Wang, X. Liu, L. Zhao, and Z. Wang, “ADHD classification by dual subspace learning using resting-state functional connectivity. Artif. Intell. Med. 103(April 2019), 101786, https://doi.org/10.1016/j.artmed.2019.101786 (2020).
Zhao, Y., Chen, H. & Ogden, R. T. Wavelet-based weighted LASSO and screening approaches in functional linear regression. J. Comput. Graph. Stat. 24(3), 655–675 (2015).
Wang, M. et al. Graph-Kernel Based Structured Feature Selection for Brain Disease Classification Using Functional Connectivity Networks. IEEE Access 7, 35001–35011. https://doi.org/10.1109/ACCESS.2019.2903332 (2019).
Xie, J., Huo, Z., Liu, X. & Wang, Z. An fMRI sequence representation learning framework for attention deficit hyperactivity disorder classification. Appl. Sci. 12(12), 6211 (2022).
Quaak, M., van de Mortel, L., Thomas, R. M. & van Wingen, G. Deep learning applications for the classification of psychiatric disorders using neuroimaging data: Systematic review and meta-analysis. NeuroImage Clin. 30, 102584 (2021).
H. W. Loh, C. P. Ooi, P. D. Barua, E. E. Palmer, F. Molinari, and U. R. Acharya, “Automated detection of ADHD: Current trends and future perspective. Comput. Biol. Med. 146. https://doi.org/10.1016/j.compbiomed.2022.105525 (2022).
Mao, Z. et al. Spatio-temporal deep learning method for ADHD fMRI classification. Inf. Sci. (Ny) 499, 1–11. https://doi.org/10.1016/j.ins.2019.05.043 (2019).
Ji, J., Xing, X., Yao, Y., Li, J. & Zhang, X. Convolutional kernels with an element-wise weighting mechanism for identifying abnormal brain connectivity patterns. Pattern Recognit. 109, 107570 (2021).
Chen, M. et al. ConCeptCNN: A novel multi-filter convolutional neural network for the prediction of neurodevelopmental disorders using brain connectome. Med. Phys. 49(5), 3171–3184 (2022).
Gao, M.-S., Tsai, F.-S. & Lee, C.-C. Learning a phenotypic-attribute attentional brain connectivity embedding for ADHD classification using rs-fMRI. In 2020 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC) 5472–5475 (2020).
Qiang, N. et al. Deep variational autoencoder for mapping functional brain networks. IEEE Trans. Cogn. Dev. Syst. 13(4), 841–852 (2020).
Bellec, P. et al. The Neuro Bureau ADHD-200 preprocessed repository. Neuroimage 144, 275–286. https://doi.org/10.1016/j.neuroimage.2016.06.034 (2017).
Friston, K. J., Ashburner, J., Frith, C. D., Poline, J. ‐B., Heather, J. D. & Frackowiak, R. S. J. Spatial registration and normalization of images. Hum. Brain Mapp. 3(3), 165–189. https://doi.org/10.1002/hbm.460030303 (1995).
Jenkinson, M. & Smith, S. A global optimisation method for robust affine registration of brain images. Med. Image Anal. 5(2), 143–156. https://doi.org/10.1016/S1361-8415(01)00036-6 (2001).
Jenkinson, M., Bannister, P., Brady, M. & Smith, S. Improved optimization for the robust and accurate linear registration and motion correction of brain images. Neuroimage 17(2), 825–841. https://doi.org/10.1006/nimg.2002.1132 (2002).
Rueckert, D. Nonrigid registration using free-form deformations: Application to breast mr images. IEEE Trans. Med. Imaging 18(8), 712–721. https://doi.org/10.1109/42.796284 (1999).
Zhang, Y., Brady, M. & Smith, S. Segmentation of brain MR images through a hidden Markov random field model and the expectation-maximization algorithm. IEEE Trans. Med. Imaging 20(1), 45–57. https://doi.org/10.1109/42.906424 (2001).
Cox, R. W. AFNI: Software for analysis and visualization of functional magnetic resonance neuroimages. Comput. Biomed. Res. 29(3), 162–173. https://doi.org/10.1006/cbmr.1996.0014 (1996).
Rolls, E. T., Huang, C. C., Lin, C. P., Feng, J. & Joliot, M. Automated anatomical labelling atlas 3. Neuroimage 206, 116189.https://doi.org/10.1016/j.neuroimage.2019.116189 (2020).
Hutchison, R. M. et al. Dynamic functional connectivity: Promise, issues, and interpretations. Neuroimage 80, 360–378. https://doi.org/10.1016/j.neuroimage.2013.05.079 (2013).
Drozdzal, C., Vorontsov, M., Chartrand, E., Kadoury, G. & Pal, S. The importance of skip connections in biomedical image segmentation, , vol. 2, 179–187 (Springer, 2016). https://doi.org/10.1007/978-3-319-46976-8_19.
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18(2), 203–211. https://doi.org/10.1038/s41592-020-01008-z (2021).
Nirthika, R., Manivannan, S., Ramanan, A. & Wang, R. Pooling in convolutional neural networks for medical image analysis: a survey and an empirical study. Neural Comput. Appl. 34(7), 5321–5347 (2022).
He, K., Zhang, X., Ren, S. & Sun, J. Identity mappings in deep residual networks. In Computer Vision -- ECCV 2016 630–645 (2016).
Pei, S., Wang, C., Cao, S. & Lv, Z. Data augmentation for fMRI-based functional connectivity and its application to cross-site ADHD classification. IEEE Trans. Instrum. Meas. 72, 1–15. https://doi.org/10.1109/TIM.2022.3232670 (2023).
Firouzi, M., Ahmadi, M., Kazemi, K., Helfroush, M. S. & Aarabi, A. Deep Convolutional Neural Network for ADHD Classification Using Resting-State fMRI. In 2023 31st International Conference on Electrical Engineering (ICEE) 877–880. https://doi.org/10.1109/ICEE59167.2023.10334844 (2023).
Wang, M., Zhu, L., Li, X., Pan, Y. & Li, L. Dynamic functional connectivity analysis with temporal convolutional network for attention deficit/hyperactivity disorder identification. Front. Neurosci. 17. https://doi.org/10.3389/fnins.2023.1322967 (2023).
Khalilian, M. et al. Age-related differences in structural and resting-state functional brain network organization across the adult lifespan: A cross-sectional study. Aging Brain 5, 100105 (2024).
Khan, N. A., Waheeb, S. A., Riaz, A. & Shang, X. A novel knowledge distillation-based feature selection for the classification of ADHD. Biomolecules 11(8), 1–18. https://doi.org/10.3390/biom11081093 (2021).
Shao, L., Zhang, D., Du, H. & Fu, D. Deep forest in ADHD data classification. IEEE Access 7(2017), 137913–137919. https://doi.org/10.1109/ACCESS.2019.2941515 (2019).
Riaz, A. et al. Deep fMRI: AN end-to-end deep network for classification of fMRI data. Proc. - Int. Symp. Biomed. Imaging 2018, 1419–1422. https://doi.org/10.1109/ISBI.2018.8363838 (2018).
Author information
Authors and Affiliations
Contributions
Conceptualization, M.H. F., K.K., M.A. M.S. H., and A.A.; methodology, MH. F., K.K., M.A., M.S. H., and A.A.; software, M.H. F., M.A., K.K. and A.A; validation, M.H. F., K.K. and A.A; formal analysis, M.H. F. K.K., A.A.; writing—original draft preparation, M.H. F., M.A., and K.K.; writing—review and editing, M.H. F. and A.A.; visualization, MH. F.; supervision, K.K. and A.A.; project administration, K.K. and A.A.; All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Firouzi, M., Kazemi, K., Ahmadi, M. et al. Enhanced ADHD classification through deep learning and dynamic resting state fMRI analysis. Sci Rep 14, 24473 (2024). https://doi.org/10.1038/s41598-024-74282-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-74282-y
Keywords
This article is cited by
-
Applying artificial intelligence in neurodevelopmental disorders management and research
European Journal of Medical Research (2026)
-
Deep Learning Approaches to Evaluating ADHD Using EEG Data: RNN, GRU, and LSTM Models
Arabian Journal for Science and Engineering (2026)
-
Symptom-based depression subtypes: brain dynamic specificity and its association with gene expression profiles
Translational Psychiatry (2025)
-
Advances in magnetic resonance imaging of the developing brain and its applications in pediatrics
World Journal of Pediatrics (2025)